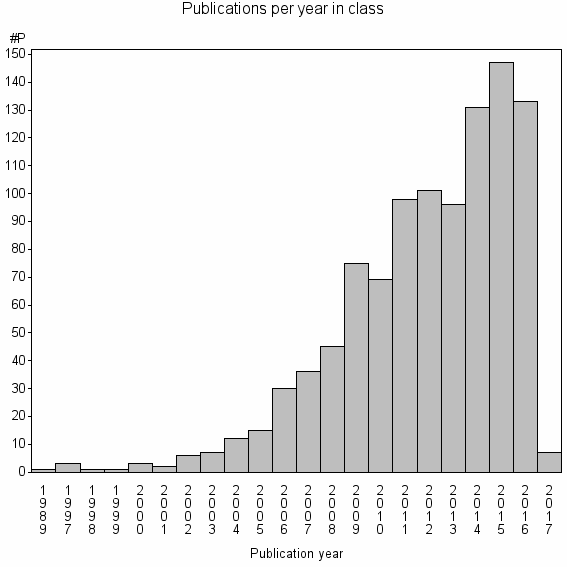
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 10975 | 1019 | 42.7 | 83% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 868 | 2 | BMC BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS | 11572 |
| 10975 | 1 | GENE PRIORITIZATION//DISEASE GENE PREDICTION//HUMAN DISEASE NETWORK | 1019 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | GENE PRIORITIZATION | authKW | 220735 | 2% | 39% | 19 |
| 2 | DISEASE GENE PREDICTION | authKW | 191769 | 1% | 80% | 8 |
| 3 | HUMAN DISEASE NETWORK | authKW | 154101 | 1% | 86% | 6 |
| 4 | SKARNES GRP | address | 149822 | 0% | 100% | 5 |
| 5 | DISEASOME | authKW | 147511 | 1% | 62% | 8 |
| 6 | DISEASE SIMILARITY | authKW | 124850 | 0% | 83% | 5 |
| 7 | DISEASE GENES | authKW | 105498 | 2% | 20% | 18 |
| 8 | ZFIN | address | 95877 | 1% | 40% | 8 |
| 9 | NETWORK MEDICINE | authKW | 86279 | 1% | 24% | 12 |
| 10 | MOUSE INFORMAT GRP | address | 81561 | 1% | 39% | 7 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 35944 | 34% | 0% | 351 |
| 2 | Biochemical Research Methods | 6422 | 27% | 0% | 273 |
| 3 | Biotechnology & Applied Microbiology | 6218 | 33% | 0% | 341 |
| 4 | Genetics & Heredity | 4725 | 29% | 0% | 294 |
| 5 | Biochemistry & Molecular Biology | 844 | 27% | 0% | 274 |
| 6 | Medical Informatics | 329 | 2% | 0% | 24 |
| 7 | Multidisciplinary Sciences | 189 | 3% | 0% | 26 |
| 8 | Computer Science, Interdisciplinary Applications | 161 | 4% | 0% | 41 |
| 9 | Cell Biology | 46 | 6% | 0% | 57 |
| 10 | Biology | 38 | 2% | 0% | 25 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | SKARNES GRP | 149822 | 0% | 100% | 5 |
| 2 | ZFIN | 95877 | 1% | 40% | 8 |
| 3 | MOUSE INFORMAT GRP | 81561 | 1% | 39% | 7 |
| 4 | BIOINFORMAT SYST ENGN BASE | 79903 | 0% | 67% | 4 |
| 5 | SYMBIOSYS COMPUTAT SYST BIOL | 59929 | 0% | 100% | 2 |
| 6 | BIOINFORMAT SCI TECHNOL | 54256 | 3% | 6% | 29 |
| 7 | MED GENET HUMAN GENET | 48917 | 1% | 12% | 14 |
| 8 | OSUCCC BIOMED INFORMAT SHARED OURCE | 39951 | 0% | 67% | 2 |
| 9 | COMPLEX NETWORKS | 35500 | 1% | 15% | 8 |
| 10 | BIOINTELLIGENCE DATA MIN | 33706 | 0% | 38% | 3 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOINFORMATICS | 25336 | 9% | 1% | 95 |
| 2 | JOURNAL OF BIOMEDICAL SEMANTICS | 24242 | 1% | 6% | 14 |
| 3 | BMC SYSTEMS BIOLOGY | 21556 | 3% | 2% | 34 |
| 4 | BMC BIOINFORMATICS | 20636 | 7% | 1% | 72 |
| 5 | DATABASE-THE JOURNAL OF BIOLOGICAL DATABASES AND CURATION | 20258 | 2% | 3% | 21 |
| 6 | NUCLEIC ACIDS RESEARCH | 7812 | 10% | 0% | 102 |
| 7 | BIODATA MINING | 6978 | 1% | 3% | 7 |
| 8 | BMC GENOMICS | 5470 | 4% | 0% | 42 |
| 9 | JOURNAL OF BIOMEDICAL INFORMATICS | 4560 | 1% | 1% | 15 |
| 10 | PLOS COMPUTATIONAL BIOLOGY | 3696 | 2% | 1% | 24 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | GENE PRIORITIZATION | 220735 | 2% | 39% | 19 | Search GENE+PRIORITIZATION | Search GENE+PRIORITIZATION |
| 2 | DISEASE GENE PREDICTION | 191769 | 1% | 80% | 8 | Search DISEASE+GENE+PREDICTION | Search DISEASE+GENE+PREDICTION |
| 3 | HUMAN DISEASE NETWORK | 154101 | 1% | 86% | 6 | Search HUMAN+DISEASE+NETWORK | Search HUMAN+DISEASE+NETWORK |
