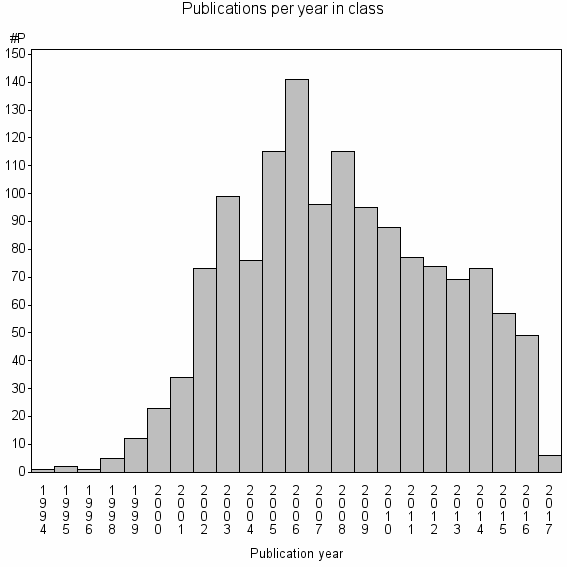
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 7323 | 1381 | 32.5 | 65% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 868 | 2 | BMC BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS | 11572 |
| 7323 | 1 | BICLUSTERING//GENE EXPRESSION DATA//GENE EXPRESSION DATA ANALYSIS | 1381 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | BICLUSTERING | authKW | 1404463 | 7% | 62% | 103 |
| 2 | GENE EXPRESSION DATA | authKW | 381283 | 5% | 25% | 69 |
| 3 | GENE EXPRESSION DATA ANALYSIS | authKW | 182543 | 1% | 49% | 17 |
| 4 | CO CLUSTERING | authKW | 134401 | 2% | 26% | 23 |
| 5 | BIOINFORMATICS | journal | 91616 | 15% | 2% | 210 |
| 6 | ORDER PRESERVING SUBMATRIX | authKW | 88438 | 0% | 100% | 4 |
| 7 | PLAID MODEL | authKW | 88438 | 0% | 100% | 4 |
| 8 | GENE EXPRESSION DATA CLUSTERING | authKW | 78959 | 0% | 71% | 5 |
| 9 | MATHEMATICAL & COMPUTATIONAL BIOLOGY | WoSSC | 70913 | 41% | 1% | 573 |
| 10 | TIME COURSE GENE EXPRESSION | authKW | 69088 | 0% | 63% | 5 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 70913 | 41% | 1% | 573 |
| 2 | Biochemical Research Methods | 17185 | 37% | 0% | 514 |
| 3 | Biotechnology & Applied Microbiology | 10491 | 37% | 0% | 513 |
| 4 | Computer Science, Artificial Intelligence | 4981 | 16% | 0% | 222 |
| 5 | Statistics & Probability | 3679 | 13% | 0% | 179 |
| 6 | Computer Science, Interdisciplinary Applications | 2795 | 13% | 0% | 174 |
| 7 | Computer Science, Theory & Methods | 970 | 8% | 0% | 109 |
| 8 | Computer Science, Information Systems | 748 | 7% | 0% | 91 |
| 9 | Medical Informatics | 692 | 3% | 0% | 40 |
| 10 | Genetics & Heredity | 296 | 7% | 0% | 103 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | KIT LANGUAGE TECHNOL DOCTORATE | 44219 | 0% | 100% | 2 |
| 2 | NACL TECNOL AGR IB INTA | 44219 | 0% | 100% | 2 |
| 3 | GRP GENE COMPUTAT | 29478 | 0% | 67% | 2 |
| 4 | MACHINE INTELLIGENCE UNIT | 29231 | 2% | 6% | 24 |
| 5 | FICH UNL | 28423 | 0% | 43% | 3 |
| 6 | LATICE | 27206 | 0% | 31% | 4 |
| 7 | ONCOL PHYSIOL BIOPHYS | 24869 | 0% | 38% | 3 |
| 8 | AIDS INFORMAT EDUC PROGRAM | 22109 | 0% | 100% | 1 |
| 9 | ARTIFICIAL INTELLIGENCE BIOINFORRNAT GRP | 22109 | 0% | 100% | 1 |
| 10 | B IT BONN AACHEN INT INFORMAT TECHNOL | 22109 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOINFORMATICS | 91616 | 15% | 2% | 210 |
| 2 | BMC BIOINFORMATICS | 57690 | 10% | 2% | 140 |
| 3 | IEEE-ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS | 31228 | 3% | 4% | 40 |
| 4 | INTERNATIONAL JOURNAL OF DATA MINING AND BIOINFORMATICS | 20481 | 1% | 5% | 20 |
| 5 | JOURNAL OF COMPUTATIONAL BIOLOGY | 11078 | 2% | 2% | 28 |
| 6 | ALGORITHMS FOR MOLECULAR BIOLOGY | 6311 | 1% | 3% | 9 |
| 7 | BIODATA MINING | 5145 | 1% | 3% | 7 |
| 8 | STATISTICAL APPLICATIONS IN GENETICS AND MOLECULAR BIOLOGY | 4359 | 1% | 2% | 10 |
| 9 | CURRENT BIOINFORMATICS | 3186 | 1% | 2% | 8 |
| 10 | JOURNAL OF BIOINFORMATICS AND COMPUTATIONAL BIOLOGY | 2163 | 0% | 2% | 6 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | BICLUSTERING | 1404463 | 7% | 62% | 103 | Search BICLUSTERING | Search BICLUSTERING |
