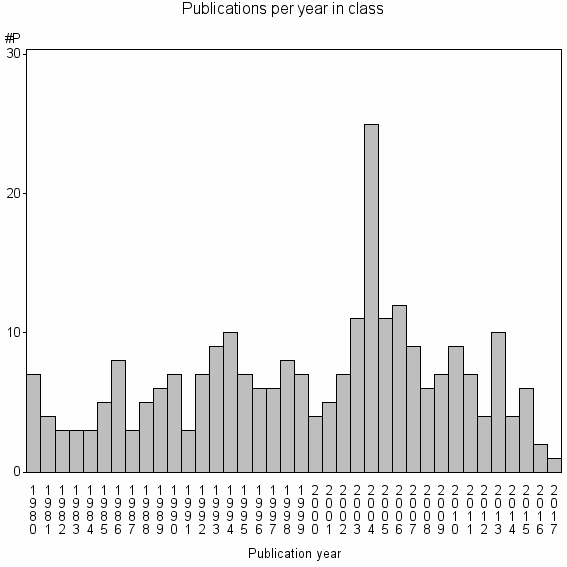
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 26332 | 257 | 31.7 | 68% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 219 | 3 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN FOLDING//PROTEIN STRUCTURE PREDICTION | 50409 |
| 365 | 2 | PROTEIN STRUCTURE PREDICTION//PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PSEUDO AMINO ACID COMPOSITION | 17331 |
| 26332 | 1 | CYSTEINE BONDING STATE//DISULFIDE CONNECTIVITY PREDICTION//MEDIUM AND LONG RANGE | 257 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | CYSTEINE BONDING STATE | authKW | 356443 | 1% | 100% | 3 |
| 2 | DISULFIDE CONNECTIVITY PREDICTION | authKW | 356443 | 1% | 100% | 3 |
| 3 | MEDIUM AND LONG RANGE | authKW | 356443 | 1% | 100% | 3 |
| 4 | BONDING STATE OF CYSTEINES | authKW | 237629 | 1% | 100% | 2 |
| 5 | CLANTOX | authKW | 237629 | 1% | 100% | 2 |
| 6 | CSU SOFTWARE | authKW | 237629 | 1% | 100% | 2 |
| 7 | CYSTEINE REDOX STATE PREDICTION | authKW | 237629 | 1% | 100% | 2 |
| 8 | DISULFIDE BONDING STATE | authKW | 237629 | 1% | 100% | 2 |
| 9 | HIDDEN NEURAL NETWORKS | authKW | 237629 | 1% | 100% | 2 |
| 10 | PAIR PREFERENCES | authKW | 237629 | 1% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 1997 | 16% | 0% | 42 |
| 2 | Biochemistry & Molecular Biology | 1082 | 54% | 0% | 138 |
| 3 | Biophysics | 851 | 21% | 0% | 55 |
| 4 | Biochemical Research Methods | 601 | 17% | 0% | 43 |
| 5 | Biology | 184 | 8% | 0% | 21 |
| 6 | Biotechnology & Applied Microbiology | 168 | 12% | 0% | 31 |
| 7 | Polymer Science | 112 | 9% | 0% | 23 |
| 8 | Computer Science, Interdisciplinary Applications | 37 | 4% | 0% | 10 |
| 9 | Evolutionary Biology | 32 | 3% | 0% | 7 |
| 10 | Medical Informatics | 18 | 1% | 0% | 3 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MACHINE LEARNING NEURAL NETWORKS GRP | 152758 | 1% | 43% | 3 |
| 2 | CIRI LIFE SCI HLTH TECHNOL | 118814 | 0% | 100% | 1 |
| 3 | EQUIPE STAT SEQUENCES BIOL | 118814 | 0% | 100% | 1 |
| 4 | GEN PROTEO | 118814 | 0% | 100% | 1 |
| 5 | POLYMER PLASMA | 118814 | 0% | 100% | 1 |
| 6 | RIKEN LIFE SCI | 118814 | 0% | 100% | 1 |
| 7 | STRUCT BIOL SBRC | 118814 | 0% | 100% | 1 |
| 8 | INTERUNIV BIOINFORMAT BRUSSELS | 97207 | 1% | 27% | 3 |
| 9 | SUDARSKY COMPUTAT BIOL | 35639 | 1% | 10% | 3 |
| 10 | SEAVER FDN BIOINFORMAT | 33944 | 1% | 14% | 2 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS | 4753 | 6% | 0% | 16 |
| 2 | PROTEIN ENGINEERING | 4246 | 4% | 0% | 9 |
| 3 | ACTA POLYMERICA SINICA | 4004 | 4% | 0% | 11 |
| 4 | PROTEIN SCIENCE | 3188 | 5% | 0% | 13 |
| 5 | BIOINFORMATICS | 2839 | 6% | 0% | 16 |
| 6 | INTERNATIONAL JOURNAL OF PEPTIDE AND PROTEIN RESEARCH | 2315 | 3% | 0% | 7 |
| 7 | JOURNAL OF BIOSCIENCES | 1755 | 2% | 0% | 6 |
| 8 | JOURNAL OF BIOLOGICAL PHYSICS | 1435 | 1% | 0% | 3 |
| 9 | INTERNATIONAL JOURNAL OF BIO-MEDICAL COMPUTING | 1314 | 1% | 0% | 3 |
| 10 | PREPARATIVE BIOCHEMISTRY & BIOTECHNOLOGY | 1303 | 1% | 0% | 3 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CYSTEINE BONDING STATE | 356443 | 1% | 100% | 3 | Search CYSTEINE+BONDING+STATE | Search CYSTEINE+BONDING+STATE |
| 2 | DISULFIDE CONNECTIVITY PREDICTION | 356443 | 1% | 100% | 3 | Search DISULFIDE+CONNECTIVITY+PREDICTION | Search DISULFIDE+CONNECTIVITY+PREDICTION |
