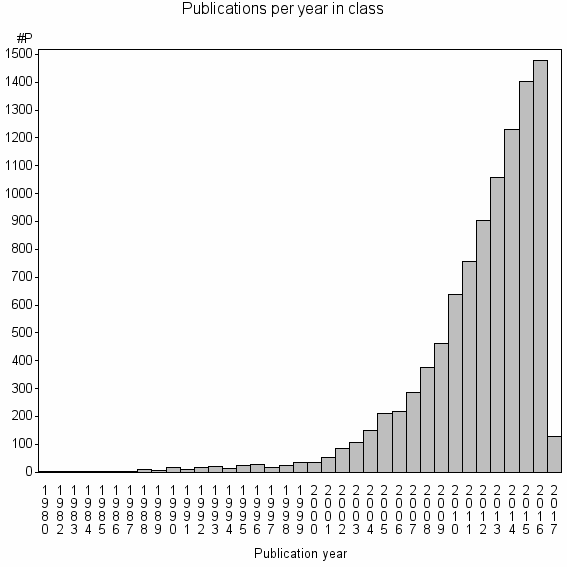
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 1101 | 9808 | 39.0 | 86% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 1101 | 2 | BIOINFORMATICS//COPY NUMBER VARIATION//NEXT GENERATION SEQUENCING | 9808 |
| 2013 | 1 | GENOME ASSEMBLY//BIOINFORMATICS//VARIANT CALLING | 2386 |
| 3965 | 1 | EVOLUT CANC//CANCER GENOME//CANCER GENOMICS | 1873 |
| 5569 | 1 | COPY NUMBER VARIATION//COPY NUMBER VARIANTS//STRUCTURAL VARIATION | 1604 |
| 7400 | 1 | RNA SEQ//TRANSCRIPTOME ASSEMBLY//RNA SEQUENCING | 1371 |
| 12230 | 1 | DNA COPY NUMBER//HLTH SOLUT GRP//ARRAY CGH | 922 |
| 16413 | 1 | WHOLE GENOME AMPLIFICATION//MULTIPLE DISPLACEMENT AMPLIFICATION//PHI29 DNA POLYMERASE | 655 |
| 19243 | 1 | TRANS SPLICING//SPLICED LEADER//CHIMERIC RNA | 515 |
| 19974 | 1 | SINGLE CELL RNA SEQ//SINGLE CELL RNA SEQUENCING//SINGLE CELL TRANSCRIPTOMICS | 482 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | BIOINFORMATICS | journal | 238336 | 9% | 9% | 904 |
| 2 | COPY NUMBER VARIATION | authKW | 216896 | 3% | 26% | 264 |
| 3 | NEXT GENERATION SEQUENCING | authKW | 167773 | 5% | 11% | 485 |
| 4 | RNA SEQ | authKW | 141260 | 3% | 14% | 330 |
| 5 | MATHEMATICAL & COMPUTATIONAL BIOLOGY | WoSSC | 140936 | 22% | 2% | 2168 |
| 6 | WHOLE GENOME AMPLIFICATION | authKW | 127831 | 1% | 46% | 90 |
| 7 | BMC BIOINFORMATICS | journal | 97394 | 5% | 7% | 486 |
| 8 | BIOTECHNOLOGY & APPLIED MICROBIOLOGY | WoSSC | 73850 | 37% | 1% | 3628 |
| 9 | BMC GENOMICS | journal | 72239 | 5% | 5% | 473 |
| 10 | CANCER GENOME | authKW | 66074 | 0% | 59% | 36 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 140936 | 22% | 2% | 2168 |
| 2 | Biotechnology & Applied Microbiology | 73850 | 37% | 1% | 3628 |
| 3 | Genetics & Heredity | 65066 | 34% | 1% | 3354 |
| 4 | Biochemical Research Methods | 56422 | 26% | 1% | 2515 |
| 5 | Biochemistry & Molecular Biology | 4806 | 22% | 0% | 2153 |
| 6 | Oncology | 3440 | 12% | 0% | 1140 |
| 7 | Statistics & Probability | 1183 | 3% | 0% | 315 |
| 8 | Cell Biology | 930 | 7% | 0% | 693 |
| 9 | Multidisciplinary Sciences | 684 | 2% | 0% | 164 |
| 10 | Pathology | 419 | 3% | 0% | 252 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | EVOLUT CANC | 42853 | 0% | 38% | 36 |
