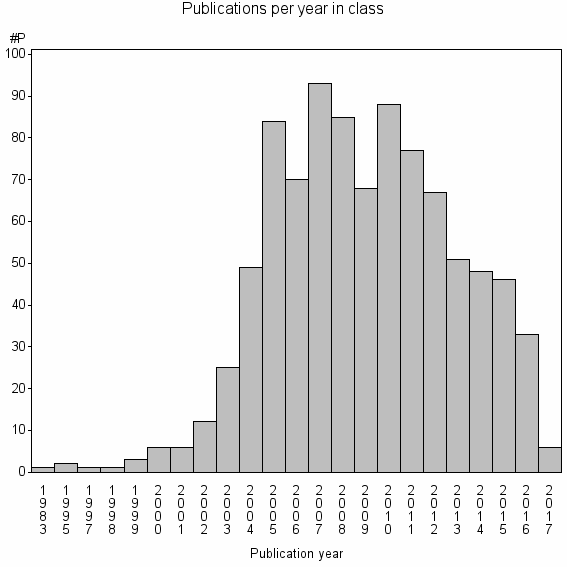
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 12230 | 922 | 34.9 | 85% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 1101 | 2 | BIOINFORMATICS//COPY NUMBER VARIATION//NEXT GENERATION SEQUENCING | 9808 |
| 12230 | 1 | DNA COPY NUMBER//HLTH SOLUT GRP//ARRAY CGH | 922 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | DNA COPY NUMBER | authKW | 264905 | 3% | 33% | 24 |
| 2 | HLTH SOLUT GRP | address | 132464 | 1% | 67% | 6 |
| 3 | ARRAY CGH | authKW | 132136 | 7% | 6% | 67 |
| 4 | COPY NUMBER ALTERATION | authKW | 108667 | 2% | 21% | 16 |
| 5 | MATRIX CGH | authKW | 88310 | 0% | 67% | 4 |
| 6 | CIRCULAR BINARY SEGMENTATION | authKW | 74512 | 0% | 75% | 3 |
| 7 | ACGH DATA | authKW | 66234 | 0% | 100% | 2 |
| 8 | ACGH MICROARRAY DATA | authKW | 66234 | 0% | 100% | 2 |
| 9 | ARRAY CGH DATA | authKW | 66234 | 0% | 100% | 2 |
| 10 | BREAST CANC FUNCT GEN | address | 66234 | 0% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 30170 | 33% | 0% | 306 |
| 2 | Biochemical Research Methods | 7262 | 30% | 0% | 275 |
| 3 | Biotechnology & Applied Microbiology | 6275 | 35% | 0% | 325 |
| 4 | Genetics & Heredity | 3405 | 26% | 0% | 239 |
| 5 | Statistics & Probability | 2064 | 12% | 0% | 110 |
| 6 | Oncology | 1650 | 23% | 0% | 214 |
| 7 | Pathology | 164 | 4% | 0% | 41 |
| 8 | Biochemistry & Molecular Biology | 130 | 14% | 0% | 132 |
| 9 | Computer Science, Interdisciplinary Applications | 118 | 4% | 0% | 34 |
| 10 | Cell Biology | 32 | 5% | 0% | 48 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | HLTH SOLUT GRP | 132464 | 1% | 67% | 6 |
| 2 | BREAST CANC FUNCT GEN | 66234 | 0% | 100% | 2 |
| 3 | INTEGRE BIOINFORMAT | 66234 | 0% | 100% | 2 |
| 4 | CANC GENET DEV BIOL | 39977 | 1% | 15% | 8 |
| 5 | ADDENBROOKES HOPSITAL | 33117 | 0% | 100% | 1 |
| 6 | ASSAY DEV BIOINFORMAT | 33117 | 0% | 100% | 1 |
| 7 | BELFER CANC GENOM | 33117 | 0% | 100% | 1 |
| 8 | BIOENGN INFORMAT RADIOL | 33117 | 0% | 100% | 1 |
| 9 | BIOINFORMAT PREDICT GENOM TEAM | 33117 | 0% | 100% | 1 |
| 10 | BIOL BIOCHEM BIOINFORMAT | 33117 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOINFORMATICS | 55861 | 15% | 1% | 134 |
| 2 | BMC BIOINFORMATICS | 22822 | 8% | 1% | 72 |
| 3 | BIOSTATISTICS | 12364 | 2% | 2% | 17 |
| 4 | BMC MEDICAL GENOMICS | 8864 | 2% | 2% | 14 |
| 5 | STATISTICAL APPLICATIONS IN GENETICS AND MOLECULAR BIOLOGY | 5294 | 1% | 2% | 9 |
| 6 | GENES CHROMOSOMES & CANCER | 5029 | 2% | 1% | 22 |
| 7 | IEEE-ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS | 4197 | 1% | 1% | 12 |
| 8 | BMC GENOMICS | 3499 | 3% | 0% | 32 |
| 9 | ANNALS OF APPLIED STATISTICS | 3230 | 1% | 1% | 9 |
| 10 | EXPERT REVIEW OF MOLECULAR DIAGNOSTICS | 2620 | 1% | 1% | 9 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DNA COPY NUMBER | 264905 | 3% | 33% | 24 | Search DNA+COPY+NUMBER | Search DNA+COPY+NUMBER |
| 2 | ARRAY CGH | 132136 | 7% | 6% | 67 | Search ARRAY+CGH | Search ARRAY+CGH |
| 3 | COPY NUMBER ALTERATION | 108667 | 2% | 21% | 16 | Search COPY+NUMBER+ALTERATION | Search COPY+NUMBER+ALTERATION |
| 4 | MATRIX CGH | 88310 | 0% | 67% | 4 | Search MATRIX+CGH | Search MATRIX+CGH |
