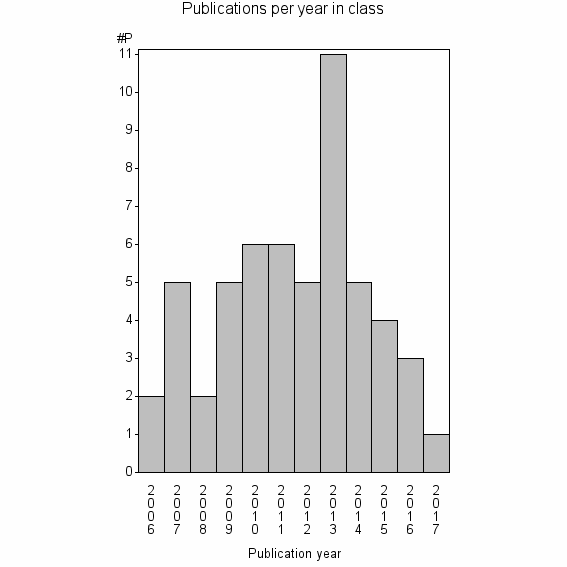
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 37522 | 55 | 32.0 | 85% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 868 | 2 | BMC BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS | 11572 |
| 37522 | 1 | DIFFERENTIAL GENE EXPRESSION DETECTION//CANC GENOME DISCOVERY CCGD//CANCER GENE ACTIVATION HETEROGENEITY | 55 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | DIFFERENTIAL GENE EXPRESSION DETECTION | authKW | 1665585 | 5% | 100% | 3 |
| 2 | CANC GENOME DISCOVERY CCGD | address | 555195 | 2% | 100% | 1 |
| 3 | CANCER GENE ACTIVATION HETEROGENEITY | authKW | 555195 | 2% | 100% | 1 |
| 4 | DEV EXCELLENCE ASIA PACIFICCHUO KU | address | 555195 | 2% | 100% | 1 |
| 5 | DMICE | address | 555195 | 2% | 100% | 1 |
| 6 | EXCRETION AND TOXICITY ADME TOX | authKW | 555195 | 2% | 100% | 1 |
| 7 | GENEGO DATABASE | authKW | 555195 | 2% | 100% | 1 |
| 8 | METADRUG TM | authKW | 555195 | 2% | 100% | 1 |
| 9 | OPEN ENDED CATEGORIES | authKW | 555195 | 2% | 100% | 1 |
| 10 | OUTLIER ROBUST T STATISTIC | authKW | 555195 | 2% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 2388 | 38% | 0% | 21 |
| 2 | Statistics & Probability | 508 | 24% | 0% | 13 |
| 3 | Biochemical Research Methods | 123 | 16% | 0% | 9 |
| 4 | Oncology | 121 | 25% | 0% | 14 |
| 5 | Biotechnology & Applied Microbiology | 112 | 20% | 0% | 11 |
| 6 | Genetics & Heredity | 94 | 18% | 0% | 10 |
| 7 | Multidisciplinary Sciences | 53 | 5% | 0% | 3 |
| 8 | Computer Science, Interdisciplinary Applications | 18 | 5% | 0% | 3 |
| 9 | Biology | 16 | 5% | 0% | 3 |
| 10 | Medical Informatics | 10 | 2% | 0% | 1 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CANC GENOME DISCOVERY CCGD | 555195 | 2% | 100% | 1 |
| 2 | DEV EXCELLENCE ASIA PACIFICCHUO KU | 555195 | 2% | 100% | 1 |
| 3 | DMICE | 555195 | 2% | 100% | 1 |
| 4 | PHPM | 185064 | 2% | 33% | 1 |
| 5 | GENE VIRAL THER Y | 138795 | 4% | 13% | 2 |
| 6 | CANC STEM CELL INNOVAT CAST | 111037 | 2% | 20% | 1 |
| 7 | UNIT 1374 | 101969 | 5% | 6% | 3 |
| 8 | RONALD A MATRICARIA MOL MED | 92531 | 2% | 17% | 1 |
| 9 | SPLEEN STOMACH DIS | 79312 | 2% | 14% | 1 |
