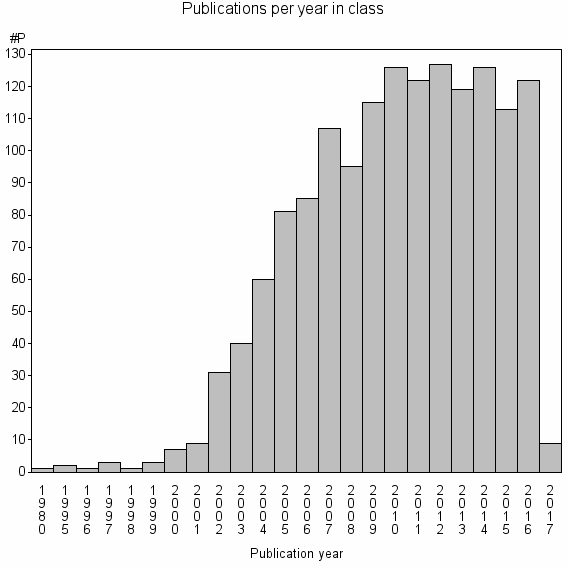
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 6275 | 1505 | 40.6 | 72% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 548 | 2 | BMC SYSTEMS BIOLOGY//MATHEMATICAL & COMPUTATIONAL BIOLOGY//BOOLEAN NETWORKS | 14655 |
| 6275 | 1 | GENE REGULATORY NETWORK//NETWORK INFERENCE//REVERSE ENGINEERING | 1505 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | GENE REGULATORY NETWORK | authKW | 612258 | 11% | 19% | 163 |
| 2 | NETWORK INFERENCE | authKW | 332799 | 3% | 37% | 44 |
| 3 | REVERSE ENGINEERING | authKW | 127950 | 6% | 8% | 83 |
| 4 | DREAM CHALLENGE | authKW | 121726 | 0% | 100% | 6 |
| 5 | MATHEMATICAL & COMPUTATIONAL BIOLOGY | WoSSC | 117554 | 51% | 1% | 769 |
| 6 | GENE NETWORKS | authKW | 115944 | 4% | 10% | 57 |
| 7 | GENE NETWORK INFERENCE | authKW | 91291 | 0% | 75% | 6 |
| 8 | BMC SYSTEMS BIOLOGY | journal | 87084 | 6% | 5% | 83 |
| 9 | GENE REGULATORY NETWORK INFERENCE | authKW | 81150 | 0% | 100% | 4 |
| 10 | BIOINFORMATICS | journal | 80854 | 14% | 2% | 206 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 117554 | 51% | 1% | 769 |
| 2 | Biochemical Research Methods | 21942 | 40% | 0% | 605 |
| 3 | Biotechnology & Applied Microbiology | 13811 | 41% | 0% | 612 |
| 4 | Statistics & Probability | 2144 | 10% | 0% | 145 |
| 5 | Computer Science, Interdisciplinary Applications | 1448 | 9% | 0% | 134 |
| 6 | Genetics & Heredity | 766 | 11% | 0% | 160 |
| 7 | Computer Science, Artificial Intelligence | 569 | 6% | 0% | 85 |
| 8 | Biology | 418 | 5% | 0% | 81 |
| 9 | Biochemistry & Molecular Biology | 271 | 15% | 0% | 233 |
| 10 | Mathematics, Interdisciplinary Applications | 262 | 3% | 0% | 52 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | COMPUTAT BIOL MACHINE LEARNING | 76173 | 1% | 29% | 13 |
| 2 | SYST BIOL BIOCHEM MOL BIOPHYS | 60863 | 0% | 100% | 3 |
| 3 | BIOENVIRONM INTEGRAT BIOSCI | 40575 | 0% | 100% | 2 |
| 4 | JBI | 36515 | 0% | 60% | 3 |
| 5 | KNOWLEDGE DISCOVERY BIOINFORMAT GRP | 30428 | 0% | 50% | 3 |
| 6 | JOINT SYST BIOL | 29496 | 1% | 18% | 8 |
| 7 | COMPUTAT ASTROBIOL FUNDAMENTAL BIOL | 27049 | 0% | 67% | 2 |
| 8 | INFECT DIS MOL MEDMED | 27049 | 0% | 67% | 2 |
| 9 | INFORMAT DECIS ALGORITHMS S | 27049 | 0% | 67% | 2 |
| 10 | BONN AACHEN INT IT | 26075 | 0% | 21% | 6 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BMC SYSTEMS BIOLOGY | 87084 | 6% | 5% | 83 |
| 2 | BIOINFORMATICS | 80854 | 14% | 2% | 206 |
| 3 | BMC BIOINFORMATICS | 52159 | 9% | 2% | 139 |
| 4 | IET SYSTEMS BIOLOGY | 38146 | 2% | 8% | 25 |
| 5 | IEEE-ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS | 36270 | 3% | 4% | 45 |
| 6 | JOURNAL OF COMPUTATIONAL BIOLOGY | 18741 | 3% | 2% | 38 |
| 7 | STATISTICAL APPLICATIONS IN GENETICS AND MOLECULAR BIOLOGY | 12982 | 1% | 4% | 18 |
| 8 | INTERNATIONAL JOURNAL OF DATA MINING AND BIOINFORMATICS | 4688 | 1% | 2% | 10 |
| 9 | JOURNAL OF BIOINFORMATICS AND COMPUTATIONAL BIOLOGY | 4472 | 1% | 2% | 9 |
| 10 | IEE PROCEEDINGS SYSTEMS BIOLOGY | 3903 | 0% | 5% | 4 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | GENE REGULATORY NETWORK | 612258 | 11% | 19% | 163 | Search GENE+REGULATORY+NETWORK | Search GENE+REGULATORY+NETWORK |
| 2 | NETWORK INFERENCE | 332799 | 3% | 37% | 44 | Search NETWORK+INFERENCE | Search NETWORK+INFERENCE |
