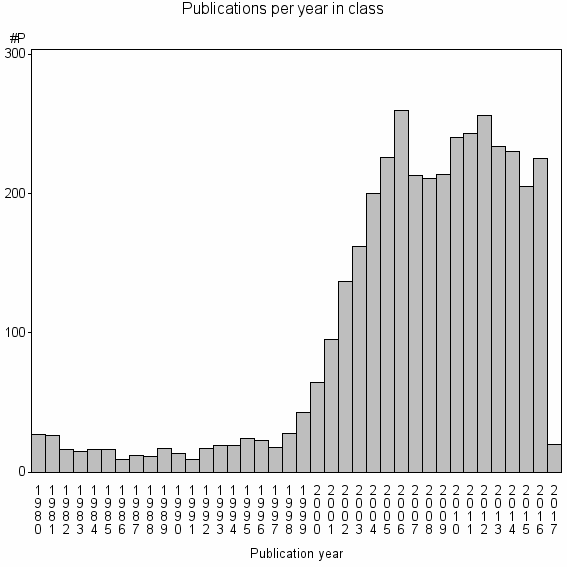
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 2525 | 3813 | 41.9 | 83% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 478 | 3 | PHAGE DISPLAY//CELL FREE PROTEIN SYNTHESIS//ANTIBODY ENGINEERING | 23687 |
| 2525 | 2 | PROTEIN MICROARRAY//DIAZIRINE//PROTEIN ARRAY | 3813 |
| 7202 | 1 | PROTEIN MICROARRAY//PROTEIN ARRAY//ANTIBODY MICROARRAY | 1394 |
| 13434 | 1 | CHEMICAL PROTEOMICS//ACTIVITY BASED PROTEIN PROFILING//ACTIVITY BASED PROBES | 837 |
| 16265 | 1 | DIVERSITY ORIENTED SYNTHESIS//SMALL MOLECULE MICROARRAYS//ICCB | 664 |
| 18286 | 1 | DIAZIRINE//PHOTOAFFINITY LABELING//BIORECOGNIT CHEM | 558 |
| 26304 | 1 | SPOT SYNTHESIS//PEPTIDE ARRAY//CHIP BASED PEPTIDE LIB | 258 |
| 35110 | 1 | OBLIQUE INCIDENCE REFLECTIVITY DIFFERENCE OIRD//OBLIQUE INCIDENCE REFLECTIVITY DIFFERENCE//SERIES PHOTODETECTOR FREQUENCY CIRCUIT SYSTEM | 102 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | PROTEIN MICROARRAY | authKW | 532329 | 5% | 38% | 177 |
| 2 | DIAZIRINE | authKW | 246270 | 2% | 49% | 63 |
| 3 | PROTEIN ARRAY | authKW | 234609 | 3% | 28% | 105 |
| 4 | CHEMICAL PROTEOMICS | authKW | 196093 | 1% | 44% | 56 |
| 5 | ANTIBODY MICROARRAY | authKW | 195507 | 1% | 46% | 53 |
| 6 | ANTIBODY MICROARRAYS | authKW | 177964 | 1% | 65% | 34 |
| 7 | ACTIVITY BASED PROTEIN PROFILING | authKW | 156546 | 1% | 53% | 37 |
| 8 | ACTIVITY BASED PROBES | authKW | 155684 | 1% | 37% | 52 |
| 9 | PHOTOAFFINITY LABELING | authKW | 149602 | 3% | 17% | 107 |
| 10 | PROTEIN CHIP | authKW | 142848 | 1% | 31% | 57 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemical Research Methods | 14750 | 21% | 0% | 809 |
| 2 | Biochemistry & Molecular Biology | 3459 | 28% | 0% | 1063 |
| 3 | Chemistry, Medicinal | 3326 | 9% | 0% | 358 |
| 4 | Chemistry, Organic | 3282 | 15% | 0% | 555 |
| 5 | Chemistry, Multidisciplinary | 2542 | 19% | 0% | 724 |
| 6 | Chemistry, Analytical | 2157 | 11% | 0% | 438 |
| 7 | Biophysics | 618 | 6% | 0% | 217 |
| 8 | Biotechnology & Applied Microbiology | 546 | 7% | 0% | 249 |
| 9 | Chemistry, Applied | 122 | 2% | 0% | 95 |
