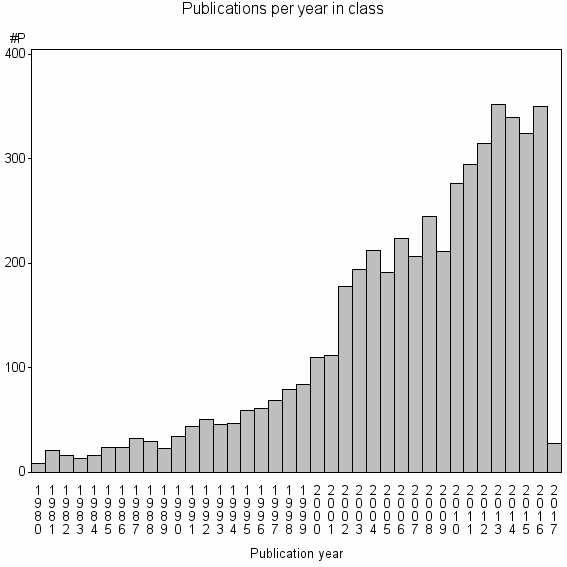
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 2179 | 4941 | 31.4 | 80% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 2179 | 2 | REFERENCE GENES//GENORM//NORMFINDER | 4941 |
| 4517 | 1 | REFERENCE GENES//GENORM//NORMFINDER | 1774 |
| 11368 | 1 | RNA AMPLIFICATION//LASER CAPTURE MICRODISSECTION//LASER MICRODISSECTION | 987 |
| 11477 | 1 | FFPE//FORMALIN FIXED PARAFFIN EMBEDDED TISSUE//METHACARN | 979 |
| 16568 | 1 | RNA ISOLATION//RNA EXTRACTION//DNA ISOLATION | 647 |
| 20679 | 1 | BRAIN BANK//BRAIN DONATION//AGONAL STATE | 449 |
| 34880 | 1 | TRIZOL//ALLEGRO//FUNCTIONAL INFORMATICS | 105 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | REFERENCE GENES | authKW | 2595959 | 11% | 81% | 519 |
| 2 | GENORM | authKW | 495075 | 2% | 83% | 96 |
| 3 | NORMFINDER | authKW | 424803 | 2% | 90% | 76 |
| 4 | HOUSEKEEPING GENES | authKW | 360081 | 3% | 46% | 128 |
| 5 | RNA ISOLATION | authKW | 252675 | 2% | 47% | 87 |
| 6 | HOUSEKEEPING GENE | authKW | 247892 | 2% | 47% | 85 |
| 7 | RNA INTEGRITY | authKW | 184579 | 1% | 85% | 35 |
| 8 | BESTKEEPER | authKW | 173755 | 1% | 94% | 30 |
| 9 | RNA EXTRACTION | authKW | 158227 | 2% | 32% | 81 |
| 10 | LASER CAPTURE MICRODISSECTION | authKW | 152341 | 3% | 18% | 136 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemical Research Methods | 12727 | 18% | 0% | 865 |
| 2 | Pathology | 8235 | 12% | 0% | 595 |
| 3 | Biochemistry & Molecular Biology | 4961 | 29% | 0% | 1435 |
| 4 | Biotechnology & Applied Microbiology | 4381 | 14% | 0% | 681 |
| 5 | Plant Sciences | 1839 | 10% | 0% | 490 |
| 6 | Genetics & Heredity | 1692 | 9% | 0% | 445 |
| 7 | Medical Laboratory Technology | 1109 | 3% | 0% | 135 |
| 8 | Cell Biology | 526 | 7% | 0% | 363 |
| 9 | Neurosciences | 497 | 8% | 0% | 418 |
| 10 | Oncology | 307 | 6% | 0% | 311 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | PATHOGENET UNIT | 63401 | 1% | 38% | 27 |
| 2 | CANC GENOME ANAT PROJECT | 23250 | 0% | 47% | 8 |
| 3 | BIOREPOSITORIES BIOSPECIMEN BRANCH | 22236 | 0% | 60% | 6 |
| 4 | NEW S WALES TISSUE OURCE | 13899 | 0% | 75% | 3 |
| 5 | MOL BIOL CORE SERV | 12356 | 0% | 100% | 2 |
| 6 | MURINE MICROARRAY AFFYMETRIX IL | 12356 | 0% | 100% | 2 |