| 2 | GENE EXPRESSION DATA | 381283 | 5% | 25% | 69 | Search GENE+EXPRESSION+DATA | Search GENE+EXPRESSION+DATA |
| 3 | GENE EXPRESSION DATA ANALYSIS | 182543 | 1% | 49% | 17 | Search GENE+EXPRESSION+DATA+ANALYSIS | Search GENE+EXPRESSION+DATA+ANALYSIS |
| 4 | CO CLUSTERING | 134401 | 2% | 26% | 23 | Search CO+CLUSTERING | Search CO+CLUSTERING |
| 5 | ORDER PRESERVING SUBMATRIX | 88438 | 0% | 100% | 4 | Search ORDER+PRESERVING+SUBMATRIX | Search ORDER+PRESERVING+SUBMATRIX |
| 6 | PLAID MODEL | 88438 | 0% | 100% | 4 | Search PLAID+MODEL | Search PLAID+MODEL |
| 7 | GENE EXPRESSION DATA CLUSTERING | 78959 | 0% | 71% | 5 | Search GENE+EXPRESSION+DATA+CLUSTERING | Search GENE+EXPRESSION+DATA+CLUSTERING |
| 8 | TIME COURSE GENE EXPRESSION | 69088 | 0% | 63% | 5 | Search TIME+COURSE+GENE+EXPRESSION | Search TIME+COURSE+GENE+EXPRESSION |
| 9 | BICLUSTERING ALGORITHM | 66328 | 0% | 100% | 3 | Search BICLUSTERING+ALGORITHM | Search BICLUSTERING+ALGORITHM |
| 10 | OPSM | 66328 | 0% | 100% | 3 | Search OPSM | Search OPSM |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | PONTES, B , GIRALDEZ, R , AGUILAR-RUIZ, JS , (2015) BICLUSTERING ON EXPRESSION DATA: A REVIEW.JOURNAL OF BIOMEDICAL INFORMATICS. VOL. 57. ISSUE . P. 163 -180 | 39 | 80% | 2 |
| 2 | MUKHOPADHYAY, A , MAULIK, U , BANDYOPADHYAY, S , (2010) ON BICLUSTERING OF GENE EXPRESSION DATA.CURRENT BIOINFORMATICS. VOL. 5. ISSUE 3. P. 204 -216 | 33 | 85% | 9 |
| 3 | FLORES, JL , INZA, I , LARRANAGA, P , CALVO, B , (2013) A NEW MEASURE FOR GENE EXPRESSION BICLUSTERING BASED ON NON-PARAMETRIC CORRELATION.COMPUTER METHODS AND PROGRAMS IN BIOMEDICINE. VOL. 112. ISSUE 3. P. 367 -397 | 33 | 67% | 13 |
| 4 | AYADI, W , ELLOURNI, M , HAO, JK , (2012) BIMINE+: AN EFFICIENT ALGORITHM FOR DISCOVERING RELEVANT BICLUSTERS OF DNA MICROARRAY DATA.KNOWLEDGE-BASED SYSTEMS. VOL. 35. ISSUE . P. 224 -234 | 25 | 86% | 4 |
| 5 | AYADI, W , ELLOUMI, M , HAO, JK , (2012) BICFINDER: A BICLUSTERING ALGORITHM FOR MICROARRAY DATA ANALYSIS.KNOWLEDGE AND INFORMATION SYSTEMS. VOL. 30. ISSUE 2. P. 341 -358 | 24 | 89% | 6 |
| 6 | OGHABIAN, A , KILPINEN, S , HAUTANIEMI, S , CZEIZLER, E , (2014) BICLUSTERING METHODS: BIOLOGICAL RELEVANCE AND APPLICATION IN GENE EXPRESSION ANALYSIS.PLOS ONE. VOL. 9. ISSUE 3. P. - | 23 | 77% | 15 |
| 7 | AYADI, W , ELLOUMI, M , HAO, JK , (2009) A BICLUSTERING ALGORITHM BASED ON A BICLUSTER ENUMERATION TREE: APPLICATION TO DNA MICROARRAY DATA.BIODATA MINING. VOL. 2. ISSUE . P. - | 22 | 92% | 17 |
| 8 | NEPOMUCENOA, JA , TRONCOSO, A , AGUILAR-RUIZ, JS , (2015) SCATTER SEARCH-BASED IDENTIFICATION OF LOCAL PATTERNS WITH POSITIVE AND NEGATIVE CORRELATIONS IN GENE EXPRESSION DATA.APPLIED SOFT COMPUTING. VOL. 35. ISSUE . P. 637 -651 | 23 | 79% | 0 |
| 9 | COOKE, EJ , SAVAGE, RS , KIRK, PDW , DARKINS, R , WILD, DL , (2011) BAYESIAN HIERARCHICAL CLUSTERING FOR MICROARRAY TIME SERIES DATA WITH REPLICATES AND OUTLIER MEASUREMENTS.BMC BIOINFORMATICS. VOL. 12. ISSUE . P. - | 22 | 79% | 14 |
| 10 | AYADI, W , HAO, JK , (2014) A MEMETIC ALGORITHM FOR DISCOVERING NEGATIVE CORRELATION BICLUSTERS OF DNA MICROARRAY DATA.NEUROCOMPUTING. VOL. 145. ISSUE . P. 14 -22 | 18 | 90% | 3 |
Classes with closest relation at Level 1 |