| 2 | GENOME SCI | 34102 | 2% | 5% | 239 |
| 3 | BIOINFORMAT COMPUTAT BIOL | 31808 | 1% | 8% | 131 |
| 4 | SIMONS QUANTITAT BIOL | 29632 | 0% | 53% | 18 |
| 5 | CANC GENOME PROJECT | 27724 | 0% | 21% | 43 |
| 6 | INTEGRATED GENOME POLYMORPHISM | 21431 | 0% | 34% | 20 |
| 7 | PERSONALIZED CANC THER Y | 18440 | 0% | 25% | 24 |
| 8 | CANC GENOME DISCOVERY | 17058 | 0% | 17% | 33 |
| 9 | PROGRAM COMPUTAT BIOL BIOINFORMAT | 16929 | 0% | 12% | 47 |
| 10 | STANLEY COGNIT GENOM | 16649 | 0% | 36% | 15 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOINFORMATICS | 238336 | 9% | 9% | 904 |
| 2 | BMC BIOINFORMATICS | 97394 | 5% | 7% | 486 |
| 3 | BMC GENOMICS | 72239 | 5% | 5% | 473 |
| 4 | GENOME RESEARCH | 56967 | 2% | 12% | 159 |
| 5 | GENOME BIOLOGY | 56797 | 2% | 8% | 234 |
| 6 | GIGASCIENCE | 32356 | 0% | 27% | 39 |
| 7 | NATURE METHODS | 28996 | 1% | 7% | 126 |
| 8 | BIOTECHFORUM ( BTF ) | 18892 | 1% | 5% | 126 |
| 9 | GENOME MEDICINE | 18769 | 1% | 10% | 63 |
| 10 | BRIEFINGS IN BIOINFORMATICS | 15476 | 1% | 8% | 59 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | COPY NUMBER VARIATION | 216896 | 3% | 26% | 264 | Search COPY+NUMBER+VARIATION | Search COPY+NUMBER+VARIATION |
| 2 | NEXT GENERATION SEQUENCING | 167773 | 5% | 11% | 485 | Search NEXT+GENERATION+SEQUENCING | Search NEXT+GENERATION+SEQUENCING |
| 3 | RNA SEQ | 141260 | 3% | 14% | 330 | Search RNA+SEQ | Search RNA+SEQ |
| 4 | WHOLE GENOME AMPLIFICATION | 127831 | 1% | 46% | 90 | Search WHOLE+GENOME+AMPLIFICATION | Search WHOLE+GENOME+AMPLIFICATION |
| 5 | CANCER GENOME | 66074 | 0% | 59% | 36 | Search CANCER+GENOME | Search CANCER+GENOME |
| 6 | GENOME ASSEMBLY | 62503 | 1% | 37% | 54 | Search GENOME+ASSEMBLY | Search GENOME+ASSEMBLY |
| 7 | MULTIPLE DISPLACEMENT AMPLIFICATION | 59222 | 0% | 48% | 40 | Search MULTIPLE+DISPLACEMENT+AMPLIFICATION | Search MULTIPLE+DISPLACEMENT+AMPLIFICATION |
| 8 | TRANS SPLICING | 55910 | 1% | 25% | 71 | Search TRANS+SPLICING | Search TRANS+SPLICING |
| 9 | VARIANT CALLING | 55333 | 0% | 68% | 26 | Search VARIANT+CALLING | Search VARIANT+CALLING |
