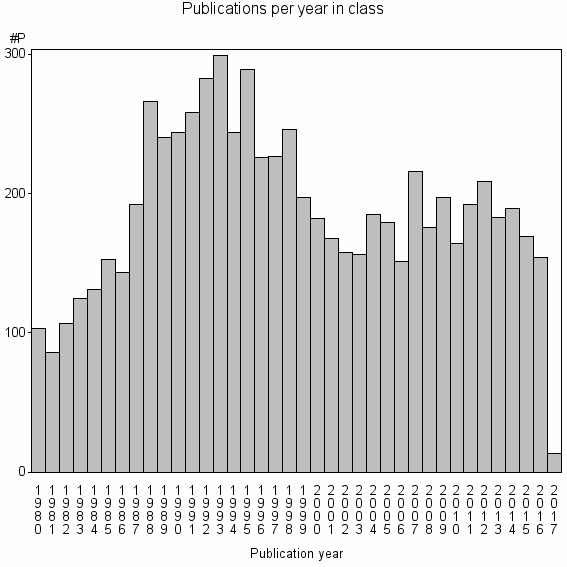
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 1609 | 7100 | 27.9 | 71% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 7 | 4 | INFECTIOUS DISEASES//MICROBIOLOGY//VIROLOGY | 1353914 |
| 33 | 3 | JOURNAL OF BACTERIOLOGY//MICROBIOLOGY//MOLECULAR MICROBIOLOGY | 107033 |
| 1609 | 2 | RESTRICTION ENDONUCLEASE//BIOTECHNIQUES//RESTRICTION MODIFICATION SYSTEM | 7100 |
| 784 | 1 | RESTRICTION ENDONUCLEASE//RESTRICTION MODIFICATION SYSTEM//RESTRICTION MODIFICATION | 3079 |
| 12652 | 1 | PLASMID PURIFICATION//PLASMID DNA//SUPERCOILED PLASMID DNA | 891 |
| 12753 | 1 | T VECTOR//TA CLONING//GENE SYNTHESIS | 883 |
| 15387 | 1 | MEGAPRIMER//MULTIPLE SITE DIRECTED MUTAGENESIS//OVERLAP EXTENSION | 715 |
| 21162 | 1 | NOTI LINKING CLONES//NOTI LINKING AND JUMPING CLONES//NOTL LINKING CLONES | 430 |
| 21956 | 1 | GENOME WALKING//FALSE POSITIVE EXCLUSION//UNKNOWN FLANKING SEQUENCES | 399 |
| 22511 | 1 | GC RICH DNA//HOT START//AU GOLD NANOPARTICLES | 379 |
| 33383 | 1 | SPEL 6//AND PLASMIDS//ASSEMBLY OF DNA FRAGMENTS | 125 |
| 35141 | 1 | CALLIGONUM COMOSUM//CALLIGONUM//CALLIGONUM SPECIES | 102 |
| 35472 | 1 | ANALYTICAL PITFALLS//ARTEFACTUAL RESULTS OF LABORATORY TESTS//AUTOMATED BLOOD COUNTERS | 97 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | RESTRICTION ENDONUCLEASE | authKW | 274358 | 2% | 39% | 163 |
| 2 | BIOTECHNIQUES | journal | 251022 | 8% | 10% | 595 |
| 3 | RESTRICTION MODIFICATION SYSTEM | authKW | 217896 | 1% | 65% | 78 |
| 4 | RESTRICTION MODIFICATION | authKW | 201451 | 1% | 63% | 75 |
| 5 | NUCLEIC ACIDS RESEARCH | journal | 140170 | 16% | 3% | 1135 |
| 6 | DNA PROT INTERACT UNIT | address | 91321 | 1% | 51% | 42 |
| 7 | ISOSCHIZOMER | authKW | 89526 | 0% | 77% | 27 |
| 8 | PLASMID PURIFICATION | authKW | 80017 | 0% | 85% | 22 |
| 9 | RESTRICTION ENZYME | authKW | 73626 | 1% | 28% | 61 |
| 10 | PLASMID DNA | authKW | 71672 | 2% | 13% | 129 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 43539 | 64% | 0% | 4520 |
| 2 | Biochemical Research Methods | 25234 | 20% | 0% | 1447 |
| 3 | Genetics & Heredity | 6831 | 14% | 0% | 995 |
| 4 | Biotechnology & Applied Microbiology | 5495 | 13% | 0% | 922 |
| 5 | Microbiology | 1166 | 7% | 0% | 481 |
| 6 | Chemistry, Analytical | 981 | 7% | 0% | 464 |
