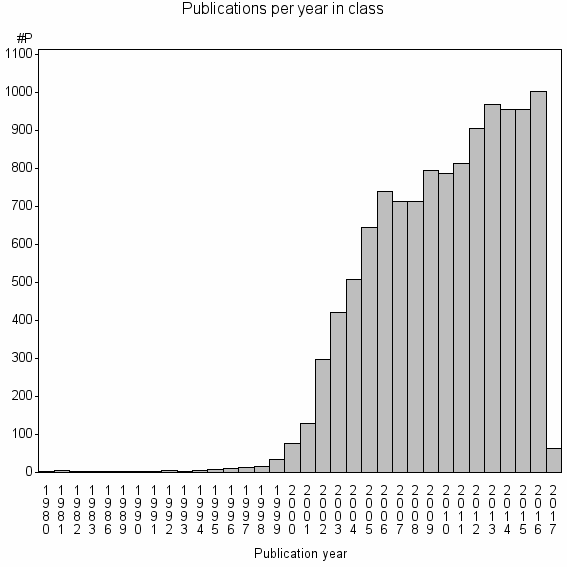
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 868 | 11572 | 39.6 | 79% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | BMC BIOINFORMATICS | journal | 456436 | 10% | 15% | 1140 |
| 2 | MATHEMATICAL & COMPUTATIONAL BIOLOGY | WoSSC | 452762 | 36% | 4% | 4196 |
| 3 | BIOINFORMATICS | journal | 441211 | 12% | 13% | 1335 |
| 4 | BICLUSTERING | authKW | 173996 | 1% | 63% | 105 |
| 5 | PROTEIN INTERACTION NETWORK | authKW | 144470 | 1% | 48% | 114 |
| 6 | BIOCHEMICAL RESEARCH METHODS | WoSSC | 124993 | 35% | 1% | 4021 |
| 7 | PROTEIN PROTEIN INTERACTION NETWORK | authKW | 124138 | 1% | 37% | 129 |
| 8 | GENE ONTOLOGY | authKW | 94748 | 1% | 21% | 170 |
| 9 | BMC SYSTEMS BIOLOGY | journal | 92047 | 2% | 15% | 237 |
| 10 | GENE EXPRESSION DATA | authKW | 78991 | 1% | 33% | 91 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 452762 | 36% | 4% | 4196 |
| 2 | Biochemical Research Methods | 124993 | 35% | 1% | 4021 |
| 3 | Biotechnology & Applied Microbiology | 77826 | 35% | 1% | 4056 |
| 4 | Genetics & Heredity | 8576 | 12% | 0% | 1446 |
| 5 | Statistics & Probability | 6134 | 6% | 0% | 706 |
| 6 | Computer Science, Interdisciplinary Applications | 5712 | 7% | 0% | 761 |
| 7 | Biochemistry & Molecular Biology | 5457 | 22% | 0% | 2504 |
| 8 | Medical Informatics | 2063 | 2% | 0% | 207 |
| 9 | Multidisciplinary Sciences | 1352 | 2% | 0% | 241 |
| 10 | Computer Science, Artificial Intelligence | 1333 | 3% | 0% | 395 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOINFORMAT | 73271 | 5% | 5% | 624 |
| 2 | BIOINFORMAT SCI TECHNOL | 57811 | 1% | 22% | 101 |
| 3 | SIBS NOVO NORDISK TRANSLAT PREDIABET | 29191 | 0% | 43% | 26 |
| 4 | COMP SCI | 23975 | 10% | 1% | 1215 |
| 5 | BIOMED INFORMAT | 22755 | 2% | 4% | 251 |
| 6 | CCSB | 22246 | 0% | 33% | 26 |
| 7 | ABORAT INNOVAT MATH MODELLING | 21915 | 0% | 40% | 21 |
| 8 | SYST BIOL | 17631 | 2% | 3% | 241 |
| 9 | SYST COHORT | 17572 | 0% | 67% | 10 |
| 10 | BIOINFORMAT PROGRAM | 16993 | 1% | 8% | 86 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BMC BIOINFORMATICS | 456436 | 10% | 15% | 1140 |
| 2 | BIOINFORMATICS | 441211 | 12% | 13% | 1335 |
| 3 | BMC SYSTEMS BIOLOGY | 92047 | 2% | 15% | 237 |
| 4 | IEEE-ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS | 46803 | 1% | 13% | 142 |
| 5 | JOURNAL OF COMPUTATIONAL BIOLOGY | 32938 | 1% | 9% | 140 |
| 6 | NUCLEIC ACIDS RESEARCH | 28508 | 6% | 2% | 663 |
| 7 | BRIEFINGS IN BIOINFORMATICS | 23538 | 1% | 11% | 79 |
| 8 | PLOS COMPUTATIONAL BIOLOGY | 22266 | 2% | 4% | 199 |
| 9 | STATISTICAL APPLICATIONS IN GENETICS AND MOLECULAR BIOLOGY | 21947 | 1% | 13% | 65 |
| 10 | INTERNATIONAL JOURNAL OF DATA MINING AND BIOINFORMATICS | 21194 | 1% | 14% | 59 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | BICLUSTERING | 173996 | 1% | 63% | 105 | Search BICLUSTERING | Search BICLUSTERING |
| 2 | PROTEIN INTERACTION NETWORK | 144470 | 1% | 48% | 114 | Search PROTEIN+INTERACTION+NETWORK | Search PROTEIN+INTERACTION+NETWORK |
| 3 | PROTEIN PROTEIN INTERACTION NETWORK | 124138 | 1% | 37% | 129 | Search PROTEIN+PROTEIN+INTERACTION+NETWORK | Search PROTEIN+PROTEIN+INTERACTION+NETWORK |
