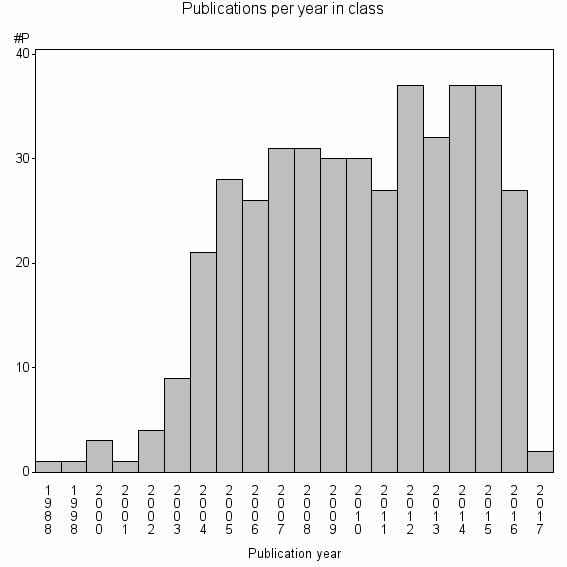
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 21530 | 415 | 41.7 | 77% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 548 | 2 | BMC SYSTEMS BIOLOGY//MATHEMATICAL & COMPUTATIONAL BIOLOGY//BOOLEAN NETWORKS | 14655 |
| 21530 | 1 | NETWORK MOTIFS//SUBGRAPH SAMPLING//PROGRAMA GENOM COMPUTAC | 415 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | NETWORK MOTIFS | authKW | 1177189 | 12% | 33% | 48 |
| 2 | SUBGRAPH SAMPLING | authKW | 165550 | 1% | 75% | 3 |
| 3 | PROGRAMA GENOM COMPUTAC | address | 162368 | 2% | 28% | 8 |
| 4 | GRAPHLET COUNTING | authKW | 147157 | 0% | 100% | 2 |
| 5 | NETWORK MOTIF DETECTION ALGORITHM | authKW | 147157 | 0% | 100% | 2 |
| 6 | REGULATORY NETWORK MOTIFS | authKW | 147157 | 0% | 100% | 2 |
| 7 | SPECTRAL PLOT | authKW | 147157 | 0% | 100% | 2 |
| 8 | STRONG WEAK TIE LINK | authKW | 147157 | 0% | 100% | 2 |
| 9 | FEED FORWARD LOOP | authKW | 147145 | 2% | 25% | 8 |
| 10 | PROGRAMA BIOL MOL COMPUTAC | address | 110364 | 1% | 50% | 3 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 8777 | 27% | 0% | 111 |
| 2 | Biochemical Research Methods | 659 | 14% | 0% | 58 |
| 3 | Biotechnology & Applied Microbiology | 351 | 13% | 0% | 56 |
| 4 | Biology | 339 | 9% | 0% | 36 |
| 5 | Biochemistry & Molecular Biology | 302 | 26% | 0% | 106 |
| 6 | Genetics & Heredity | 283 | 12% | 0% | 50 |
| 7 | Physics, Mathematical | 250 | 7% | 0% | 31 |
| 8 | Physics, Fluids & Plasmas | 176 | 6% | 0% | 23 |
| 9 | Multidisciplinary Sciences | 121 | 3% | 0% | 13 |
| 10 | Computer Science, Information Systems | 87 | 4% | 0% | 18 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | PROGRAMA GENOM COMPUTAC | 162368 | 2% | 28% | 8 |
| 2 | PROGRAMA BIOL MOL COMPUTAC | 110364 | 1% | 50% | 3 |
| 3 | PROGRAM COMPUTAT GENOM | 94591 | 1% | 21% | 6 |
| 4 | CIFN | 90552 | 1% | 31% | 4 |
| 5 | AD T SYST BIOCOMPUTAT GRP | 73578 | 0% | 100% | 1 |
| 6 | ANAL GENOM EVOLUT FUNCT | 73578 | 0% | 100% | 1 |
| 7 | BIONANOTECNOL NANOCIENCIAS NANOTECNOL | 73578 | 0% | 100% | 1 |
| 8 | C ITALIO NACL | 73578 | 0% | 100% | 1 |
| 9 | CATALANA RECERCA ESTUDIS AVANCATS COMPLEX | 73578 | 0% | 100% | 1 |
| 10 | CHANGZHOU BIOMED INFORMAT TECHNOL | 73578 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BMC SYSTEMS BIOLOGY | 20213 | 5% | 1% | 21 |
| 2 | MOLECULAR BIOSYSTEMS | 6950 | 4% | 1% | 16 |
| 3 | IET SYSTEMS BIOLOGY | 1989 | 1% | 1% | 3 |
| 4 | BMC BIOINFORMATICS | 1637 | 3% | 0% | 13 |
| 5 | MOLECULAR SYSTEMS BIOLOGY | 1549 | 1% | 1% | 4 |
| 6 | NANO COMMUNICATION NETWORKS | 1413 | 0% | 2% | 1 |
| 7 | BIOSYSTEMS | 1321 | 2% | 0% | 7 |
| 8 | IEEE-ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS | 1034 | 1% | 0% | 4 |
| 9 | DATABASE-THE JOURNAL OF BIOLOGICAL DATABASES AND CURATION | 1011 | 1% | 0% | 3 |
| 10 | PLOS COMPUTATIONAL BIOLOGY | 1005 | 2% | 0% | 8 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | NETWORK MOTIFS | 1177189 | 12% | 33% | 48 | Search NETWORK+MOTIFS | Search NETWORK+MOTIFS |