| 7 | RUMENTAT DEV OURCE BIOENGN PHYS SC | 12356 | 0% | 100% | 2 |
| 8 | SPIDIA CONSORTIUM | 12356 | 0% | 100% | 2 |
| 9 | HORMONE RECEPTOR | 11875 | 0% | 38% | 5 |
| 10 | LASER MOL BIOL | 11118 | 0% | 60% | 3 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BMC MOLECULAR BIOLOGY | 68916 | 2% | 12% | 92 |
| 2 | BIOTECHNIQUES | 35045 | 4% | 3% | 186 |
| 3 | DIAGNOSTIC MOLECULAR PATHOLOGY | 25629 | 1% | 7% | 57 |
| 4 | PLANT MOLECULAR BIOLOGY REPORTER | 25004 | 2% | 5% | 75 |
| 5 | BIOPRESERVATION AND BIOBANKING | 17915 | 1% | 9% | 32 |
| 6 | JOURNAL OF MOLECULAR DIAGNOSTICS | 10417 | 1% | 4% | 44 |
| 7 | MOLECULAR BIOTECHNOLOGY | 7298 | 1% | 3% | 47 |
| 8 | ANALYTICAL BIOCHEMISTRY | 5379 | 3% | 1% | 129 |
| 9 | CELL AND TISSUE BANKING | 3961 | 0% | 4% | 18 |
| 10 | PLANT METHODS | 3281 | 0% | 4% | 15 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | REFERENCE GENES | 2595959 | 11% | 81% | 519 | Search REFERENCE+GENES | Search REFERENCE+GENES |
| 2 | GENORM | 495075 | 2% | 83% | 96 | Search GENORM | Search GENORM |
| 3 | NORMFINDER | 424803 | 2% | 90% | 76 | Search NORMFINDER | Search NORMFINDER |
| 4 | HOUSEKEEPING GENES | 360081 | 3% | 46% | 128 | Search HOUSEKEEPING+GENES | Search HOUSEKEEPING+GENES |
| 5 | RNA ISOLATION | 252675 | 2% | 47% | 87 | Search RNA+ISOLATION | Search RNA+ISOLATION |
| 6 | HOUSEKEEPING GENE | 247892 | 2% | 47% | 85 | Search HOUSEKEEPING+GENE | Search HOUSEKEEPING+GENE |
| 7 | RNA INTEGRITY | 184579 | 1% | 85% | 35 | Search RNA+INTEGRITY | Search RNA+INTEGRITY |
| 8 | BESTKEEPER | 173755 | 1% | 94% | 30 | Search BESTKEEPER | Search BESTKEEPER |
| 9 | RNA EXTRACTION | 158227 | 2% | 32% | 81 | Search RNA+EXTRACTION | Search RNA+EXTRACTION |
| 10 | LASER CAPTURE MICRODISSECTION | 152341 | 3% | 18% | 136 | Search LASER+CAPTURE+MICRODISSECTION | Search LASER+CAPTURE+MICRODISSECTION |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | KOZERA, B , RAPACZ, M , (2013) REFERENCE GENES IN REAL-TIME PCR.JOURNAL OF APPLIED GENETICS. VOL. 54. ISSUE 4. P. 391 -406 | 61 | 82% | 102 |
| 2 | CHAPMAN, JR , WALDENSTROM, J , (2015) WITH REFERENCE TO REFERENCE GENES: A SYSTEMATIC REVIEW OF ENDOGENOUS CONTROLS IN GENE EXPRESSION STUDIES.PLOS ONE. VOL. 10. ISSUE 11. P. - | 69 | 91% | 4 |
