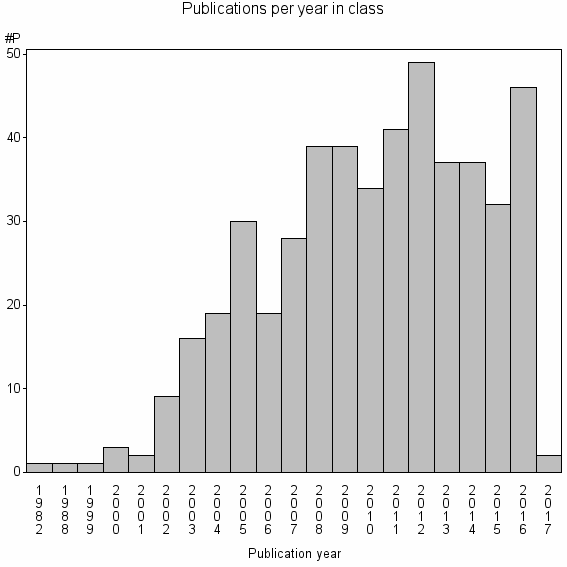
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 19900 | 485 | 45.2 | 85% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 126 | 2 | MOLECULAR BIOLOGY AND EVOLUTION//BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY | 23963 |
| 19900 | 1 | GENE INDEXED MUTANT//NAA PLANT SOIL NUTR UNIT//ORYZABASE | 485 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | GENE INDEXED MUTANT | authKW | 125917 | 0% | 100% | 2 |
| 2 | NAA PLANT SOIL NUTR UNIT | address | 125917 | 0% | 100% | 2 |
| 3 | ORYZABASE | authKW | 125917 | 0% | 100% | 2 |
| 4 | FUNCTIONAL GENE ANNOTATION | authKW | 83943 | 0% | 67% | 2 |
| 5 | NAA PLANT | address | 83943 | 0% | 67% | 2 |
| 6 | GENE CO EXPRESSION NETWORKS | authKW | 70825 | 1% | 38% | 3 |
| 7 | GENE COEXPRESSION | authKW | 65574 | 1% | 21% | 5 |
| 8 | ABERTAY ENVIRON | address | 62958 | 0% | 100% | 1 |
| 9 | AGRICULTURAL OMICS | authKW | 62958 | 0% | 100% | 1 |
| 10 | APPLEGFDB | authKW | 62958 | 0% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Plant Sciences | 4700 | 43% | 0% | 210 |
| 2 | Mathematical & Computational Biology | 3276 | 15% | 0% | 74 |
| 3 | Biotechnology & Applied Microbiology | 1344 | 23% | 0% | 112 |
| 4 | Biochemical Research Methods | 591 | 12% | 0% | 60 |
| 5 | Biochemistry & Molecular Biology | 550 | 31% | 0% | 148 |
| 6 | Genetics & Heredity | 531 | 15% | 0% | 72 |
| 7 | Cell Biology | 67 | 8% | 0% | 39 |
| 8 | Agronomy | 29 | 3% | 0% | 13 |
| 9 | Horticulture | 26 | 1% | 0% | 7 |
| 10 | Multidisciplinary Sciences | 16 | 1% | 0% | 6 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | NAA PLANT SOIL NUTR UNIT | 125917 | 0% | 100% | 2 |
| 2 | NAA PLANT | 83943 | 0% | 67% | 2 |
| 3 | ABERTAY ENVIRON | 62958 | 0% | 100% | 1 |
| 4 | BAKER COMPUTAT BIOL ELECT COMP ENGN | 62958 | 0% | 100% | 1 |
| 5 | CBFP | 62958 | 0% | 100% | 1 |
| 6 | CENT GENET BRONNEN NEDERLAND | 62958 | 0% | 100% | 1 |
| 7 | CNRS 1165 | 62958 | 0% | 100% | 1 |
| 8 | DEEP SEQ LIFE SCI | 62958 | 0% | 100% | 1 |
| 9 | DEP ENERGY GREAT LAKES BIOENERGY | 62958 | 0% | 100% | 1 |
| 10 | EXCELLENCE INTERACT LEGUME BIOINFORMAT | 62958 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | DATABASE-THE JOURNAL OF BIOLOGICAL DATABASES AND CURATION | 16319 | 3% | 2% | 13 |
| 2 | RICE | 14024 | 2% | 3% | 8 |
| 3 | COMPARATIVE AND FUNCTIONAL GENOMICS | 10477 | 2% | 2% | 8 |
| 4 | NUCLEIC ACIDS RESEARCH | 10199 | 16% | 0% | 80 |
| 5 | PLANT PHYSIOLOGY | 6581 | 10% | 0% | 47 |
| 6 | PLANT AND CELL PHYSIOLOGY | 6134 | 5% | 0% | 26 |
| 7 | PLANT BIOTECHNOLOGY | 4530 | 1% | 1% | 7 |
| 8 | BMC PLANT BIOLOGY | 3876 | 2% | 1% | 12 |
| 9 | BMC BIOINFORMATICS | 3329 | 4% | 0% | 20 |
| 10 | BMC GENOMICS | 2875 | 4% | 0% | 21 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | GENE INDEXED MUTANT | 125917 | 0% | 100% | 2 | Search GENE+INDEXED+MUTANT | Search GENE+INDEXED+MUTANT |
| 2 | ORYZABASE | 125917 | 0% | 100% | 2 | Search ORYZABASE | Search ORYZABASE |
| 3 | FUNCTIONAL GENE ANNOTATION | 83943 | 0% | 67% | 2 | Search FUNCTIONAL+GENE+ANNOTATION | Search FUNCTIONAL+GENE+ANNOTATION |
