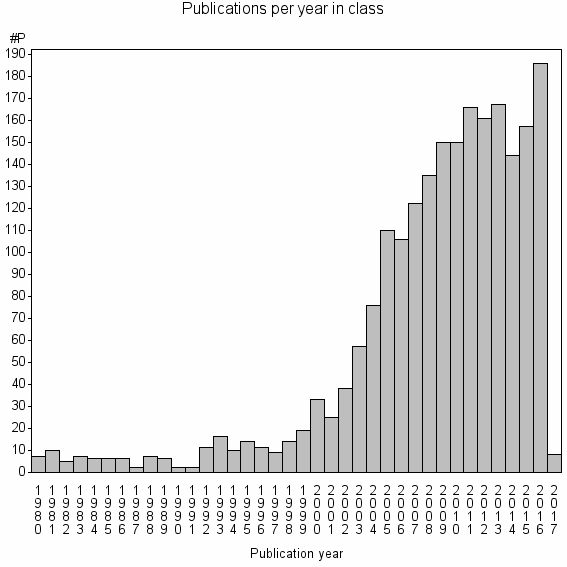
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 2741 | 2161 | 50.1 | 79% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 219 | 3 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN FOLDING//PROTEIN STRUCTURE PREDICTION | 50409 |
| 365 | 2 | PROTEIN STRUCTURE PREDICTION//PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PSEUDO AMINO ACID COMPOSITION | 17331 |
| 2741 | 1 | PSEUDO AMINO ACID COMPOSITION//PROTEIN STRUCTURAL CLASS//JACKKNIFE TEST | 2161 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | PSEUDO AMINO ACID COMPOSITION | authKW | 1656956 | 6% | 95% | 123 |
| 2 | PROTEIN STRUCTURAL CLASS | authKW | 516072 | 2% | 83% | 44 |
| 3 | JACKKNIFE TEST | authKW | 429794 | 2% | 89% | 34 |
| 4 | COMPUTAT MUTAT PROJECT | address | 427379 | 2% | 92% | 33 |
| 5 | PROTEIN SUBCELLULAR LOCALIZATION | authKW | 357289 | 1% | 82% | 31 |
| 6 | PSEAAC | authKW | 282568 | 1% | 100% | 20 |
| 7 | COVARIANT DISCRIMINANT ALGORITHM | authKW | 240183 | 1% | 100% | 17 |
| 8 | INCREMENT OF DIVERSITY | authKW | 232498 | 1% | 69% | 24 |
| 9 | NETWORK ORIENTED INTELLIGENT COMPUTAT | address | 215483 | 1% | 49% | 31 |
| 10 | NEAREST NEIGHBOR ALGORITHM | authKW | 214552 | 1% | 56% | 27 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 47158 | 27% | 1% | 587 |
| 2 | Biochemical Research Methods | 7813 | 21% | 0% | 444 |
| 3 | Biochemistry & Molecular Biology | 5766 | 44% | 0% | 948 |
| 4 | Biology | 4286 | 13% | 0% | 283 |
| 5 | Biotechnology & Applied Microbiology | 2755 | 16% | 0% | 350 |
| 6 | Biophysics | 1295 | 10% | 0% | 212 |
| 7 | Computer Science, Interdisciplinary Applications | 1186 | 7% | 0% | 149 |
| 8 | Computer Science, Artificial Intelligence | 604 | 5% | 0% | 107 |
| 9 | Chemistry, Medicinal | 318 | 4% | 0% | 94 |
| 10 | Mathematics, Interdisciplinary Applications | 102 | 2% | 0% | 44 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | COMPUTAT MUTAT PROJECT | 427379 | 2% | 92% | 33 |
| 2 | NETWORK ORIENTED INTELLIGENT COMPUTAT | 215483 | 1% | 49% | 31 |
| 3 | NEUROINFORMATMINIST EDUC | 180177 | 1% | 49% | 26 |
| 4 | GENOM COMPUTAT BIOL | 146140 | 2% | 21% | 49 |
| 5 | COMP AIDED DRUG DISCOVERY | 135231 | 1% | 32% | 30 |
| 6 | CEGMR | 130973 | 2% | 24% | 39 |
| 7 | ENGN NONFOOD BIOREFINERY | 75459 | 1% | 23% | 23 |
| 8 | UPJOHN S | 71219 | 1% | 46% | 11 |
| 9 | THEORET BIOPHYS | 70958 | 1% | 18% | 28 |
| 10 | BIOINFORMAT DRUG DISCOVERY | 64580 | 0% | 57% | 8 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | PROTEIN AND PEPTIDE LETTERS | 161007 | 8% | 7% | 170 |
| 2 | JOURNAL OF THEORETICAL BIOLOGY | 55999 | 10% | 2% | 207 |
| 3 | CURRENT BIOINFORMATICS | 28712 | 1% | 7% | 30 |