| 7 | Biophysics | 709 | 5% | 0% | 336 |
| 8 | Cell Biology | 206 | 5% | 0% | 352 |
| 9 | Biology | 25 | 1% | 0% | 91 |
| 10 | Virology | 2 | 1% | 0% | 48 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | DNA PROT INTERACT UNIT | 91321 | 1% | 51% | 42 |
| 2 | BIOINFORMAT PROT ENGN | 28408 | 0% | 19% | 35 |
| 3 | NIPPONKO KU | 21494 | 0% | 100% | 5 |
| 4 | PROGRAM BIOPHYS BIOCHEM | 20013 | 0% | 27% | 17 |
| 5 | CICS UBI INVEST CIENCIAS SAUDE | 19954 | 0% | 39% | 12 |
| 6 | BIOPHYS S | 19615 | 0% | 24% | 19 |
| 7 | KEY HORMONE DEV | 17910 | 0% | 83% | 5 |
| 8 | ABT BIOPHYS BIOCHEM VERFAHREN | 13755 | 0% | 80% | 4 |
| 9 | IBBS BIOPHYS S | 13430 | 0% | 63% | 5 |
| 10 | ABT STRUKTURANAL | 12896 | 0% | 100% | 3 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOTECHNIQUES | 251022 | 8% | 10% | 595 |
| 2 | NUCLEIC ACIDS RESEARCH | 140170 | 16% | 3% | 1135 |
| 3 | GENE | 54410 | 7% | 3% | 497 |
| 4 | PCR-METHODS AND APPLICATIONS | 17676 | 0% | 16% | 26 |
| 5 | ANALYTICAL BIOCHEMISTRY | 16019 | 4% | 1% | 265 |
| 6 | GENE ANALYSIS TECHNIQUES | 14330 | 0% | 19% | 18 |
| 7 | MOLECULAR BIOTECHNOLOGY | 12263 | 1% | 4% | 73 |
| 8 | MOLECULAR BIOLOGY | 11112 | 2% | 2% | 111 |
| 9 | JOURNAL OF MOLECULAR BIOLOGY | 7364 | 3% | 1% | 204 |
| 10 | ACS SYNTHETIC BIOLOGY | 5998 | 0% | 5% | 28 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | RESTRICTION ENDONUCLEASE | 274358 | 2% | 39% | 163 | Search RESTRICTION+ENDONUCLEASE | Search RESTRICTION+ENDONUCLEASE |
| 2 | RESTRICTION MODIFICATION SYSTEM | 217896 | 1% | 65% | 78 | Search RESTRICTION+MODIFICATION+SYSTEM | Search RESTRICTION+MODIFICATION+SYSTEM |
| 3 | RESTRICTION MODIFICATION | 201451 | 1% | 63% | 75 | Search RESTRICTION+MODIFICATION | Search RESTRICTION+MODIFICATION |
| 4 | ISOSCHIZOMER | 89526 | 0% | 77% | 27 | Search ISOSCHIZOMER | Search ISOSCHIZOMER |
| 5 | PLASMID PURIFICATION | 80017 | 0% | 85% | 22 | Search PLASMID+PURIFICATION | Search PLASMID+PURIFICATION |
| 6 | RESTRICTION ENZYME | 73626 | 1% | 28% | 61 | Search RESTRICTION+ENZYME | Search RESTRICTION+ENZYME |
| 7 | PLASMID DNA | 71672 | 2% | 13% | 129 | Search PLASMID+DNA | Search PLASMID+DNA |
| 8 | DNA METHYLTRANSFERASE | 65625 | 1% | 14% | 106 | Search DNA+METHYLTRANSFERASE | Search DNA+METHYLTRANSFERASE |