| 4 | DISEASOME | 147511 | 1% | 62% | 8 | Search DISEASOME | Search DISEASOME |
| 5 | DISEASE SIMILARITY | 124850 | 0% | 83% | 5 | Search DISEASE+SIMILARITY | Search DISEASE+SIMILARITY |
| 6 | DISEASE GENES | 105498 | 2% | 20% | 18 | Search DISEASE+GENES | Search DISEASE+GENES |
| 7 | NETWORK MEDICINE | 86279 | 1% | 24% | 12 | Search NETWORK+MEDICINE | Search NETWORK+MEDICINE |
| 8 | DISEASE GENE IDENTIFICATION | 68096 | 0% | 45% | 5 | Search DISEASE+GENE+IDENTIFICATION | Search DISEASE+GENE+IDENTIFICATION |
| 9 | CANDIDATE DISEASE GENES | 67419 | 0% | 75% | 3 | Search CANDIDATE+DISEASE+GENES | Search CANDIDATE+DISEASE+GENES |
| 10 | CANDIDATE GENE PREDICTION | 67419 | 0% | 75% | 3 | Search CANDIDATE+GENE+PREDICTION | Search CANDIDATE+GENE+PREDICTION |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | DONCHEVA, NT , KACPROWSKI, T , ALBRECHT, M , (2012) RECENT APPROACHES TO THE PRIORITIZATION OF CANDIDATE DISEASE GENES.WILEY INTERDISCIPLINARY REVIEWS-SYSTEMS BIOLOGY AND MEDICINE. VOL. 4. ISSUE 5. P. 429-442 | 66 | 62% | 18 |
| 2 | GILL, N , SINGH, S , ASERI, TC , (2014) COMPUTATIONAL DISEASE GENE PRIORITIZATION: AN APPRAISAL.JOURNAL OF COMPUTATIONAL BIOLOGY. VOL. 21. ISSUE 6. P. 456 -465 | 40 | 87% | 8 |
| 3 | PIRO, RM , DI CUNTO, F , (2012) COMPUTATIONAL APPROACHES TO DISEASE-GENE PREDICTION: RATIONALE, CLASSIFICATION AND SUCCESSES.FEBS JOURNAL. VOL. 279. ISSUE 5. P. 678 -696 | 51 | 50% | 51 |
| 4 | ZHU, C , WU, C , ARONOW, BJ , JEGGA, AG , (2014) COMPUTATIONAL APPROACHES FOR HUMAN DISEASE GENE PREDICTION AND RANKING.SYSTEMS ANALYSIS OF HUMAN MULTIGENE DISORDERS. VOL. 799. ISSUE . P. 69-84 | 43 | 66% | 2 |
| 5 | MOREAU, Y , TRANCHEVENT, LC , (2012) COMPUTATIONAL TOOLS FOR PRIORITIZING CANDIDATE GENES: BOOSTING DISEASE GENE DISCOVERY.NATURE REVIEWS GENETICS. VOL. 13. ISSUE 8. P. 523-536 | 46 | 41% | 140 |
| 6 | XIAO, Y , XU, CH , PING, YY , GUAN, JX , FAN, HH , LI, YQ , LI, X , (2011) DIFFERENTIAL EXPRESSION PATTERN-BASED PRIORITIZATION OF CANDIDATE GENES THROUGH INTEGRATING DISEASE-SPECIFIC EXPRESSION DATA.GENOMICS. VOL. 98. ISSUE 1. P. 64-71 | 35 | 80% | 1 |
| 7 | WANG, JX , LI, M , PENG, W , WU, FX , LAN, W , (2015) COMPUTATIONAL APPROACHES FOR PRIORITIZING CANDIDATE DISEASE GENES BASED ON PPI NETWORKS.TSINGHUA SCIENCE AND TECHNOLOGY. VOL. 20. ISSUE 5. P. 500 -512 | 48 | 48% | 1 |
| 8 | BARABASI, AL , GULBAHCE, N , LOSCALZO, J , (2011) NETWORK MEDICINE: A NETWORK-BASED APPROACH TO HUMAN DISEASE.NATURE REVIEWS GENETICS. VOL. 12. ISSUE 1. P. 56 -68 | 31 | 26% | 1018 |
| 9 | ZHANG, SW , SHAO, DD , ZHANG, SY , WANG, YB , (2014) PRIORITIZATION OF CANDIDATE DISEASE GENES BY ENLARGING THE SEED SET AND FUSING INFORMATION OF THE NETWORK TOPOLOGY AND GENE EXPRESSION.MOLECULAR BIOSYSTEMS. VOL. 10. ISSUE 6. P. 1400 -1408 | 44 | 49% | 4 |
| 10 | TRANCHEVENT, LC , CAPDEVILA, FB , NITSCH, D , DE MOOR, B , DE CAUSMAECKER, P , MOREAU, Y , (2011) A GUIDE TO WEB TOOLS TO PRIORITIZE CANDIDATE GENES.BRIEFINGS IN BIOINFORMATICS. VOL. 12. ISSUE 1. P. 22-32 | 34 | 57% | 66 |
Classes with closest relation at Level 1 |