| 10 | Pharmacology & Pharmacy | 112 | 6% | 0% | 228 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | PLANT CHEMET | 76832 | 1% | 40% | 24 |
| 2 | CHIP BASED PEPTIDE LIB | 64051 | 0% | 100% | 8 |
| 3 | CREATE HLTH | 59518 | 1% | 28% | 27 |
| 4 | IMMUNOTECHNOL | 44832 | 1% | 12% | 45 |
| 5 | ICCB | 44471 | 0% | 56% | 10 |
| 6 | BIORECOGNIT CHEM | 43587 | 0% | 78% | 7 |
| 7 | NUS MEDCHEM PROGRAM | 42825 | 0% | 31% | 17 |
| 8 | HARVARD CHEM CELL BIOL | 36382 | 0% | 45% | 10 |
| 9 | ROBOT CHEM GRP | 32025 | 0% | 100% | 4 |
| 10 | PL PROTE MOL MED | 30846 | 1% | 15% | 25 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CURRENT OPINION IN CHEMICAL BIOLOGY | 20766 | 2% | 4% | 70 |
| 2 | PROTEOMICS | 19328 | 3% | 2% | 118 |
| 3 | MOLECULAR & CELLULAR PROTEOMICS | 11557 | 2% | 2% | 67 |
| 4 | EXPERT REVIEW OF PROTEOMICS | 11139 | 1% | 4% | 32 |
| 5 | CHEMISTRY & BIOLOGY | 10723 | 2% | 2% | 60 |
| 6 | NATURE CHEMICAL BIOLOGY | 7935 | 1% | 3% | 37 |
| 7 | CHEMBIOCHEM | 6384 | 2% | 1% | 60 |
| 8 | PROTEOMICS CLINICAL APPLICATIONS | 6292 | 1% | 3% | 27 |
| 9 | JOURNAL OF PROTEOME RESEARCH | 6139 | 2% | 1% | 69 |
| 10 | JOURNAL OF COMBINATORIAL CHEMISTRY | 5224 | 1% | 2% | 30 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PROTEIN MICROARRAY | 532329 | 5% | 38% | 177 | Search PROTEIN+MICROARRAY | Search PROTEIN+MICROARRAY |
| 2 | DIAZIRINE | 246270 | 2% | 49% | 63 | Search DIAZIRINE | Search DIAZIRINE |
| 3 | PROTEIN ARRAY | 234609 | 3% | 28% | 105 | Search PROTEIN+ARRAY | Search PROTEIN+ARRAY |
| 4 | CHEMICAL PROTEOMICS | 196093 | 1% | 44% | 56 | Search CHEMICAL+PROTEOMICS | Search CHEMICAL+PROTEOMICS |
| 5 | ANTIBODY MICROARRAY | 195507 | 1% | 46% | 53 | Search ANTIBODY+MICROARRAY | Search ANTIBODY+MICROARRAY |
| 6 | ANTIBODY MICROARRAYS | 177964 | 1% | 65% | 34 | Search ANTIBODY+MICROARRAYS | Search ANTIBODY+MICROARRAYS |
| 7 | ACTIVITY BASED PROTEIN PROFILING | 156546 | 1% | 53% | 37 | Search ACTIVITY+BASED+PROTEIN+PROFILING | Search ACTIVITY+BASED+PROTEIN+PROFILING |
| 8 | ACTIVITY BASED PROBES | 155684 | 1% | 37% | 52 | Search ACTIVITY+BASED+PROBES | Search ACTIVITY+BASED+PROBES |
| 9 | PHOTOAFFINITY LABELING | 149602 | 3% | 17% | 107 | Search PHOTOAFFINITY+LABELING | Search PHOTOAFFINITY+LABELING |
