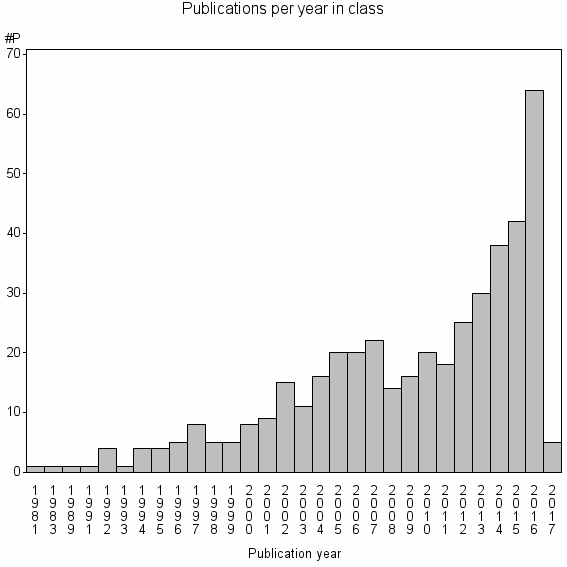
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 21112 | 433 | 38.2 | 84% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 868 | 2 | BMC BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS | 11572 |
| 21112 | 1 | PAULLONES//ADV BIOINFORMAT SYST MED//NCI 60 | 433 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | PAULLONES | authKW | 511906 | 3% | 52% | 14 |
| 2 | ADV BIOINFORMAT SYST MED | address | 317335 | 1% | 75% | 6 |
| 3 | NCI 60 | authKW | 218776 | 3% | 28% | 11 |
| 4 | CLUSTERED IMAGE MAP | authKW | 211559 | 1% | 100% | 3 |
| 5 | DRUG SENSITIVITY PREDICTION | authKW | 211559 | 1% | 100% | 3 |
| 6 | INFORMAT TECHNOL BRANCH | address | 147101 | 3% | 19% | 11 |
| 7 | CANCER CELL LINE MODELING | authKW | 141039 | 0% | 100% | 2 |
| 8 | CELL LINE PROFILING | authKW | 141039 | 0% | 100% | 2 |
| 9 | DARPONES | authKW | 141039 | 0% | 100% | 2 |
| 10 | FREDERICK CANC DEV SAIC | address | 141039 | 0% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Oncology | 2052 | 36% | 0% | 157 |
| 2 | Mathematical & Computational Biology | 1210 | 10% | 0% | 43 |
| 3 | Chemistry, Medicinal | 756 | 13% | 0% | 56 |
| 4 | Biochemical Research Methods | 495 | 12% | 0% | 52 |
| 5 | Biotechnology & Applied Microbiology | 332 | 13% | 0% | 56 |
| 6 | Computer Science, Interdisciplinary Applications | 159 | 6% | 0% | 25 |
| 7 | Pharmacology & Pharmacy | 136 | 13% | 0% | 55 |
| 8 | Chemistry, Organic | 77 | 8% | 0% | 33 |
| 9 | Genetics & Heredity | 64 | 6% | 0% | 28 |
| 10 | Cell Biology | 56 | 8% | 0% | 34 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ADV BIOINFORMAT SYST MED | 317335 | 1% | 75% | 6 |
| 2 | INFORMAT TECHNOL BRANCH | 147101 | 3% | 19% | 11 |
| 3 | FREDERICK CANC DEV SAIC | 141039 | 0% | 100% | 2 |
| 4 | GENOM BIOINFORMAT GRP | 124161 | 2% | 20% | 9 |
| 5 | COURS ARGONNE 229 | 94025 | 0% | 67% | 2 |
| 6 | INFORMAT TECHNOL BRANCHDEV THER EUT PROGRAM | 94025 | 0% | 67% | 2 |
| 7 | BASIC SCI MOL PHARMACOL | 70520 | 0% | 100% | 1 |
| 8 | CANC THER Y EVALUAT PROGRAMBIOMETR BRANCH | 70520 | 0% | 100% | 1 |
| 9 | CANCER EPIGENET DRUG DISCOVERY | 70520 | 0% | 100% | 1 |
| 10 | CAND GENET DEV BIOL | 70520 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOLECULAR CANCER THERAPEUTICS | 4420 | 4% | 0% | 16 |
| 2 | CANCER DISCOVERY | 2445 | 1% | 1% | 4 |
| 3 | BIOINFORMATICS | 1890 | 4% | 0% | 17 |
| 4 | ASSAY AND DRUG DEVELOPMENT TECHNOLOGIES | 1644 | 1% | 1% | 4 |
| 5 | NATURE REVIEWS CANCER | 1049 | 1% | 0% | 4 |
| 6 | NATURE BIOTECHNOLOGY | 959 | 1% | 0% | 6 |
| 7 | JOURNAL OF CHEMICAL INFORMATION AND MODELING | 807 | 1% | 0% | 6 |
| 8 | DOKLADY BIOCHEMISTRY AND BIOPHYSICS | 706 | 1% | 0% | 3 |
| 9 | EUROPEAN JOURNAL OF MEDICINAL CHEMISTRY | 599 | 2% | 0% | 9 |