| 9 | SITE SPECIFIC ENDONUCLEASE | 61400 | 0% | 71% | 20 | Search SITE+SPECIFIC+ENDONUCLEASE | Search SITE+SPECIFIC+ENDONUCLEASE |
| 10 | T VECTOR | 59153 | 0% | 81% | 17 | Search T+VECTOR | Search T+VECTOR |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | PINGOUD, A , WILSON, GG , WENDE, W , (2014) TYPE II RESTRICTION ENDONUCLEASES-A HISTORICAL PERSPECTIVE AND MORE.NUCLEIC ACIDS RESEARCH. VOL. 42. ISSUE 12. P. 7489 -7527 | 306 | 82% | 26 |
| 2 | LOENEN, WAM , DRYDEN, DTF , RALEIGH, EA , WILSON, GG , (2014) TYPE I RESTRICTION ENZYMES AND THEIR RELATIVES.NUCLEIC ACIDS RESEARCH. VOL. 42. ISSUE 1. P. 20 -44 | 167 | 83% | 31 |
| 3 | MCCLELLAND, M , NELSON, M , RASCHKE, E , (1994) EFFECT OF SITE-SPECIFIC MODIFICATION ON RESTRICTION ENDONUCLEASES AND DNA MODIFICATION METHYLTRANSFERASES.NUCLEIC ACIDS RESEARCH. VOL. 22. ISSUE 17. P. 3640-3659 | 270 | 84% | 276 |
| 4 | KESSLER, C , MANTA, V , (1990) SPECIFICITY OF RESTRICTION ENDONUCLEASES AND DNA MODIFICATION METHYLTRANSFERASES - A REVIEW.GENE. VOL. 92. ISSUE 1-2. P. 1-248 | 382 | 82% | 112 |
| 5 | MADHUSOODANAN, UK , RAO, DN , (2010) DIVERSITY OF DNA METHYLTRANSFERASES THAT RECOGNIZE ASYMMETRIC TARGET SEQUENCES.CRITICAL REVIEWS IN BIOCHEMISTRY AND MOLECULAR BIOLOGY. VOL. 45. ISSUE 2. P. 125 -145 | 155 | 90% | 11 |
| 6 | NELSON, M , RASCHKE, E , MCCLELLAND, M , (1993) EFFECT OF SITE-SPECIFIC METHYLATION ON RESTRICTION ENDONUCLEASES AND DNA MODIFICATION METHYLTRANSFERASES.NUCLEIC ACIDS RESEARCH. VOL. 21. ISSUE 13. P. 3139-3154 | 246 | 84% | 62 |
| 7 | PINGOUD, A , FUXREITER, M , PINGOUD, V , WENDE, W , (2005) TYPE II RESTRICTION ENDONUCLEASES: STRUCTURE AND MECHANISM.CELLULAR AND MOLECULAR LIFE SCIENCES. VOL. 62. ISSUE 6. P. 685-707 | 141 | 74% | 275 |
| 8 | LOENEN, WAM , DRYDEN, DTF , RALEIGH, EA , WILSON, GG , MURRAY, NE , (2014) HIGHLIGHTS OF THE DNA CUTTERS: A SHORT HISTORY OF THE RESTRICTION ENZYMES.NUCLEIC ACIDS RESEARCH. VOL. 42. ISSUE 1. P. 3 -19 | 121 | 70% | 39 |
| 9 | PINGOUD, A , JELTSCH, A , (2001) STRUCTURE AND FUNCTION OF TYPE II RESTRICTION ENDONUCLEASES.NUCLEIC ACIDS RESEARCH. VOL. 29. ISSUE 18. P. 3705-3727 | 135 | 77% | 357 |
| 10 | RAO, DN , SAHA, S , KRISHNAMURTHY, V , (2000) ATP-DEPENDENT RESTRICTION ENZYMES.PROGRESS IN NUCLEIC ACID RESEARCH AND MOLECULAR BIOLOGY, VOL 64. VOL. 64. ISSUE . P. 1 -63 | 168 | 88% | 27 |
Classes with closest relation at Level 2 |