| 4 | GENE CO EXPRESSION NETWORKS | 70825 | 1% | 38% | 3 | Search GENE+CO+EXPRESSION+NETWORKS | Search GENE+CO+EXPRESSION+NETWORKS |
| 5 | GENE COEXPRESSION | 65574 | 1% | 21% | 5 | Search GENE+COEXPRESSION | Search GENE+COEXPRESSION |
| 6 | AGRICULTURAL OMICS | 62958 | 0% | 100% | 1 | Search AGRICULTURAL+OMICS | Search AGRICULTURAL+OMICS |
| 7 | APPLEGFDB | 62958 | 0% | 100% | 1 | Search APPLEGFDB | Search APPLEGFDB |
| 8 | APPLICATIONS OF INTEGRATION | 62958 | 0% | 100% | 1 | Search APPLICATIONS+OF+INTEGRATION | Search APPLICATIONS+OF+INTEGRATION |
| 9 | ARABIDOPOSIS | 62958 | 0% | 100% | 1 | Search ARABIDOPOSIS | Search ARABIDOPOSIS |
| 10 | ARABIDOPSIS HORMONE RELATED GENE | 62958 | 0% | 100% | 1 | Search ARABIDOPSIS+HORMONE+RELATED+GENE | Search ARABIDOPSIS+HORMONE+RELATED+GENE |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | SERIN, EAR , NIJVEEN, H , HILHORST, HWM , LIGTERINK, W , (2016) LEARNING FROM CO-EXPRESSION NETWORKS: POSSIBILITIES AND CHALLENGES.FRONTIERS IN PLANT SCIENCE. VOL. 7. ISSUE . P. - | 43 | 29% | 5 |
| 2 | OBAYASHI, T , OKAMURA, Y , ITO, S , TADAKA, S , AOKI, Y , SHIROTA, M , KINOSHITA, K , (2014) ATTED-II IN 2014: EVALUATION OF GENE COEXPRESSION IN AGRICULTURALLY IMPORTANT PLANTS.PLANT AND CELL PHYSIOLOGY. VOL. 55. ISSUE 1. P. - | 22 | 56% | 40 |
| 3 | TOHGE, T , FERNIE, AR , (2012) CO-EXPRESSION AND CO-RESPONSES: WITHIN AND BEYOND TRANSCRIPTION.FRONTIERS IN PLANT SCIENCE. VOL. 3. ISSUE . P. - | 33 | 46% | 7 |
| 4 | SCHAEFER, RJ , MICHNO, JM , MYERS, CL , (2017) UNRAVELING GENE FUNCTION IN AGRICULTURAL SPECIES USING GENE CO-EXPRESSION NETWORKS.BIOCHIMICA ET BIOPHYSICA ACTA-GENE REGULATORY MECHANISMS. VOL. 1860. ISSUE 1. P. 53 -63 | 26 | 30% | 1 |
| 5 | TZFADIA, O , DIELS, T , DE MEYER, S , VANDEPOELE, K , AHARONI, A , VAN DE PEER, Y , (2016) COEXPNETVIZ: COMPARATIVE CO-EXPRESSION NETWORKS CONSTRUCTION AND VISUALIZATION TOOL.FRONTIERS IN PLANT SCIENCE. VOL. 6. ISSUE . P. - | 16 | 62% | 1 |
| 6 | DE BODT, S , HOLLUNDER, J , NELISSEN, H , MEULEMEESTER, N , INZE, D , (2012) CORNET 2.0: INTEGRATING PLANT COEXPRESSION, PROTEIN-PROTEIN INTERACTIONS, REGULATORY INTERACTIONS, GENE ASSOCIATIONS AND FUNCTIONAL ANNOTATIONS.NEW PHYTOLOGIST. VOL. 195. ISSUE 3. P. 707-720 | 25 | 40% | 31 |
| 7 | FICKLIN, SP , FELTUS, FA , (2011) GENE COEXPRESSION NETWORK ALIGNMENT AND CONSERVATION OF GENE MODULES BETWEEN TWO GRASS SPECIES: MAIZE AND RICE.PLANT PHYSIOLOGY. VOL. 156. ISSUE 3. P. 1244 -1256 | 22 | 43% | 55 |
| 8 | OBAYASHI, T , KINOSHITA, K , (2010) COEXPRESSION LANDSCAPE IN ATTED-II: USAGE OF GENE LIST AND GENE NETWORK FOR VARIOUS TYPES OF PATHWAYS.JOURNAL OF PLANT RESEARCH. VOL. 123. ISSUE 3. P. 311-319 | 21 | 50% | 35 |
| 9 | OBAYASHI, T , NISHIDA, K , KASAHARA, K , KINOSHITA, K , (2011) ATTED-II UPDATES: CONDITION-SPECIFIC GENE COEXPRESSION TO EXTEND COEXPRESSION ANALYSES AND APPLICATIONS TO A BROAD RANGE OF FLOWERING PLANTS.PLANT AND CELL PHYSIOLOGY. VOL. 52. ISSUE 2. P. 213 -219 | 14 | 61% | 89 |
| 10 | CAO, PJ , JUNG, KH , CHOI, D , HWANG, D , ZHU, J , RONALD, PC , (2012) THE RICE OLIGONUCLEOTIDE ARRAY DATABASE: AN ATLAS OF RICE GENE EXPRESSION.RICE. VOL. 5. ISSUE . P. - | 19 | 46% | 39 |
Classes with closest relation at Level 1 |