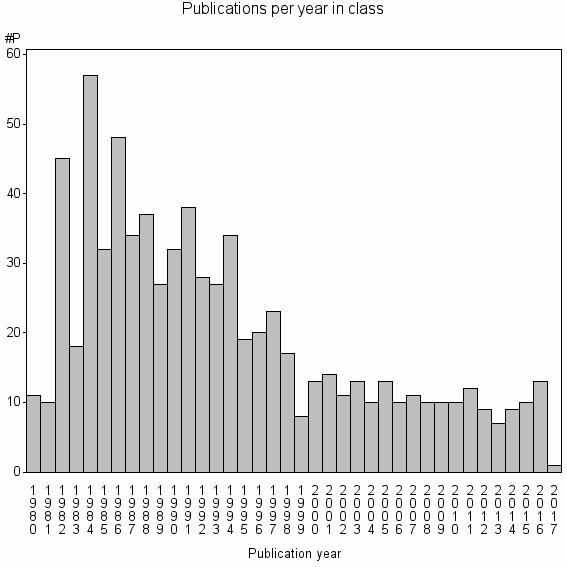
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 3958 | 751 | 17.0 | 50% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 3958 | 2 | COMPUTER APPLICATIONS IN THE BIOSCIENCES//ADAPTIVE WEIGHTED LEAST SQUARES//AMBIGRAM | 751 |
| 21589 | 1 | COMPUTER APPLICATIONS IN THE BIOSCIENCES//ADAPTIVE WEIGHTED LEAST SQUARES//AMBIGRAM | 413 |
| 29833 | 1 | FACTORISATIO NUMERORUM//IMPROPER EDGE//ORDERED FACTORIZATIONS | 179 |
| 35525 | 1 | ANCIENT GREEN COLOR YARN//BLUE COLORANT//CHINESE BROCADE | 96 |
| 37241 | 1 | LANE SEGMENTATION//BAND DETECTION//1D GEL ELECTROPHORESIS ANALYSIS | 63 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | COMPUTER APPLICATIONS IN THE BIOSCIENCES | journal | 188857 | 9% | 7% | 65 |
| 2 | ADAPTIVE WEIGHTED LEAST SQUARES | authKW | 81316 | 0% | 100% | 2 |
| 3 | AMBIGRAM | authKW | 81316 | 0% | 100% | 2 |
| 4 | FACTORISATIO NUMERORUM | authKW | 81316 | 0% | 100% | 2 |
| 5 | IMPROPER EDGE | authKW | 81316 | 0% | 100% | 2 |
| 6 | LANE SEGMENTATION | authKW | 81316 | 0% | 100% | 2 |
| 7 | ORDERED FACTORIZATIONS | authKW | 81316 | 0% | 100% | 2 |
| 8 | HOOK LENGTH FORMULAS | authKW | 73182 | 0% | 60% | 3 |
| 9 | BAND DETECTION | authKW | 54210 | 0% | 67% | 2 |
| 10 | PARTIAL DIGEST | authKW | 54210 | 0% | 67% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Computer Science, Interdisciplinary Applications | 1637 | 13% | 0% | 98 |
| 2 | Biochemical Research Methods | 1321 | 15% | 0% | 110 |
| 3 | Biochemistry & Molecular Biology | 1251 | 36% | 0% | 269 |
| 4 | Mathematics | 466 | 12% | 0% | 91 |
| 5 | Mathematical & Computational Biology | 427 | 5% | 0% | 35 |
| 6 | Medical Informatics | 223 | 2% | 0% | 17 |
| 7 | Computer Science, Theory & Methods | 159 | 5% | 0% | 35 |
| 8 | Biotechnology & Applied Microbiology | 96 | 6% | 0% | 47 |
| 9 | Chemistry, Analytical | 92 | 6% | 0% | 47 |
| 10 | Engineering, Biomedical | 74 | 3% | 0% | 23 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CENTRALIZED STERILE SERV | 40658 | 0% | 100% | 1 |
| 2 | DBVBAA | 40658 | 0% | 100% | 1 |
| 3 | DIRECC PROYECTOS INVEST | 40658 | 0% | 100% | 1 |
| 4 | DISCRETE SIMULAT SCI CCS5 | 40658 | 0% | 100% | 1 |
| 5 | HOSP INFORMAT NUMER IMAGING GRP | 40658 | 0% | 100% | 1 |
| 6 | IISE | 40658 | 0% | 100% | 1 |
| 7 | MICROBIAL POPULAT DYNAM | 40658 | 0% | 100% | 1 |
| 8 | QUAL IMPROVEMENT AREA | 40658 | 0% | 100% | 1 |
| 9 | SECT MICROBIOL AGROALIMENTARE AMBIENTALE | 40658 | 0% | 100% | 1 |
| 10 | SPACE ORGANIZAT | 40658 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | COMPUTER APPLICATIONS IN THE BIOSCIENCES | 188857 | 9% | 7% | 65 |
| 2 | NUCLEIC ACIDS RESEARCH | 16871 | 17% | 0% | 128 |
| 3 | BIOTECHNIQUES | 12367 | 6% | 1% | 43 |
| 4 | COMPUTER PROGRAMS IN BIOMEDICINE | 6523 | 1% | 3% | 6 |
| 5 | BIOCHEMICAL EDUCATION | 6415 | 1% | 2% | 9 |
| 6 | AUSTRALASIAN BIOTECHNOLOGY | 2969 | 0% | 2% | 3 |
| 7 | FIBONACCI QUARTERLY | 2263 | 1% | 1% | 8 |
| 8 | AMERICAN MATHEMATICAL MONTHLY | 1802 | 1% | 0% | 11 |
| 9 | JOURNAL OF BIOINFORMATICS AND COMPUTATIONAL BIOLOGY | 1770 | 1% | 1% | 4 |
