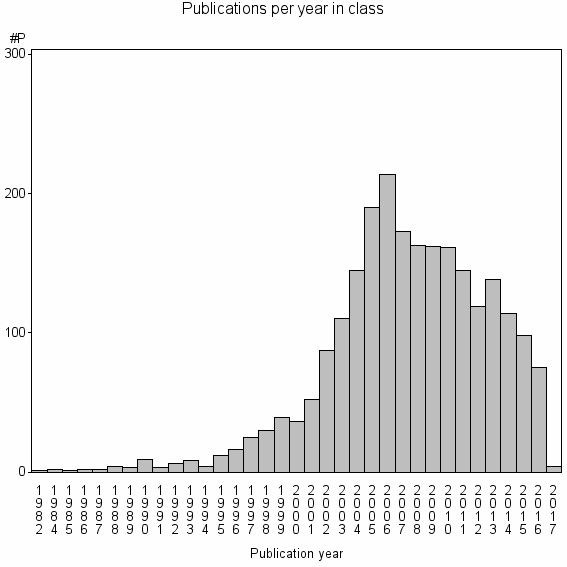
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 2111 | 2353 | 40.0 | 81% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 126 | 2 | MOLECULAR BIOLOGY AND EVOLUTION//BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY | 23963 |
| 2111 | 1 | MOTIF DISCOVERY//TRANSCRIPTION FACTOR BINDING SITE//MOTIF FINDING | 2353 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | MOTIF DISCOVERY | authKW | 419446 | 2% | 58% | 56 |
| 2 | TRANSCRIPTION FACTOR BINDING SITE | authKW | 364118 | 3% | 35% | 80 |
| 3 | MOTIF FINDING | authKW | 184693 | 1% | 65% | 22 |
| 4 | POSITION WEIGHT MATRIX | authKW | 118619 | 1% | 57% | 16 |
| 5 | BIOINFORMATICS | journal | 102432 | 12% | 3% | 290 |
| 6 | BIOINFORMAT GENOMES EAUX BIGRE | address | 90814 | 1% | 50% | 14 |
| 7 | UKG | address | 87579 | 0% | 75% | 9 |
| 8 | DNA MOTIFS | authKW | 75601 | 1% | 45% | 13 |
| 9 | PROMOTER PREDICTION | authKW | 73080 | 1% | 43% | 13 |
| 10 | MATHEMATICAL & COMPUTATIONAL BIOLOGY | WoSSC | 69249 | 31% | 1% | 741 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 69249 | 31% | 1% | 741 |
| 2 | Biotechnology & Applied Microbiology | 20901 | 40% | 0% | 942 |
| 3 | Biochemical Research Methods | 20799 | 32% | 0% | 742 |
| 4 | Genetics & Heredity | 5334 | 21% | 0% | 486 |
| 5 | Biochemistry & Molecular Biology | 4657 | 39% | 0% | 907 |
| 6 | Computer Science, Interdisciplinary Applications | 1183 | 7% | 0% | 156 |
| 7 | Statistics & Probability | 631 | 5% | 0% | 106 |
| 8 | Computer Science, Artificial Intelligence | 312 | 4% | 0% | 85 |
| 9 | Computer Science, Theory & Methods | 169 | 3% | 0% | 71 |
| 10 | Biology | 133 | 3% | 0% | 67 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOINFORMAT GENOMES EAUX BIGRE | 90814 | 1% | 50% | 14 |
| 2 | UKG | 87579 | 0% | 75% | 9 |
| 3 | BIOINFORMAT INTELLIGENT COMP | 46338 | 0% | 71% | 5 |
| 4 | THEORET GENET | 43697 | 0% | 42% | 8 |
| 5 | KNOWLEDGE EXTRACT | 42379 | 0% | 47% | 7 |
| 6 | HARVARD MIT HLTH SCI TECHNOL HST | 41512 | 0% | 40% | 8 |
| 7 | GRP ALGORISM GENET | 38926 | 0% | 100% | 3 |
| 8 | ARCO GRP | 31134 | 0% | 40% | 6 |
| 9 | STUDIES PHYS BIOL | 29337 | 1% | 9% | 25 |
| 10 | EEBI COMP | 25951 | 0% | 100% | 2 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOINFORMATICS | 102432 | 12% | 3% | 290 |
| 2 | BMC BIOINFORMATICS | 54644 | 8% | 2% | 178 |
| 3 | NUCLEIC ACIDS RESEARCH | 52317 | 17% | 1% | 399 |
| 4 | BIOTECHFORUM ( BTF ) | 37001 | 4% | 3% | 86 |