| 3 | MA, R , XU, S , ZHAO, YC , XIA, B , WANG, R , (2016) SELECTION AND VALIDATION OF APPROPRIATE REFERENCE GENES FOR QUANTITATIVE REAL-TIME PCR ANALYSIS OF GENE EXPRESSION IN LYCORIS AUREA.FRONTIERS IN PLANT SCIENCE. VOL. 7. ISSUE . P. - | 64 | 77% | 2 |
| 4 | REDDY, DS , BHATNAGAR-MATHUR, P , REDDY, PS , CINDHURI, KS , GANESH, AS , SHARMA, KK , (2016) IDENTIFICATION AND VALIDATION OF REFERENCE GENES AND THEIR IMPACT ON NORMALIZED GENE EXPRESSION STUDIES ACROSS CULTIVATED AND WILD CICER SPECIES.PLOS ONE. VOL. 11. ISSUE 2. P. - | 52 | 88% | 3 |
| 5 | WANG, T , HAO, RJ , PAN, HT , CHENG, TR , ZHANG, QX , (2014) SELECTION OF SUITABLE REFERENCE GENES FOR QUANTITATIVE REAL-TIME POLYMERASE CHAIN REACTION IN PRUNUS MUME DURING FLOWERING STAGES AND UNDER DIFFERENT ABIOTIC STRESS CONDITIONS.JOURNAL OF THE AMERICAN SOCIETY FOR HORTICULTURAL SCIENCE. VOL. 139. ISSUE 2. P. 113 -122 | 67 | 83% | 5 |
| 6 | BONIN, S , STANTA, G , (2013) NUCLEIC ACID EXTRACTION METHODS FROM FIXED AND PARAFFIN-EMBEDDED TISSUES IN CANCER DIAGNOSTICS.EXPERT REVIEW OF MOLECULAR DIAGNOSTICS. VOL. 13. ISSUE 3. P. 271 -282 | 65 | 82% | 7 |
| 7 | LEGRES, LG , JANIN, A , MASSELON, C , BERTHEAU, P , (2014) BEYOND LASER MICRODISSECTION TECHNOLOGY: FOLLOW THE YELLOW BRICK ROAD FOR CANCER RESEARCH.AMERICAN JOURNAL OF CANCER RESEARCH. VOL. 4. ISSUE 1. P. 1-28 | 93 | 53% | 8 |
| 8 | MANOLI, A , STURARO, A , TREVISAN, S , QUAGGIOTTI, S , NONIS, A , (2012) EVALUATION OF CANDIDATE REFERENCE GENES FOR QPCR IN MAIZE.JOURNAL OF PLANT PHYSIOLOGY. VOL. 169. ISSUE 8. P. 807 -815 | 51 | 88% | 50 |
| 9 | RUIJTER, JM , RAMAKERS, C , HOOGAARS, WMH , KARLEN, Y , BAKKER, O , VAN DEN HOFF, MJB , MOORMAN, AFM , (2009) AMPLIFICATION EFFICIENCY: LINKING BASELINE AND BIAS IN THE ANALYSIS OF QUANTITATIVE PCR DATA.NUCLEIC ACIDS RESEARCH. VOL. 37. ISSUE 6. P. - | 27 | 93% | 744 |
| 10 | XIAO, XL , WU, XM , MA, JB , LI, PB , LI, TT , YAO, YA , (2015) SYSTEMATIC ASSESSMENT OF REFERENCE GENES FOR RT-QPCR ACROSS PLANT SPECIES UNDER SALT STRESS AND DROUGHT STRESS.ACTA PHYSIOLOGIAE PLANTARUM. VOL. 37. ISSUE 9. P. - | 53 | 88% | 1 |
Classes with closest relation at Level 2 |