| 10 | GENE ANALYSIS TECHNIQUES | 1673 | 0% | 2% | 2 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ADAPTIVE WEIGHTED LEAST SQUARES | 81316 | 0% | 100% | 2 | Search ADAPTIVE+WEIGHTED+LEAST+SQUARES | Search ADAPTIVE+WEIGHTED+LEAST+SQUARES |
| 2 | AMBIGRAM | 81316 | 0% | 100% | 2 | Search AMBIGRAM | Search AMBIGRAM |
| 3 | FACTORISATIO NUMERORUM | 81316 | 0% | 100% | 2 | Search FACTORISATIO+NUMERORUM | Search FACTORISATIO+NUMERORUM |
| 4 | IMPROPER EDGE | 81316 | 0% | 100% | 2 | Search IMPROPER+EDGE | Search IMPROPER+EDGE |
| 5 | LANE SEGMENTATION | 81316 | 0% | 100% | 2 | Search LANE+SEGMENTATION | Search LANE+SEGMENTATION |
| 6 | ORDERED FACTORIZATIONS | 81316 | 0% | 100% | 2 | Search ORDERED+FACTORIZATIONS | Search ORDERED+FACTORIZATIONS |
| 7 | HOOK LENGTH FORMULAS | 73182 | 0% | 60% | 3 | Search HOOK+LENGTH+FORMULAS | Search HOOK+LENGTH+FORMULAS |
| 8 | BAND DETECTION | 54210 | 0% | 67% | 2 | Search BAND+DETECTION | Search BAND+DETECTION |
| 9 | PARTIAL DIGEST | 54210 | 0% | 67% | 2 | Search PARTIAL+DIGEST | Search PARTIAL+DIGEST |
| 10 | TURNPIKE PROBLEM | 54210 | 0% | 67% | 2 | Search TURNPIKE+PROBLEM | Search TURNPIKE+PROBLEM |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | HOU, QH , (2016) AN INSERTION ALGORITHM AND LEADERS OF ROOTED TREES.EUROPEAN JOURNAL OF COMBINATORICS. VOL. 53. ISSUE . P. 35 -44 | 10 | 91% | 1 |
| 2 | BISHOP, MJ , (1984) SOFTWARE FOR MOLECULAR-BIOLOGY .3. GENERAL-ANALYSIS PROGRAMS CLASSIFIED ACCORDING TO THE COMPUTER ON WHICH THEY RUN.BIOESSAYS. VOL. 1. ISSUE 3. P. 126-129 | 42 | 76% | 0 |
| 3 | FOMIN, E , (2016) A SIMPLE APPROACH TO THE RECONSTRUCTION OF A SET OF POINTS FROM THE MULTISET OF N(2) PAIRWISE DISTANCES IN N(2) STEPS FOR THE SEQUENCING PROBLEM: II. ALGORITHM.JOURNAL OF COMPUTATIONAL BIOLOGY. VOL. 23. ISSUE 12. P. 934 -942 | 18 | 53% | 0 |
| 4 | KORN, LJ , QUEEN, C , (1984) ANALYSIS OF BIOLOGICAL SEQUENCES ON SMALL COMPUTERS.DNA-A JOURNAL OF MOLECULAR & CELLULAR BIOLOGY. VOL. 3. ISSUE 6. P. 421-436 | 51 | 58% | 30 |
| 5 | PEVZNER, PA , (1994) COMBINATORIAL METHODS FOR DNA PHYSICAL MAPPING.IEEE ENGINEERING IN MEDICINE AND BIOLOGY MAGAZINE. VOL. 13. ISSUE 1. P. 146-& | 16 | 100% | 1 |
| 6 | PEVZNER, PA , (1995) DNA PHYSICAL MAPPING AND ALTERNATING EULERIAN CYCLES IN COLORED GRAPHS.ALGORITHMICA. VOL. 13. ISSUE 1-2. P. 77-105 | 17 | 81% | 51 |
| 7 | WRIGHT, LW , LICHTER, JB , REINITZ, J , SHIFMAN, MA , KIDD, KK , MILLER, PL , (1994) COMPUTER-ASSISTED RESTRICTION MAPPING - AN INTEGRATED APPROACH TO HANDLING EXPERIMENTAL UNCERTAINTY.COMPUTER APPLICATIONS IN THE BIOSCIENCES. VOL. 10. ISSUE 4. P. 435-442 | 14 | 100% | 6 |
| 8 | ABBAS, MM , BAHIG, HM , (2016) A FAST EXACT SEQUENTIAL ALGORITHM FOR THE PARTIAL DIGEST PROBLEM.BMC BIOINFORMATICS. VOL. 17. ISSUE . P. - | 9 | 75% | 1 |
| 9 | CIELIEBAK, M , EIDENBENZ, S , PENNA, P , (2005) PARTIAL DIGEST IS HARD TO SOLVE FOR ERRONEOUS INPUT DATA.THEORETICAL COMPUTER SCIENCE. VOL. 349. ISSUE 3. P. 361-381 | 11 | 85% | 5 |
| 10 | INGLEHART, J , NELSON, PC , (1994) ON THE LIMITATIONS OF AUTOMATED RESTRICTION MAPPING.COMPUTER APPLICATIONS IN THE BIOSCIENCES. VOL. 10. ISSUE 3. P. 249-261 | 13 | 100% | 8 |
Classes with closest relation at Level 2 |