| 5 | JOURNAL OF COMPUTATIONAL BIOLOGY | 19108 | 2% | 3% | 48 |
| 6 | GENOME BIOLOGY | 14049 | 2% | 2% | 57 |
| 7 | ALGORITHMS FOR MOLECULAR BIOLOGY | 13219 | 1% | 6% | 17 |
| 8 | BMC GENOMICS | 10147 | 4% | 1% | 87 |
| 9 | COMPUTER APPLICATIONS IN THE BIOSCIENCES | 9599 | 1% | 3% | 26 |
| 10 | IEEE-ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS | 7128 | 1% | 2% | 25 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | GUHATHAKURTA, D , (2006) COMPUTATIONAL IDENTIFICATION OF TRANSCRIPTIONAL REGULATORY ELEMENTS IN DNA SEQUENCE.NUCLEIC ACIDS RESEARCH. VOL. 34. ISSUE 12. P. 3585 -3598 | 110 | 72% | 64 |
| 2 | SANDVE, GK , DRABLOS, F , (2006) A SURVEY OF MOTIF DISCOVERY METHODS IN AN INTEGRATED FRAMEWORK.BIOLOGY DIRECT. VOL. 1. ISSUE . P. - | 74 | 90% | 61 |
| 3 | MACISAAC, KD , FRAENKEL, E , (2006) PRACTICAL STRATEGIES FOR DISCOVERING REGULATORY DNA SEQUENCE MOTIFS.PLOS COMPUTATIONAL BIOLOGY. VOL. 2. ISSUE 4. P. 201 -210 | 76 | 81% | 79 |
| 4 | DAI, HK , DAS, MK , (2007) A SURVEY OF DNA MOTIF FINDING ALGORITHMS.BMC BIOINFORMATICS. VOL. 8. ISSUE . P. - | 61 | 86% | 155 |
| 5 | ZAMBELLI, F , PESOLE, G , PAVESI, G , (2013) MOTIF DISCOVERY AND TRANSCRIPTION FACTOR BINDING SITES BEFORE AND AFTER THE NEXT-GENERATION SEQUENCING ERA.BRIEFINGS IN BIOINFORMATICS. VOL. 14. ISSUE 2. P. 225-237 | 71 | 72% | 22 |
| 6 | LIHU, A , HOLBAN, S , (2015) A REVIEW OF ENSEMBLE METHODS FOR DE NOVO MOTIF DISCOVERY IN CHIP-SEQ DATA.BRIEFINGS IN BIOINFORMATICS. VOL. 16. ISSUE 6. P. 964 -973 | 61 | 80% | 4 |
| 7 | AERTS, S , (2012) COMPUTATIONAL STRATEGIES FOR THE GENOME-WIDE IDENTIFICATION OF CIS-REGULATORY ELEMENTS AND TRANSCRIPTIONAL TARGETS.TRANSCRIPTIONAL SWITCHES DURING DEVELOPMENT. VOL. 98. ISSUE . P. 121 -145 | 80 | 65% | 10 |
| 8 | VINGRON, M , BRAZMA, A , COULSON, R , VAN HELDEN, J , MANKE, T , PALIN, K , SAND, O , UKKONEN, E , (2009) INTEGRATING SEQUENCE, EVOLUTION AND FUNCTIONAL GENOMICS IN REGULATORY GENOMICS.GENOME BIOLOGY. VOL. 10. ISSUE 1. P. - | 71 | 78% | 13 |
| 9 | ABNIZOVA, I , SUBHANKULOVA, T , GILKS, WR , (2007) RECENT COMPUTATIONAL APPROACHES TO UNDERSTAND GENE REGULATION: MINING GENE REGULATION IN SILICO.CURRENT GENOMICS. VOL. 8. ISSUE 2. P. 79 -91 | 92 | 58% | 1 |
| 10 | VAN LOO, P , MARYNEN, P , (2009) COMPUTATIONAL METHODS FOR THE DETECTION OF CIS-REGULATORY MODULES.BRIEFINGS IN BIOINFORMATICS. VOL. 10. ISSUE 5. P. 509 -524 | 46 | 92% | 36 |
Classes with closest relation at Level 1 |