| 3 | REVERSE ENGINEERING | 127950 | 6% | 8% | 83 | Search REVERSE+ENGINEERING | Search REVERSE+ENGINEERING |
| 4 | DREAM CHALLENGE | 121726 | 0% | 100% | 6 | Search DREAM+CHALLENGE | Search DREAM+CHALLENGE |
| 5 | GENE NETWORKS | 115944 | 4% | 10% | 57 | Search GENE+NETWORKS | Search GENE+NETWORKS |
| 6 | GENE NETWORK INFERENCE | 91291 | 0% | 75% | 6 | Search GENE+NETWORK+INFERENCE | Search GENE+NETWORK+INFERENCE |
| 7 | GENE REGULATORY NETWORK INFERENCE | 81150 | 0% | 100% | 4 | Search GENE+REGULATORY+NETWORK+INFERENCE | Search GENE+REGULATORY+NETWORK+INFERENCE |
| 8 | MULTIPLE CHANGEPOINT PROCESS | 60863 | 0% | 100% | 3 | Search MULTIPLE+CHANGEPOINT+PROCESS | Search MULTIPLE+CHANGEPOINT+PROCESS |
| 9 | NESTED EFFECTS MODELS | 60863 | 0% | 100% | 3 | Search NESTED+EFFECTS+MODELS | Search NESTED+EFFECTS+MODELS |
| 10 | NETGENERATOR | 60863 | 0% | 100% | 3 | Search NETGENERATOR | Search NETGENERATOR |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | LIU, ZP , (2015) REVERSE ENGINEERING OF GENOME-WIDE GENE REGULATORY NETWORKS FROM GENE EXPRESSION DATA.CURRENT GENOMICS. VOL. 16. ISSUE 1. P. 3 -22 | 81 | 49% | 9 |
| 2 | LINDE, J , SCHULZE, S , HENKEL, SG , GUTHKE, R , (2015) DATA-AND KNOWLEDGE-BASED MODELING OF GENE REGULATORY NETWORKS: AN UPDATE.EXCLI JOURNAL. VOL. 14. ISSUE . P. 346 -378 | 79 | 55% | 2 |
| 3 | HECKER, M , LAMBECK, S , TOEPFER, S , VAN SOMEREN, E , GUTHKE, R , (2009) GENE REGULATORY NETWORK INFERENCE: DATA INTEGRATION IN DYNAMIC MODELS-A REVIEW.BIOSYSTEMS. VOL. 96. ISSUE 1. P. 86 -103 | 46 | 61% | 238 |
| 4 | VASHISHTHA, S , BRODERICK, G , CRADDOCK, TJA , FLETCHER, MA , KLIMAS, NG , (2015) INFERRING BROAD REGULATORY BIOLOGY FROM TIME COURSE DATA: HAVE WE REACHED AN UPPER BOUND UNDER CONSTRAINTS TYPICAL OF IN VIVO STUDIES?.PLOS ONE. VOL. 10. ISSUE 5. P. - | 50 | 72% | 1 |
| 5 | LIU, LZ , WU, FX , ZHANG, WJ , (2012) REVERSE ENGINEERING OF GENE REGULATORY NETWORKS FROM BIOLOGICAL DATA.WILEY INTERDISCIPLINARY REVIEWS-DATA MINING AND KNOWLEDGE DISCOVERY. VOL. 2. ISSUE 5. P. 365 -385 | 48 | 75% | 2 |
| 6 | HE, F , BALLING, R , ZENG, AP , (2009) REVERSE ENGINEERING AND VERIFICATION OF GENE NETWORKS: PRINCIPLES, ASSUMPTIONS, AND LIMITATIONS OF PRESENT METHODS AND FUTURE PERSPECTIVES.JOURNAL OF BIOTECHNOLOGY. VOL. 144. ISSUE 3. P. 190 -203 | 57 | 50% | 38 |
| 7 | ZHANG, XJ , LIU, KQ , LIU, ZP , DUVAL, B , RICHER, JM , ZHAO, XM , HAO, JK , CHEN, LN , (2013) NARROMI: A NOISE AND REDUNDANCY REDUCTION TECHNIQUE IMPROVES ACCURACY OF GENE REGULATORY NETWORK INFERENCE.BIOINFORMATICS. VOL. 29. ISSUE 1. P. 106-113 | 32 | 76% | 29 |
| 8 | LIU, F , ZHANG, SW , GUO, WF , WEI, ZG , CHEN, LN , (2016) INFERENCE OF GENE REGULATORY NETWORK BASED ON LOCAL BAYESIAN NETWORKS.PLOS COMPUTATIONAL BIOLOGY. VOL. 12. ISSUE 8. P. - | 39 | 67% | 0 |
| 9 | SIMA, C , HUA, JP , JUNG, SW , (2009) INFERENCE OF GENE REGULATORY NETWORKS USING TIME-SERIES DATA: A SURVEY.CURRENT GENOMICS. VOL. 10. ISSUE 6. P. 416-429 | 44 | 62% | 37 |
| 10 | STUDHAM, ME , TJARNBERG, A , NORDLING, TEM , NELANDER, S , SONNHAMMER, ELL , (2014) FUNCTIONAL ASSOCIATION NETWORKS AS PRIORS FOR GENE REGULATORY NETWORK INFERENCE.BIOINFORMATICS. VOL. 30. ISSUE 12. P. 130 -138 | 30 | 81% | 5 |
Classes with closest relation at Level 1 |