| 5 | CIRCULAR BINARY SEGMENTATION | 74512 | 0% | 75% | 3 | Search CIRCULAR+BINARY+SEGMENTATION | Search CIRCULAR+BINARY+SEGMENTATION |
| 6 | ACGH DATA | 66234 | 0% | 100% | 2 | Search ACGH+DATA | Search ACGH+DATA |
| 7 | ACGH MICROARRAY DATA | 66234 | 0% | 100% | 2 | Search ACGH+MICROARRAY+DATA | Search ACGH+MICROARRAY+DATA |
| 8 | ARRAY CGH DATA | 66234 | 0% | 100% | 2 | Search ARRAY+CGH+DATA | Search ARRAY+CGH+DATA |
| 9 | GENOTYPE CALLING ERROR | 66234 | 0% | 100% | 2 | Search GENOTYPE+CALLING+ERROR | Search GENOTYPE+CALLING+ERROR |
| 10 | COPY NUMBER PROFILE | 44155 | 0% | 67% | 2 | Search COPY+NUMBER+PROFILE | Search COPY+NUMBER+PROFILE |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | VAN DE WIEL, MA , PICARD, F , VAN WIERINGEN, WN , YLSTRA, B , (2011) PREPROCESSING AND DOWNSTREAM ANALYSIS OF MICROARRAY DNA COPY NUMBER PROFILES.BRIEFINGS IN BIOINFORMATICS. VOL. 12. ISSUE 1. P. 10 -21 | 41 | 75% | 19 |
| 2 | CHEN, H , XING, HP , ZHANG, NR , (2011) ESTIMATION OF PARENT SPECIFIC DNA COPY NUMBER IN TUMORS USING HIGH-DENSITY GENOTYPING ARRAYS.PLOS COMPUTATIONAL BIOLOGY. VOL. 7. ISSUE 1. P. - | 30 | 81% | 19 |
| 3 | SEIFERT, M , GOHR, A , STRICKERT, M , GROSSE, I , (2012) PARSIMONIOUS HIGHER-ORDER HIDDEN MARKOV MODELS FOR IMPROVED ARRAY-CGH ANALYSIS WITH APPLICATIONS TO ARABIDOPSIS THALIANA.PLOS COMPUTATIONAL BIOLOGY. VOL. 8. ISSUE 1. P. - | 45 | 51% | 4 |
| 4 | NILSEN, G , LIESTOL, K , VAN LOO, P , VOLLAN, HKM , EIDE, MB , RUEDA, OM , CHIN, SF , RUSSELL, R , BAUMBUSCH, LO , CALDAS, C , ET AL (2012) COPYNUMBER: EFFICIENT ALGORITHMS FOR SINGLE- AND MULTI-TRACK COPY NUMBER SEGMENTATION.BMC GENOMICS. VOL. 13. ISSUE . P. - | 26 | 79% | 25 |
| 5 | PICARD, F , LEBARBIER, E , HOEBEKE, M , RIGAILL, G , THIAM, B , ROBIN, S , (2011) JOINT SEGMENTATION, CALLING, AND NORMALIZATION OF MULTIPLE CGH PROFILES.BIOSTATISTICS. VOL. 12. ISSUE 3. P. 413 -428 | 24 | 86% | 33 |
| 6 | YU, TW , YE, H , SUN, W , LI, KC , CHEN, ZG , JACOBS, S , BAILEY, DK , WONG, DT , ZHOU, XF , (2007) A FORWARD-BACKWARD FRAGMENT ASSEMBLING ALGORITHM FOR THE IDENTIFICATION OF GENOMIC AMPLIFICATION AND DELETION BREAKPOINTS USING HIGH-DENSITY SINGLE NUCLEOTIDE POLYMORPHISM (SNP) ARRAY.BMC BIOINFORMATICS. VOL. 8. ISSUE . P. - | 25 | 96% | 12 |
| 7 | OLSHEN, AB , BENGTSSON, H , NEUVIAL, P , SPELLMAN, PT , OLSHEN, RA , SESHAN, VE , (2011) PARENT-SPECIFIC COPY NUMBER IN PAIRED TUMOR-NORMAL STUDIES USING CIRCULAR BINARY SEGMENTATION.BIOINFORMATICS. VOL. 27. ISSUE 15. P. 2038 -2046 | 27 | 75% | 26 |
| 8 | DALMASSO, C , BROET, P , (2011) DETECTION OF CHROMOSOMAL ABNORMALITIES USING HIGH RESOLUTION ARRAYS IN CLINICAL CANCER RESEARCH.JOURNAL OF BIOMEDICAL INFORMATICS. VOL. 44. ISSUE 6. P. 936-942 | 31 | 70% | 1 |
| 9 | GREENMAN, CD , BIGNELL, G , BUTLER, A , EDKINS, S , HINTON, J , BEARE, D , SWAMY, S , SANTARIUS, T , CHEN, L , WIDAA, S , ET AL (2010) PICNIC: AN ALGORITHM TO PREDICT ABSOLUTE ALLELIC COPY NUMBER VARIATION WITH MICROARRAY CANCER DATA.BIOSTATISTICS. VOL. 11. ISSUE 1. P. 164 -175 | 25 | 71% | 87 |
| 10 | LAI, WR , JOHNSON, MD , KUCHERLAPATI, R , PARK, PJ , (2005) COMPARATIVE ANALYSIS OF ALGORITHMS FOR IDENTIFYING AMPLIFICATIONS AND DELETIONS IN ARRAY CGH DATA.BIOINFORMATICS. VOL. 21. ISSUE 19. P. 3763 -3770 | 23 | 82% | 230 |
Classes with closest relation at Level 1 |