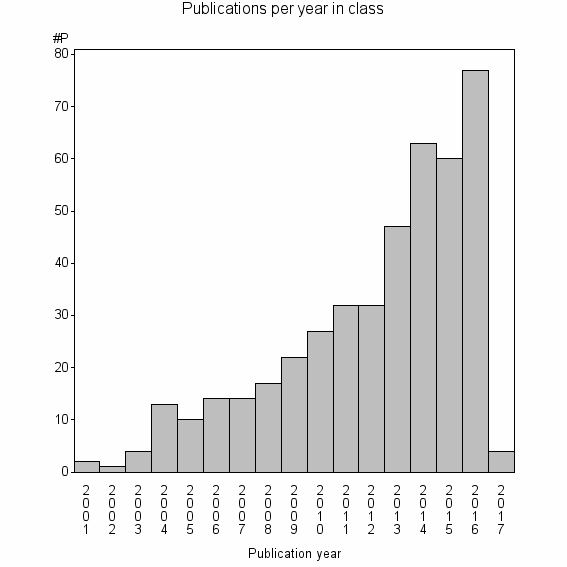
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 20966 | 439 | 44.9 | 85% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 868 | 2 | BMC BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS | 11572 |
| 20966 | 1 | DYNAMICAL NETWORK BIOMARKER//EDGE BIOMARKER//ABORAT INNOVAT MATH MODELLING | 439 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | DYNAMICAL NETWORK BIOMARKER | authKW | 486890 | 2% | 100% | 7 |
| 2 | EDGE BIOMARKER | authKW | 417334 | 1% | 100% | 6 |
| 3 | ABORAT INNOVAT MATH MODELLING | address | 335945 | 4% | 30% | 16 |
| 4 | SIBS NOVO NORDISK TRANSLAT PREDIABET | address | 291882 | 4% | 26% | 16 |
| 5 | NETWORK BIOMARKER | authKW | 227208 | 2% | 47% | 7 |
| 6 | WEIGHTED GENE CO EXPRESSION NETWORK ANALYSIS | authKW | 227208 | 2% | 47% | 7 |
| 7 | DIFFERENTIAL NETWORKING | authKW | 139111 | 0% | 100% | 2 |
| 8 | MODULE BIOMARKERS | authKW | 139111 | 0% | 100% | 2 |
| 9 | MULTIPLE DIFFERENTIAL MODULES | authKW | 139111 | 0% | 100% | 2 |
| 10 | PSYCHIAT CLIN MORPHOL | address | 139111 | 0% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 13833 | 33% | 0% | 143 |
| 2 | Biochemical Research Methods | 2885 | 27% | 0% | 120 |
| 3 | Biotechnology & Applied Microbiology | 2568 | 33% | 0% | 144 |
| 4 | Genetics & Heredity | 956 | 20% | 0% | 89 |
| 5 | Multidisciplinary Sciences | 320 | 5% | 0% | 21 |
| 6 | Biochemistry & Molecular Biology | 107 | 17% | 0% | 75 |
| 7 | Medicine, Research & Experimental | 79 | 6% | 0% | 27 |
| 8 | Statistics & Probability | 72 | 4% | 0% | 16 |
| 9 | Oncology | 66 | 8% | 0% | 37 |
| 10 | Cell Biology | 27 | 6% | 0% | 27 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ABORAT INNOVAT MATH MODELLING | 335945 | 4% | 30% | 16 |
| 2 | SIBS NOVO NORDISK TRANSLAT PREDIABET | 291882 | 4% | 26% | 16 |
| 3 | PSYCHIAT CLIN MORPHOL | 139111 | 0% | 100% | 2 |
| 4 | SYST BIOLINNOVAT CELL SIGNALING NETW | 139103 | 1% | 33% | 6 |
| 5 | BIOSTAT PUBL HLTH PREVENTAT MED | 133755 | 1% | 38% | 5 |
| 6 | TCM COMPLEX SYST | 123650 | 1% | 44% | 4 |
| 7 | HUMAN CENTRIFUGE MED TRAINING | 92740 | 0% | 67% | 2 |
| 8 | AFFILIATED DA YI HOSP | 69556 | 0% | 100% | 1 |
| 9 | ASBESTOS RELATED DISEASES | 69556 | 0% | 100% | 1 |
| 10 | BARIC | 69556 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BMC SYSTEMS BIOLOGY | 22923 | 5% | 1% | 23 |
| 2 | BIOINFORMATICS | 11497 | 10% | 0% | 42 |
| 3 | BMC MEDICAL GENOMICS | 9508 | 2% | 1% | 10 |
| 4 | BMC BIOINFORMATICS | 9464 | 7% | 0% | 32 |