| 10 | BLOOD DNA PROFILING IL | 69398 | 2% | 13% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOSTATISTICS | 11499 | 7% | 1% | 4 |
| 2 | BMC MEDICAL GENOMICS | 3038 | 4% | 0% | 2 |
| 3 | CANCER GENOMICS & PROTEOMICS | 2642 | 2% | 0% | 1 |
| 4 | COMPUTATIONAL BIOLOGY AND CHEMISTRY | 2534 | 4% | 0% | 2 |
| 5 | INTERNATIONAL JOURNAL OF BIOSTATISTICS | 2412 | 2% | 0% | 1 |
| 6 | ONCOGENESIS | 2109 | 2% | 0% | 1 |
| 7 | COMPUTATIONAL AND MATHEMATICAL METHODS IN MEDICINE | 1724 | 4% | 0% | 2 |
| 8 | JOURNAL OF BIOINFORMATICS AND COMPUTATIONAL BIOLOGY | 1515 | 2% | 0% | 1 |
| 9 | BMC SYSTEMS BIOLOGY | 1382 | 4% | 0% | 2 |
| 10 | BIOINFORMATICS | 1300 | 9% | 0% | 5 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DIFFERENTIAL GENE EXPRESSION DETECTION | 1665585 | 5% | 100% | 3 | Search DIFFERENTIAL+GENE+EXPRESSION+DETECTION | Search DIFFERENTIAL+GENE+EXPRESSION+DETECTION |
| 2 | CANCER GENE ACTIVATION HETEROGENEITY | 555195 | 2% | 100% | 1 | Search CANCER+GENE+ACTIVATION+HETEROGENEITY | Search CANCER+GENE+ACTIVATION+HETEROGENEITY |
| 3 | EXCRETION AND TOXICITY ADME TOX | 555195 | 2% | 100% | 1 | Search EXCRETION+AND+TOXICITY+ADME+TOX | Search EXCRETION+AND+TOXICITY+ADME+TOX |
| 4 | GENEGO DATABASE | 555195 | 2% | 100% | 1 | Search GENEGO+DATABASE | Search GENEGO+DATABASE |
| 5 | METADRUG TM | 555195 | 2% | 100% | 1 | Search METADRUG+TM | Search METADRUG+TM |
| 6 | OPEN ENDED CATEGORIES | 555195 | 2% | 100% | 1 | Search OPEN+ENDED+CATEGORIES | Search OPEN+ENDED+CATEGORIES |
| 7 | OUTLIER ROBUST T STATISTIC | 555195 | 2% | 100% | 1 | Search OUTLIER+ROBUST+T+STATISTIC | Search OUTLIER+ROBUST+T+STATISTIC |
| 8 | STATISTIC METHODOLOGY | 555195 | 2% | 100% | 1 | Search STATISTIC+METHODOLOGY | Search STATISTIC+METHODOLOGY |
| 9 | SUBPATH | 555195 | 2% | 100% | 1 | Search SUBPATH | Search SUBPATH |
| 10 | TRI MEAN | 555195 | 2% | 100% | 1 | Search TRI+MEAN | Search TRI+MEAN |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | EMERSON, SC , EMERSON, SS , (2013) DETECTING DIFFERENTIAL GENE EXPRESSION IN SUBGROUPS OF A DISEASE POPULATION.INTERNATIONAL JOURNAL OF BIOSTATISTICS. VOL. 9. ISSUE 1. P. 95-108 | 4 | 80% | 0 |