| 10 | COPY NUMBER VARIANTS | 48012 | 1% | 23% | 68 | Search COPY+NUMBER+VARIANTS | Search COPY+NUMBER+VARIANTS |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | CONESA, A , MADRIGAL, P , TARAZONA, S , GOMEZ-CABRERO, D , CERVERA, A , MCPHERSON, A , SZCZESNIAK, MW , GAFFNEY, DJ , ELO, LL , ZHANG, XG , ET AL (2016) A SURVEY OF BEST PRACTICES FOR RNA-SEQ DATA ANALYSIS.GENOME BIOLOGY. VOL. 17. ISSUE . P. - | 121 | 63% | 23 |
| 2 | LAEHNEMANN, D , BORKHARDT, A , MCHARDY, AC , (2016) DENOISING DNA DEEP SEQUENCING DATA-HIGH-THROUGHPUT SEQUENCING ERRORS AND THEIR CORRECTION.BRIEFINGS IN BIOINFORMATICS. VOL. 17. ISSUE 1. P. 154 -179 | 78 | 90% | 16 |
| 3 | TRAPNELL, C , ROBERTS, A , GOFF, L , PERTEA, G , KIM, D , KELLEY, DR , PIMENTEL, H , SALZBERG, SL , RINN, JL , PACHTER, L , (2012) DIFFERENTIAL GENE AND TRANSCRIPT EXPRESSION ANALYSIS OF RNA-SEQ EXPERIMENTS WITH TOPHAT AND CUFFLINKS.NATURE PROTOCOLS. VOL. 7. ISSUE 3. P. 562 -578 | 34 | 83% | 2089 |
| 4 | GAWAD, C , KOH, W , QUAKE, SR , (2016) SINGLE-CELL GENOME SEQUENCING: CURRENT STATE OF THE SCIENCE.NATURE REVIEWS GENETICS. VOL. 17. ISSUE 3. P. 175 -188 | 63 | 59% | 49 |
| 5 | METZKER, ML , (2010) APPLICATIONS OF NEXT-GENERATION SEQUENCING SEQUENCING TECHNOLOGIES - THE NEXT GENERATION.NATURE REVIEWS GENETICS. VOL. 11. ISSUE 1. P. 31 -46 | 54 | 59% | 2273 |
| 6 | ALKAN, C , COE, BP , EICHLER, EE , (2011) APPLICATIONS OF NEXT-GENERATION SEQUENCING GENOME STRUCTURAL VARIATION DISCOVERY AND GENOTYPING.NATURE REVIEWS GENETICS. VOL. 12. ISSUE 5. P. 363 -375 | 78 | 74% | 371 |
| 7 | BACHER, R , KENDZIORSKI, C , (2016) DESIGN AND COMPUTATIONAL ANALYSIS OF SINGLE-CELL RNA-SEQUENCING EXPERIMENTS.GENOME BIOLOGY. VOL. 17. ISSUE . P. - | 77 | 92% | 6 |
| 8 | DING, L , WENDL, MC , MCMICHAEL, JF , RAPHAEL, BJ , (2014) EXPANDING THE COMPUTATIONAL TOOLBOX FOR MINING CANCER GENOMES.NATURE REVIEWS GENETICS. VOL. 15. ISSUE 8. P. 556 -570 | 102 | 66% | 54 |
| 9 | HAAS, BJ , PAPANICOLAOU, A , YASSOUR, M , GRABHERR, M , BLOOD, PD , BOWDEN, J , COUGER, MB , ECCLES, D , LI, B , LIEBER, M , ET AL (2013) DE NOVO TRANSCRIPT SEQUENCE RECONSTRUCTION FROM RNA-SEQ USING THE TRINITY PLATFORM FOR REFERENCE GENERATION AND ANALYSIS.NATURE PROTOCOLS. VOL. 8. ISSUE 8. P. 1494 -1512 | 36 | 84% | 734 |
| 10 | LIU, GE , BICKHART, DM , (2012) COPY NUMBER VARIATION IN THE CATTLE GENOME.FUNCTIONAL & INTEGRATIVE GENOMICS. VOL. 12. ISSUE 4. P. 609 -624 | 117 | 80% | 17 |
Classes with closest relation at Level 2 |