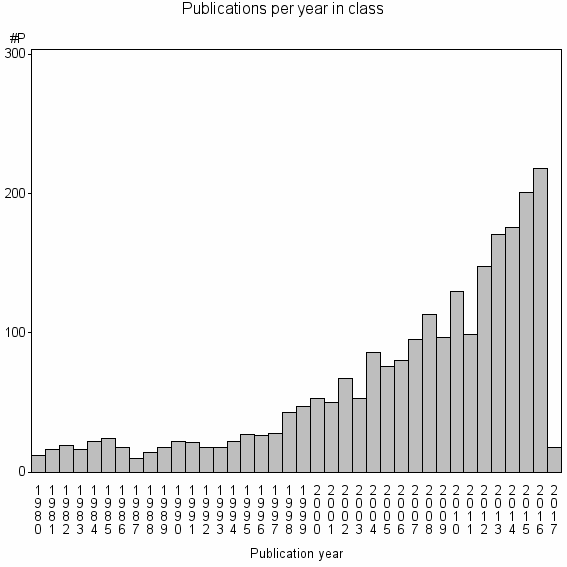
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 3084 | 2372 | 51.7 | 84% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 71 | 3 | GENETICS & HEREDITY//CHROMATIN//CELL BIOLOGY | 83302 |
| 3084 | 2 | CHROMOSOME TERRITORY//NUCLEAR ARCHITECTURE//NUCLEAR ORGANIZATION | 2372 |
| 3065 | 1 | CHROMOSOME TERRITORY//NUCLEAR ARCHITECTURE//NUCLEAR ORGANIZATION | 2075 |
| 27340 | 1 | GENOME THEORY//GENOME CHAOS//GENOM CANC DIAG | 232 |
| 37159 | 1 | GRP ONCOGENET//INTESTINE DIFFUSE POLYPOSIS//INTESTINE MALIGNANT POLYPOSIS | 65 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | CHROMOSOME TERRITORY | authKW | 1030068 | 4% | 78% | 102 |
| 2 | NUCLEAR ARCHITECTURE | authKW | 535114 | 4% | 45% | 93 |
| 3 | NUCLEAR ORGANIZATION | authKW | 390313 | 4% | 33% | 91 |
| 4 | CHROMOSOME CONFORMATION CAPTURE | authKW | 267218 | 1% | 59% | 35 |
| 5 | HI C | authKW | 212203 | 1% | 63% | 26 |
| 6 | 3D FISH | authKW | 208514 | 1% | 90% | 18 |
| 7 | GENOME THEORY | authKW | 168185 | 1% | 93% | 14 |
| 8 | NUCL DYNAM PROGRAMME | address | 165938 | 1% | 68% | 19 |
| 9 | GENE POSITIONING | authKW | 148395 | 1% | 82% | 14 |
| 10 | TRANSCRIPTION FACTORY | authKW | 141888 | 1% | 53% | 21 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Cell Biology | 17508 | 44% | 0% | 1043 |
| 2 | Genetics & Heredity | 15446 | 34% | 0% | 804 |
| 3 | Biochemistry & Molecular Biology | 2998 | 32% | 0% | 757 |
| 4 | Biochemical Research Methods | 856 | 7% | 0% | 171 |
| 5 | Biotechnology & Applied Microbiology | 812 | 9% | 0% | 218 |
| 6 | Mathematical & Computational Biology | 781 | 4% | 0% | 86 |
| 7 | Developmental Biology | 502 | 3% | 0% | 78 |
| 8 | Oncology | 345 | 8% | 0% | 198 |
| 9 | Microscopy | 179 | 1% | 0% | 25 |
| 10 | Biophysics | 136 | 4% | 0% | 92 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | NUCL DYNAM PROGRAMME | 165938 | 1% | 68% | 19 |
| 2 | PROGRAM SYST BIOL | 94540 | 1% | 28% | 26 |
| 3 | NUCL GENOM HLTH | 65154 | 0% | 56% | 9 |
| 4 | GENOME FUNCT GRP | 63066 | 0% | 70% | 7 |
| 5 | MANITOBA CELL BIOL | 61740 | 2% | 11% | 43 |
| 6 | CHROMATIN GENE EXP S | 57901 | 1% | 30% | 15 |
| 7 | GRP REGULAT SPATIALE GENOMES | 57331 | 0% | 64% | 7 |
| 8 | GENOM CANC DIAG | 52553 | 0% | 58% | 7 |
| 9 | HOWARD HUGHES MED PROGRAM SYST BIOL | 51486 | 0% | 100% | 4 |
| 10 | RG DEV DIS | 51486 | 0% | 100% | 4 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | NUCLEUS | 40491 | 1% | 11% | 29 |
| 2 | CHROMOSOME RESEARCH | 35039 | 3% | 4% | 67 |
| 3 | CHROMOSOMA | 27834 | 3% | 3% | 76 |
| 4 | CURRENT OPINION IN GENETICS & DEVELOPMENT | 16388 | 2% | 3% | 50 |
| 5 | NUCLEUS-AUSTIN | 15264 | 1% | 8% | 14 |
| 6 | CURRENT OPINION IN CELL BIOLOGY | 10285 | 2% | 2% | 46 |
| 7 | GENOME BIOLOGY | 9867 | 2% | 2% | 48 |
| 8 | JOURNAL OF CELL SCIENCE | 5897 | 3% | 1% | 78 |
| 9 | EPIGENETICS & CHROMATIN | 5368 | 0% | 4% | 11 |
| 10 | GENOME RESEARCH | 5349 | 1% | 2% | 24 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CHROMOSOME TERRITORY | 1030068 | 4% | 78% | 102 | Search CHROMOSOME+TERRITORY | Search CHROMOSOME+TERRITORY |