| 10 | PROTEIN CHIP | 142848 | 1% | 31% | 57 | Search PROTEIN+CHIP | Search PROTEIN+CHIP |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | NIPHAKIS, MJ , CRAVATT, BF , (2014) ENZYME INHIBITOR DISCOVERY BY ACTIVITY-BASED PROTEIN PROFILING.ANNUAL REVIEW OF BIOCHEMISTRY, VOL 83. VOL. 83. ISSUE . P. 341 -377 | 88 | 38% | 36 |
| 2 | SERIM, S , HAEDKE, U , VERHELST, SHL , (2012) ACTIVITY-BASED PROBES FOR THE STUDY OF PROTEASES: RECENT ADVANCES AND DEVELOPMENTS.CHEMMEDCHEM. VOL. 7. ISSUE 7. P. 1146-1159 | 68 | 56% | 41 |
| 3 | BORREBAECK, CAK , WINGREN, C , (2007) HIGH-THROUGHPUT PROTEOMICS USING ANTIBODY MICROARRAYS: AN UPDATE.EXPERT REVIEW OF MOLECULAR DIAGNOSTICS. VOL. 7. ISSUE 5. P. 673-686 | 76 | 59% | 57 |
| 4 | WANG, K , YANG, T , WU, Q , ZHAO, X , NICE, EC , HUANG, CH , (2012) CHEMISTRY-BASED FUNCTIONAL PROTEOMICS FOR DRUG TARGET DECONVOLUTION.EXPERT REVIEW OF PROTEOMICS. VOL. 9. ISSUE 3. P. 293 -310 | 77 | 55% | 9 |
| 5 | CRETICH, M , DAMIN, F , PIRRI, G , CHIARI, M , (2006) PROTEIN AND PEPTIDE ARRAYS: RECENT TRENDS AND NEW DIRECTIONS.BIOMOLECULAR ENGINEERING. VOL. 23. ISSUE 2-3. P. 77-88 | 60 | 66% | 155 |
| 6 | ZHENG, WL , LI, G , LI, XY , (2015) AFFINITY PURIFICATION IN TARGET IDENTIFICATION: THE SPECIFICITY CHALLENGE.ARCHIVES OF PHARMACAL RESEARCH. VOL. 38. ISSUE 9. P. 1661 -1685 | 90 | 40% | 4 |
| 7 | WRIGHT, MH , SIEBER, SA , (2016) CHEMICAL PROTEOMICS APPROACHES FOR IDENTIFYING THE CELLULAR TARGETS OF NATURAL PRODUCTS.NATURAL PRODUCT REPORTS. VOL. 33. ISSUE 5. P. 681 -708 | 53 | 44% | 9 |
| 8 | HATANAKA, Y , (2015) DEVELOPMENT AND LEADING-EDGE APPLICATION OF INNOVATIVE PHOTOAFFINITY LABELING.CHEMICAL & PHARMACEUTICAL BULLETIN. VOL. 63. ISSUE 1. P. 1 -12 | 52 | 68% | 3 |
| 9 | YANG, PY , LIU, K , (2015) ACTIVITY-BASED PROTEIN PROFILING: RECENT ADVANCES IN PROBE DEVELOPMENT AND APPLICATIONS.CHEMBIOCHEM. VOL. 16. ISSUE 5. P. 712 -724 | 56 | 54% | 9 |
| 10 | UTTAMCHANDANI, M , LI, JQ , SUN, H , YAO, SQ , (2008) ACTIVITY-BASED PROTEIN PROFILING: NEW DEVELOPMENTS AND DIRECTIONS IN FUNCTIONAL PROTEOMICS.CHEMBIOCHEM. VOL. 9. ISSUE 5. P. 667-675 | 53 | 70% | 41 |
Classes with closest relation at Level 2 |