| 10 | BULLETIN DU CANCER | 594 | 1% | 0% | 6 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | CORTES-CIRIANO, I , MERVIN, LH , BENDER, A , (2016) CURRENT TRENDS IN DRUG SENSITIVITY PREDICTION.CURRENT PHARMACEUTICAL DESIGN. VOL. 22. ISSUE 46. P. 6918 -6927 | 41 | 44% | 1 |
| 2 | WAN, P , LI, QY , LARSEN, JEP , EKLUND, AC , PARLESAK, A , RIGINA, O , NIELSEN, SJ , BJORKLING, F , JONSDOTTIR, SO , (2012) PREDICTION OF DRUG EFFICACY FOR CANCER TREATMENT BASED ON COMPARATIVE ANALYSIS OF CHEMOSENSITIVITY AND GENE EXPRESSION DATA.BIOORGANIC & MEDICINAL CHEMISTRY. VOL. 20. ISSUE 1. P. 167 -176 | 25 | 60% | 2 |
| 3 | BELIZARIO, JE , SANGIULIANO, BA , PEREZ-SOSA, M , NEYRA, JM , MOREIRA, DF , (2016) USING PHARMACOGENOMIC DATABASES FOR DISCOVERING PATIENT-TARGET GENES AND SMALL MOLECULE CANDIDATES TO CANCER THERAPY.FRONTIERS IN PHARMACOLOGY. VOL. 7. ISSUE . P. - | 33 | 31% | 1 |
| 4 | REINHOLD, WC , VARMA, S , RAJAPAKSE, VN , LUNA, A , SOUSA, FG , KOHN, KW , POMMIER, YG , (2015) USING DRUG RESPONSE DATA TO IDENTIFY MOLECULAR EFFECTORS, AND MOLECULAR "OMIC" DATA TO IDENTIFY CANDIDATE DRUGS IN CANCER.HUMAN GENETICS. VOL. 134. ISSUE 1. P. 3 -11 | 24 | 39% | 7 |
| 5 | TOLLE, N , KUNICK, C , (2011) PAULLONES AS INHIBITORS OF PROTEIN KINASES.CURRENT TOPICS IN MEDICINAL CHEMISTRY. VOL. 11. ISSUE 11. P. 1320-1332 | 33 | 32% | 12 |
| 6 | CORTES-CIRIANO, I , VAN WESTEN, GJP , BOUVIER, G , NILGES, M , OVERINGTON, JP , BENDER, A , MALLIAVIN, TE , (2016) IMPROVED LARGE-SCALE PREDICTION OF GROWTH INHIBITION PATTERNS USING THE NCI60 CANCER CELL LINE PANEL.BIOINFORMATICS. VOL. 32. ISSUE 1. P. 85 -95 | 18 | 41% | 4 |
| 7 | YUAN, H , PASKOV, I , PASKOV, H , GONZALEZ, AJ , LESLIE, CS , (2016) MULTITASK LEARNING IMPROVES PREDICTION OF CANCER DRUG SENSITIVITY.SCIENTIFIC REPORTS. VOL. 6. ISSUE . P. - | 16 | 55% | 0 |
| 8 | UITDEHAAG, JCM , DE ROOS, JADM , PRINSEN, MBW , WILLEMSEN-SEEGERS, N , DE VETTER, JRF , DYLUS, J , VAN DOORNMALEN, AM , KOOIJMAN, J , SAWA, M , VAN GERWEN, SJC , ET AL (2016) CELL PANEL PROFILING REVEALS CONSERVED THERAPEUTIC CLUSTERS AND DIFFERENTIATES THE MECHANISM OF ACTION OF DIFFERENT PI3K/MTOR, AURORA KINASE AND EZH2 INHIBITORS.MOLECULAR CANCER THERAPEUTICS. VOL. 15. ISSUE 12. P. 3097 -3109 | 18 | 45% | 0 |
| 9 | SCHERF, U , ROSS, DT , WALTHAM, M , SMITH, LH , LEE, JK , TANABE, L , KOHN, KW , REINHOLD, WC , MYERS, TG , ANDREWS, DT , ET AL (2000) A GENE EXPRESSION DATABASE FOR THE MOLECULAR PHARMACOLOGY OF CANCER.NATURE GENETICS. VOL. 24. ISSUE 3. P. 236-244 | 15 | 41% | 935 |
| 10 | KULESSKIY, E , SAARELA, J , TURUNEN, L , WENNERBERG, K , (2016) PRECISION CANCER MEDICINE IN THE ACOUSTIC DISPENSING ERA: EX VIVO PRIMARY CELL DRUG SENSITIVITY TESTING.JALA. VOL. 21. ISSUE 1. P. 27 -36 | 11 | 58% | 3 |
Classes with closest relation at Level 1 |