| 4 | AMINO ACIDS | 28532 | 4% | 2% | 85 |
| 5 | MEDICINAL CHEMISTRY | 17480 | 2% | 4% | 33 |
| 6 | BMC BIOINFORMATICS | 16162 | 4% | 1% | 93 |
| 7 | BIOINFORMATICS | 13672 | 5% | 1% | 102 |
| 8 | COMPUTATIONAL BIOLOGY AND CHEMISTRY | 12605 | 1% | 3% | 28 |
| 9 | CURRENT PROTEOMICS | 8598 | 1% | 5% | 12 |
| 10 | INTERDISCIPLINARY SCIENCES-COMPUTATIONAL LIFE SCIENCES | 6561 | 1% | 4% | 12 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | CHOU, KC , (2015) IMPACTS OF BIOINFORMATICS TO MEDICINAL CHEMISTRY.MEDICINAL CHEMISTRY. VOL. 11. ISSUE 3. P. 218 -234 | 215 | 87% | 81 |
| 2 | CHOU, KC , (2011) SOME REMARKS ON PROTEIN ATTRIBUTE PREDICTION AND PSEUDO AMINO ACID COMPOSITION.JOURNAL OF THEORETICAL BIOLOGY. VOL. 273. ISSUE 1. P. 236 -247 | 174 | 88% | 446 |
| 3 | CHEN, W , LIN, H , CHOU, KC , (2015) PSEUDO NUCLEOTIDE COMPOSITION OR PSEKNC: AN EFFECTIVE FORMULATION FOR ANALYZING GENOMIC SEQUENCES.MOLECULAR BIOSYSTEMS. VOL. 11. ISSUE 10. P. 2620 -2634 | 173 | 93% | 28 |
| 4 | CHEN, W , DING, H , FENG, PM , LIN, H , CHOU, KC , (2016) IACP: A SEQUENCE-BASED TOOL FOR IDENTIFYING ANTICANCER PEPTIDES.ONCOTARGET. VOL. 7. ISSUE 13. P. 16895 -16909 | 108 | 84% | 16 |
| 5 | XU, Y , CHOU, KC , (2016) RECENT PROGRESS IN PREDICTING POSTTRANSLATIONAL MODIFICATION SITES IN PROTEINS.CURRENT TOPICS IN MEDICINAL CHEMISTRY. VOL. 16. ISSUE 6. P. 591 -603 | 144 | 87% | 4 |
| 6 | JIA, JH , LIU, Z , XIAO, X , LIU, BX , CHOU, KC , (2015) IPPI-ESML: AN ENSEMBLE CLASSIFIER FOR IDENTIFYING THE INTERACTIONS OF PROTEINS BY INCORPORATING THEIR PHYSICOCHEMICAL PROPERTIES AND WAVELET TRANSFORMS INTO PSEAAC.JOURNAL OF THEORETICAL BIOLOGY. VOL. 377. ISSUE . P. 47 -56 | 106 | 80% | 56 |
| 7 | CHOU, KC , (2009) PSEUDO AMINO ACID COMPOSITION AND ITS APPLICATIONS IN BIOINFORMATICS, PROTEOMICS AND SYSTEM BIOLOGY.CURRENT PROTEOMICS. VOL. 6. ISSUE 4. P. 262-274 | 146 | 83% | 160 |
| 8 | FAN, YN , XIAO, X , MIN, JL , CHOU, KC , (2014) INR-DRUG: PREDICTING THE INTERACTION OF DRUGS WITH NUCLEAR RECEPTORS IN CELLULAR NETWORKING.INTERNATIONAL JOURNAL OF MOLECULAR SCIENCES. VOL. 15. ISSUE 3. P. 4915 -4937 | 136 | 86% | 28 |
| 9 | JIA, JH , LIU, Z , XIAO, X , LIU, BX , CHOU, KC , (2016) PSUC-LYS: PREDICT LYSINE SUCCINYLATION SITES IN PROTEINS WITH PSEAAC AND ENSEMBLE RANDOM FOREST APPROACH.JOURNAL OF THEORETICAL BIOLOGY. VOL. 394. ISSUE . P. 223 -230 | 79 | 85% | 17 |
| 10 | QIU, WR , XIAO, X , CHOU, KC , (2014) IRSPOT-TNCPSEAAC: IDENTIFY RECOMBINATION SPOTS WITH TRINUCLEOTIDE COMPOSITION AND PSEUDO AMINO ACID COMPONENTS.INTERNATIONAL JOURNAL OF MOLECULAR SCIENCES. VOL. 15. ISSUE 2. P. 1746 -1766 | 97 | 88% | 79 |
Classes with closest relation at Level 1 |