| 3 | MEDIUM AND LONG RANGE | 356443 | 1% | 100% | 3 | Search MEDIUM+AND+LONG+RANGE | Search MEDIUM+AND+LONG+RANGE |
| 4 | BONDING STATE OF CYSTEINES | 237629 | 1% | 100% | 2 | Search BONDING+STATE+OF+CYSTEINES | Search BONDING+STATE+OF+CYSTEINES |
| 5 | CLANTOX | 237629 | 1% | 100% | 2 | Search CLANTOX | Search CLANTOX |
| 6 | CSU SOFTWARE | 237629 | 1% | 100% | 2 | Search CSU+SOFTWARE | Search CSU+SOFTWARE |
| 7 | CYSTEINE REDOX STATE PREDICTION | 237629 | 1% | 100% | 2 | Search CYSTEINE+REDOX+STATE+PREDICTION | Search CYSTEINE+REDOX+STATE+PREDICTION |
| 8 | DISULFIDE BONDING STATE | 237629 | 1% | 100% | 2 | Search DISULFIDE+BONDING+STATE | Search DISULFIDE+BONDING+STATE |
| 9 | HIDDEN NEURAL NETWORKS | 237629 | 1% | 100% | 2 | Search HIDDEN+NEURAL+NETWORKS | Search HIDDEN+NEURAL+NETWORKS |
| 10 | PAIR PREFERENCES | 237629 | 1% | 100% | 2 | Search PAIR+PREFERENCES | Search PAIR+PREFERENCES |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | THANGUDU, RR , SHARMA, P , SRINIVASAN, N , OFFMANN, B , (2007) ANALYCYS: A DATABASE FOR CONSERVATION AND CONFORMATION OF DISULPHIDE BONDS IN HOMOLOGOUS PROTEIN DOMAINS.PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS. VOL. 67. ISSUE 2. P. 255 -261 | 31 | 72% | 7 |
| 2 | ELUMALAI, P , WU, JW , LIU, HL , (2010) CURRENT ADVANCES IN DISULFIDE CONNECTIVITY PREDICTIONS.JOURNAL OF THE TAIWAN INSTITUTE OF CHEMICAL ENGINEERS. VOL. 41. ISSUE 5. P. 525 -539 | 32 | 63% | 2 |
| 3 | MARQUEZ-CHAMORRO, AE , AGUILAR-RUIZ, JS , (2015) SOFT COMPUTING METHODS FOR DISULFIDE CONNECTIVITY PREDICTION.EVOLUTIONARY BIOINFORMATICS. VOL. 11. ISSUE . P. 223 -229 | 20 | 80% | 0 |
| 4 | VINCENT, M , PASSERINI, A , LABBE, M , FRASCONI, P , (2008) A SIMPLIFIED APPROACH TO DISULFIDE CONNECTIVITY PREDICTION FROM PROTEIN SEQUENCES.BMC BIOINFORMATICS. VOL. 9. ISSUE . P. - | 19 | 83% | 11 |
| 5 | CERONI, A , PASSERINI, A , VULLO, A , FRASCONI, P , (2006) DISULFIND: A DISULFIDE BONDING STATE AND CYSTEINE CONNECTIVITY PREDICTION SERVER.NUCLEIC ACIDS RESEARCH. VOL. 34. ISSUE . P. W177-W181 | 14 | 88% | 110 |
| 6 | RUBINSTEIN, R , FISER, A , (2008) PREDICTING DISULFIDE BOND CONNECTIVITY IN PROTEINS BY CORRELATED MUTATIONS ANALYSIS.BIOINFORMATICS. VOL. 24. ISSUE 4. P. 498 -504 | 22 | 58% | 23 |
| 7 | LIN, HH , TSENG, LY , (2010) DBCP: A WEB SERVER FOR DISULFIDE BONDING CONNECTIVITY PATTERN PREDICTION WITHOUT THE PRIOR KNOWLEDGE OF THE BONDING STATE OF CYSTEINES.NUCLEIC ACIDS RESEARCH. VOL. 38. ISSUE . P. W503-W507 | 14 | 82% | 19 |
| 8 | SINGH, R , MURAD, W , (2013) PROTEIN DISULFIDE TOPOLOGY DETERMINATION THROUGH THE FUSION OF MASS SPECTROMETRIC ANALYSIS AND SEQUENCE-BASED PREDICTION USING DEMPSTER-SHAFER THEORY.BMC BIOINFORMATICS. VOL. 14. ISSUE . P. - | 16 | 70% | 1 |
| 9 | THANGUDU, RR , MANOHARAN, M , SRINIVASAN, N , CADET, F , SOWDHAMINI, R , OFFMANN, B , (2008) ANALYSIS ON CONSERVATION OF DISULPHIDE BONDS AND THEIR STRUCTURAL FEATURES IN HOMOLOGOUS PROTEIN DOMAIN FAMILIES.BMC STRUCTURAL BIOLOGY. VOL. 8. ISSUE . P. - | 33 | 36% | 16 |
| 10 | LIN, HH , HSU, JC , HSU, YN , PAN, RH , CHEN, YF , TSENG, LY , (2013) DISULFIDE CONNECTIVITY PREDICTION BASED ON STRUCTURAL INFORMATION WITHOUT A PRIOR KNOWLEDGE OF THE BONDING STATE OF CYSTEINES.COMPUTERS IN BIOLOGY AND MEDICINE. VOL. 43. ISSUE 11. P. 1941-1948 | 17 | 61% | 0 |
Classes with closest relation at Level 1 |