| 2 | SUBGRAPH SAMPLING | 165550 | 1% | 75% | 3 | Search SUBGRAPH+SAMPLING | Search SUBGRAPH+SAMPLING |
| 3 | GRAPHLET COUNTING | 147157 | 0% | 100% | 2 | Search GRAPHLET+COUNTING | Search GRAPHLET+COUNTING |
| 4 | NETWORK MOTIF DETECTION ALGORITHM | 147157 | 0% | 100% | 2 | Search NETWORK+MOTIF+DETECTION+ALGORITHM | Search NETWORK+MOTIF+DETECTION+ALGORITHM |
| 5 | REGULATORY NETWORK MOTIFS | 147157 | 0% | 100% | 2 | Search REGULATORY+NETWORK+MOTIFS | Search REGULATORY+NETWORK+MOTIFS |
| 6 | SPECTRAL PLOT | 147157 | 0% | 100% | 2 | Search SPECTRAL+PLOT | Search SPECTRAL+PLOT |
| 7 | STRONG WEAK TIE LINK | 147157 | 0% | 100% | 2 | Search STRONG+WEAK+TIE+LINK | Search STRONG+WEAK+TIE+LINK |
| 8 | FEED FORWARD LOOP | 147145 | 2% | 25% | 8 | Search FEED+FORWARD+LOOP | Search FEED+FORWARD+LOOP |
| 9 | ASSOCIATED MATRIX | 98103 | 0% | 67% | 2 | Search ASSOCIATED+MATRIX | Search ASSOCIATED+MATRIX |
| 10 | NETWORK ORIENTED APPROACH | 98103 | 0% | 67% | 2 | Search NETWORK+ORIENTED+APPROACH | Search NETWORK+ORIENTED+APPROACH |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | TRAN, NTL , MOHAN, S , XU, ZQ , HUANG, CH , (2015) CURRENT INNOVATIONS AND FUTURE CHALLENGES OF NETWORK MOTIF DETECTION.BRIEFINGS IN BIOINFORMATICS. VOL. 16. ISSUE 3. P. 497 -525 | 25 | 61% | 2 |
| 2 | ALON, U , (2007) NETWORK MOTIFS: THEORY AND EXPERIMENTAL APPROACHES.NATURE REVIEWS GENETICS. VOL. 8. ISSUE 6. P. 450 -461 | 24 | 27% | 1152 |
| 3 | WONG, E , BAUR, B , QUADER, S , HUANG, CH , (2012) BIOLOGICAL NETWORK MOTIF DETECTION: PRINCIPLES AND PRACTICE.BRIEFINGS IN BIOINFORMATICS. VOL. 13. ISSUE 2. P. 202 -215 | 20 | 65% | 21 |
| 4 | PRATAP, A , TALIYAN, S , SINGH, TR , (2014) NMDB: NETWORK MOTIF DATABASE ENVISAGED AND EXPLICATED FROM HUMAN DISEASE SPECIFIC PATHWAYS.JOURNAL OF BIOLOGICAL SYSTEMS. VOL. 22. ISSUE 1. P. 89-100 | 12 | 92% | 0 |
| 5 | LI, X , STONES, DS , WANG, HD , DENG, HL , LIU, XG , WANG, G , (2012) NETMODE: NETWORK MOTIF DETECTION WITHOUT NAUTY.PLOS ONE. VOL. 7. ISSUE 12. P. - | 13 | 81% | 9 |
| 6 | WANG, JX , HUANG, YN , WU, FX , PAN, Y , (2012) SYMMETRY COMPRESSION METHOD FOR DISCOVERING NETWORK MOTIFS.IEEE-ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS. VOL. 9. ISSUE 6. P. 1776 -1789 | 14 | 74% | 2 |
| 7 | FRETTER, C , MULLER-HANNEMANN, M , HUTT, MT , (2012) SUBGRAPH FLUCTUATIONS IN RANDOM GRAPHS.PHYSICAL REVIEW E. VOL. 85. ISSUE 5. P. - | 16 | 64% | 2 |
| 8 | CAMAS, FM , POYATOS, JF , (2008) WHAT DETERMINES THE ASSEMBLY OF TRANSCRIPTIONAL NETWORK MOTIFS IN ESCHERICHIA COLI?.PLOS ONE. VOL. 3. ISSUE 11. P. - | 21 | 50% | 9 |
| 9 | WIDDER, S , SOLE, R , MACIA, J , (2012) EVOLVABILITY OF FEED-FORWARD LOOP ARCHITECTURE BIASES ITS ABUNDANCE IN TRANSCRIPTION NETWORKS.BMC SYSTEMS BIOLOGY. VOL. 6. ISSUE . P. - | 16 | 59% | 3 |
| 10 | MASOUDI-NEJAD, A , SCHREIBER, F , KASHANI, ZRM , (2012) BUILDING BLOCKS OF BIOLOGICAL NETWORKS: A REVIEW ON MAJOR NETWORK MOTIF DISCOVERY ALGORITHMS.IET SYSTEMS BIOLOGY. VOL. 6. ISSUE 5. P. 164-174 | 13 | 68% | 17 |
Classes with closest relation at Level 1 |