| 2 | BOTTOMLY, D , RYABININ, PA , TYNER, JW , CHANG, BH , LORIAUX, MM , DRUKER, BJ , MCWEENEY, SK , WILMOT, B , (2013) COMPARISON OF METHODS TO IDENTIFY ABERRANT EXPRESSION PATTERNS IN INDIVIDUAL PATIENTS: AUGMENTING OUR TOOLKIT FOR PRECISION MEDICINE.GENOME MEDICINE. VOL. 5. ISSUE . P. - | 12 | 25% | 1 |
| 3 | JI, ZH , WU, CG , WANG, Y , GUAN, RC , TU, HW , WU, XZ , LIANG, YC , (2012) TRI-MEAN-BASED STATISTICAL DIFFERENTIAL GENE EXPRESSION DETECTION.INTERNATIONAL JOURNAL OF DATA MINING AND BIOINFORMATICS. VOL. 6. ISSUE 3. P. 255-271 | 6 | 38% | 0 |
| 4 | WANG, Y , WU, CG , JI, ZH , WANG, BH , LIANG, YC , (2011) NON-PARAMETRIC CHANGE-POINT METHOD FOR DIFFERENTIAL GENE EXPRESSION DETECTION.PLOS ONE. VOL. 6. ISSUE 5. P. - | 6 | 33% | 16 |
| 5 | HSU, CL , LEE, WC , (2010) DETECTING DIFFERENTIALLY EXPRESSED GENES IN HETEROGENEOUS DISEASES USING HALF STUDENT'S T-TEST.INTERNATIONAL JOURNAL OF EPIDEMIOLOGY. VOL. 39. ISSUE 6. P. 1597 -1604 | 5 | 31% | 2 |
| 6 | DE RONDE, JJ , RIGAILL, G , ROTTENBERG, S , RODENHUIS, S , WESSELS, LFA , (2013) IDENTIFYING SUBGROUP MARKERS IN HETEROGENEOUS POPULATIONS.NUCLEIC ACIDS RESEARCH. VOL. 41. ISSUE 21. P. - | 4 | 31% | 5 |
| 7 | RODEN, DL , SEWELL, GW , LOBLEY, A , LEVINE, AP , SMITH, AM , SEGAL, AW , (2014) ZODET: SOFTWARE FOR THE IDENTIFICATION, ANALYSIS AND VISUALISATION OF OUTLIER GENES IN MICROARRAY EXPRESSION DATA.PLOS ONE. VOL. 9. ISSUE 1. P. - | 4 | 27% | 1 |
| 8 | WANG, HW , SUN, Q , ZHAO, WY , QI, LS , GU, YY , LI, PF , ZHANG, MM , LI, Y , LIU, SL , GUO, Z , (2015) INDIVIDUAL-LEVEL ANALYSIS OF DIFFERENTIAL EXPRESSION OF GENES AND PATHWAYS FOR PERSONALIZED MEDICINE.BIOINFORMATICS. VOL. 31. ISSUE 1. P. 62 -68 | 6 | 16% | 4 |
| 9 | MPINDI, JP , SARA, H , HAAPA-PAANANEN, S , KILPINEN, S , PISTO, T , BUCHER, E , OJALA, K , ILJIN, K , VAINIO, P , BJORKMAN, M , ET AL (2011) GTI: A NOVEL ALGORITHM FOR IDENTIFYING OUTLIER GENE EXPRESSION PROFILES FROM INTEGRATED MICROARRAY DATASETS.PLOS ONE. VOL. 6. ISSUE 2. P. - | 5 | 20% | 12 |
| 10 | GLEISS, A , SANCHEZ-CABO, F , PERCO, P , TONG, D , HEINZE, G , (2011) ADAPTIVE TRIMMED T-STATISTICS FOR IDENTIFYING PREDOMINANTLY HIGH EXPRESSION IN A MICROARRAY EXPERIMENT.STATISTICS IN MEDICINE. VOL. 30. ISSUE 1. P. 52-61 | 4 | 25% | 4 |
Classes with closest relation at Level 1 |