| 5 | BMC GENOMICS | 3493 | 5% | 0% | 22 |
| 6 | JOURNAL OF MOLECULAR CELL BIOLOGY | 2356 | 1% | 1% | 3 |
| 7 | PLOS COMPUTATIONAL BIOLOGY | 1803 | 3% | 0% | 11 |
| 8 | ADVANCES IN ANATOMY EMBRYOLOGY AND CELL BIOLOGY | 1662 | 0% | 1% | 2 |
| 9 | IEEE-ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS | 977 | 1% | 0% | 4 |
| 10 | MOLECULAR SYSTEMS BIOLOGY | 822 | 1% | 0% | 3 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | KAYANO, M , SHIGA, M , MAMITSUKA, H , (2014) DETECTING DIFFERENTIALLY COEXPRESSED GENES FROM LABELED EXPRESSION DATA: A BRIEF REVIEW.IEEE-ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS. VOL. 11. ISSUE 1. P. 154-167 | 18 | 64% | 4 |
| 2 | WANG, DL , WANG, JX , JIANG, YX , LIANG, YC , XU, D , (2017) BFDCA: A COMPREHENSIVE TOOL OF USING BAYES FACTOR FOR DIFFERENTIAL CO-EXPRESSION ANALYSIS.JOURNAL OF MOLECULAR BIOLOGY. VOL. 429. ISSUE 3. P. 446 -453 | 13 | 45% | 1 |
| 3 | LANGFELDER, P , MISCHEL, PS , HORVATH, S , (2013) WHEN IS HUB GENE SELECTION BETTER THAN STANDARD META-ANALYSIS?.PLOS ONE. VOL. 8. ISSUE 4. P. - | 25 | 32% | 42 |
| 4 | YU, H , LIU, BH , YE, ZQ , LI, C , LI, YX , LI, YY , (2011) LINK-BASED QUANTITATIVE METHODS TO IDENTIFY DIFFERENTIALLY COEXPRESSED GENES AND GENE PAIRS.BMC BIOINFORMATICS. VOL. 12. ISSUE . P. - | 19 | 48% | 28 |
| 5 | YANG, J , YU, H , LIU, BH , ZHAO, ZM , LIU, L , MA, LX , LI, YX , LI, YY , (2013) DCGL V2.0: AN R PACKAGE FOR UNVEILING DIFFERENTIAL REGULATION FROM DIFFERENTIAL CO-EXPRESSION.PLOS ONE. VOL. 8. ISSUE 11. P. - | 16 | 55% | 12 |
| 6 | LANGFELDER, P , LUO, R , OLDHAM, MC , HORVATH, S , (2011) IS MY NETWORK MODULE PRESERVED AND REPRODUCIBLE?.PLOS COMPUTATIONAL BIOLOGY. VOL. 7. ISSUE 1. P. - | 19 | 36% | 114 |
| 7 | FUKUSHIMA, A , (2013) DIFFCORR: AN R PACKAGE TO ANALYZE AND VISUALIZE DIFFERENTIAL CORRELATIONS IN BIOLOGICAL NETWORKS.GENE. VOL. 518. ISSUE 1. P. 209 -214 | 18 | 44% | 14 |
| 8 | LI, YY , LI, JY , LI, YX , (2016) DIFFERENTIAL REGULATORY ANALYSIS BASED ON COEXPRESSION NETWORK IN CANCER RESEARCH.BIOMED RESEARCH INTERNATIONAL. VOL. . ISSUE . P. - | 21 | 36% | 0 |
| 9 | LANGFELDER, P , HORVATH, S , (2008) WGCNA: AN R PACKAGE FOR WEIGHTED CORRELATION NETWORK ANALYSIS.BMC BIOINFORMATICS. VOL. 9. ISSUE . P. - | 9 | 35% | 1064 |
| 10 | CAI, CC , LANGFELDER, P , FULLER, TF , OLDHAM, MC , LUO, R , VAN DEN BERG, LH , OPHOFF, RA , HORVATH, S , (2010) IS HUMAN BLOOD A GOOD SURROGATE FOR BRAIN TISSUE IN TRANSCRIPTIONAL STUDIES?.BMC GENOMICS. VOL. 11. ISSUE . P. - | 15 | 45% | 32 |
Classes with closest relation at Level 1 |