| 2 | NUCLEAR ARCHITECTURE | 535114 | 4% | 45% | 93 | Search NUCLEAR+ARCHITECTURE | Search NUCLEAR+ARCHITECTURE |
| 3 | NUCLEAR ORGANIZATION | 390313 | 4% | 33% | 91 | Search NUCLEAR+ORGANIZATION | Search NUCLEAR+ORGANIZATION |
| 4 | CHROMOSOME CONFORMATION CAPTURE | 267218 | 1% | 59% | 35 | Search CHROMOSOME+CONFORMATION+CAPTURE | Search CHROMOSOME+CONFORMATION+CAPTURE |
| 5 | HI C | 212203 | 1% | 63% | 26 | Search HI+C | Search HI+C |
| 6 | 3D FISH | 208514 | 1% | 90% | 18 | Search 3D+FISH | Search 3D+FISH |
| 7 | GENOME THEORY | 168185 | 1% | 93% | 14 | Search GENOME+THEORY | Search GENOME+THEORY |
| 8 | GENE POSITIONING | 148395 | 1% | 82% | 14 | Search GENE+POSITIONING | Search GENE+POSITIONING |
| 9 | TRANSCRIPTION FACTORY | 141888 | 1% | 53% | 21 | Search TRANSCRIPTION+FACTORY | Search TRANSCRIPTION+FACTORY |
| 10 | INTERPHASE CHROMOSOMES | 131630 | 1% | 68% | 15 | Search INTERPHASE+CHROMOSOMES | Search INTERPHASE+CHROMOSOMES |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | DENKER, A , DE LAAT, W , (2016) THE SECOND DECADE OF 3C TECHNOLOGIES: DETAILED INSIGHTS INTO NUCLEAR ORGANIZATION.GENES & DEVELOPMENT. VOL. 30. ISSUE 12. P. 1357 -1382 | 133 | 67% | 8 |
| 2 | CREMER, T , CREMER, M , (2010) CHROMOSOME TERRITORIES.COLD SPRING HARBOR PERSPECTIVES IN BIOLOGY. VOL. 2. ISSUE 3. P. - | 106 | 80% | 288 |
| 3 | FRASER, J , WILLIAMSON, I , BICKMORE, WA , DOSTIE, J , (2015) AN OVERVIEW OF GENOME ORGANIZATION AND HOW WE GOT THERE: FROM FISH TO HI-C.MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS. VOL. 79. ISSUE 3. P. 347 -372 | 161 | 55% | 17 |
| 4 | DEKKER, J , MARTI-RENOM, MA , MIRNY, LA , (2013) EXPLORING THE THREE-DIMENSIONAL ORGANIZATION OF GENOMES: INTERPRETING CHROMATIN INTERACTION DATA.NATURE REVIEWS GENETICS. VOL. 14. ISSUE 6. P. 390 -403 | 72 | 69% | 246 |
| 5 | SCHMITT, AD , HU, M , REN, B , (2016) GENOME-WIDE MAPPING AND ANALYSIS OF CHROMOSOME ARCHITECTURE.NATURE REVIEWS MOLECULAR CELL BIOLOGY. VOL. 17. ISSUE 12. P. 743 -755 | 92 | 83% | 3 |
| 6 | SOLOVEI, I , THANISCH, K , FEODOROVA, Y , (2016) HOW TO RULE THE NUCLEUS: DIVIDE ET IMPERA.CURRENT OPINION IN CELL BIOLOGY. VOL. 40. ISSUE . P. 47 -59 | 84 | 78% | 4 |
| 7 | DEKKER, J , MIRNY, L , (2016) THE 3D GENOME AS MODERATOR OF CHROMOSOMAL COMMUNICATION.CELL. VOL. 164. ISSUE 6. P. 1110 -1121 | 61 | 57% | 28 |
| 8 | ROUQUETTE, J , CREMER, C , CREMER, T , FAKAN, S , (2010) FUNCTIONAL NUCLEAR ARCHITECTURE STUDIED BY MICROSCOPY: PRESENT AND FUTURE.INTERNATIONAL REVIEW OF CELL AND MOLECULAR BIOLOGY, VOL 282. VOL. 282. ISSUE . P. 1-90 | 192 | 45% | 33 |
| 9 | AY, F , NOBLE, WS , (2015) ANALYSIS METHODS FOR STUDYING THE 3D ARCHITECTURE OF THE GENOME.GENOME BIOLOGY. VOL. 16. ISSUE . P. - | 88 | 80% | 10 |
| 10 | PUESCHEL, R , CORAGGIO, F , MEISTER, P , (2016) FROM SINGLE GENES TO ENTIRE GENOMES: THE SEARCH FOR A FUNCTION OF NUCLEAR ORGANIZATION.DEVELOPMENT. VOL. 143. ISSUE 6. P. 910 -923 | 94 | 78% | 1 |
Classes with closest relation at Level 2 |