| 4 | GENE ONTOLOGY | 94748 | 1% | 21% | 170 | Search GENE+ONTOLOGY | Search GENE+ONTOLOGY |
| 5 | GENE EXPRESSION DATA | 78991 | 1% | 33% | 91 | Search GENE+EXPRESSION+DATA | Search GENE+EXPRESSION+DATA |
| 6 | ESSENTIAL PROTEINS | 74332 | 0% | 83% | 34 | Search ESSENTIAL+PROTEINS | Search ESSENTIAL+PROTEINS |
| 7 | NETWORK PHARMACOLOGY | 67131 | 0% | 58% | 44 | Search NETWORK+PHARMACOLOGY | Search NETWORK+PHARMACOLOGY |
| 8 | NETWORK ALIGNMENT | 60791 | 0% | 82% | 28 | Search NETWORK+ALIGNMENT | Search NETWORK+ALIGNMENT |
| 9 | MICROARRAY | 59931 | 5% | 4% | 585 | Search MICROARRAY | Search MICROARRAY |
| 10 | PROTEIN PROTEIN INTERACTION | 47628 | 3% | 5% | 343 | Search PROTEIN+PROTEIN+INTERACTION | Search PROTEIN+PROTEIN+INTERACTION |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | CSERMELY, P , KORCSMAROS, T , KISS, HJM , LONDON, G , NUSSINOV, R , (2013) STRUCTURE AND DYNAMICS OF MOLECULAR NETWORKS: A NOVEL PARADIGM OF DRUG DISCOVERY A COMPREHENSIVE REVIEW.PHARMACOLOGY & THERAPEUTICS. VOL. 138. ISSUE 3. P. 333 -408 | 389 | 36% | 205 |
| 2 | BARABASI, AL , GULBAHCE, N , LOSCALZO, J , (2011) NETWORK MEDICINE: A NETWORK-BASED APPROACH TO HUMAN DISEASE.NATURE REVIEWS GENETICS. VOL. 12. ISSUE 1. P. 56 -68 | 71 | 61% | 1018 |
| 3 | SNIDER, J , KOTLYAR, M , SARAON, P , YAO, Z , JURISICA, I , STAGLJAR, I , (2015) FUNDAMENTALS OF PROTEIN INTERACTION NETWORK MAPPING.MOLECULAR SYSTEMS BIOLOGY. VOL. 11. ISSUE 12. P. - | 130 | 67% | 9 |
| 4 | AL-HARAZI, O , AL INSAIF, S , AL-AJLAN, MA , KAYA, N , DZIMIRI, N , COLAK, D , (2016) INTEGRATED GENOMIC AND NETWORK-BASED ANALYSES OF COMPLEX DISEASES AND HUMAN DISEASE NETWORK.JOURNAL OF GENETICS AND GENOMICS. VOL. 43. ISSUE 6. P. 349 -367 | 125 | 68% | 1 |
| 5 | MITRA, K , CARVUNIS, AR , RAMESH, SK , IDEKER, T , (2013) INTEGRATIVE APPROACHES FOR FINDING MODULAR STRUCTURE IN BIOLOGICAL NETWORKS.NATURE REVIEWS GENETICS. VOL. 14. ISSUE 10. P. 719 -732 | 98 | 62% | 109 |
| 6 | SZKLARCZYK, D , FRANCESCHINI, A , WYDER, S , FORSLUND, K , HELLER, D , HUERTA-CEPAS, J , SIMONOVIC, M , ROTH, A , SANTOS, A , TSAFOU, KP , ET AL (2015) STRING V10: PROTEIN-PROTEIN INTERACTION NETWORKS, INTEGRATED OVER THE TREE OF LIFE.NUCLEIC ACIDS RESEARCH. VOL. 43. ISSUE D1. P. D447 -D452 | 25 | 68% | 556 |
| 7 | PETREY, D , HONIG, B , (2014) STRUCTURAL BIOINFORMATICS OF THE INTERACTOME.ANNUAL REVIEW OF BIOPHYSICS, VOL 43. VOL. 43. ISSUE . P. 193 -210 | 114 | 63% | 16 |
| 8 | CHEN, BL , FAN, WW , LIU, J , WU, FX , (2014) IDENTIFYING PROTEIN COMPLEXES AND FUNCTIONAL MODULES-FROM STATIC PPI NETWORKS TO DYNAMIC PPI NETWORKS.BRIEFINGS IN BIOINFORMATICS. VOL. 15. ISSUE 2. P. 177 -194 | 75 | 89% | 23 |
| 9 | WANG, JX , LI, M , PENG, W , WU, FX , LAN, W , (2015) COMPUTATIONAL APPROACHES FOR PRIORITIZING CANDIDATE DISEASE GENES BASED ON PPI NETWORKS.TSINGHUA SCIENCE AND TECHNOLOGY. VOL. 20. ISSUE 5. P. 500 -512 | 86 | 85% | 1 |
| 10 | SHARAN, R , ULITSKY, I , SHAMIR, R , (2007) NETWORK-BASED PREDICTION OF PROTEIN FUNCTION.MOLECULAR SYSTEMS BIOLOGY. VOL. 3. ISSUE . P. - | 66 | 81% | 388 |
Classes with closest relation at Level 2 |