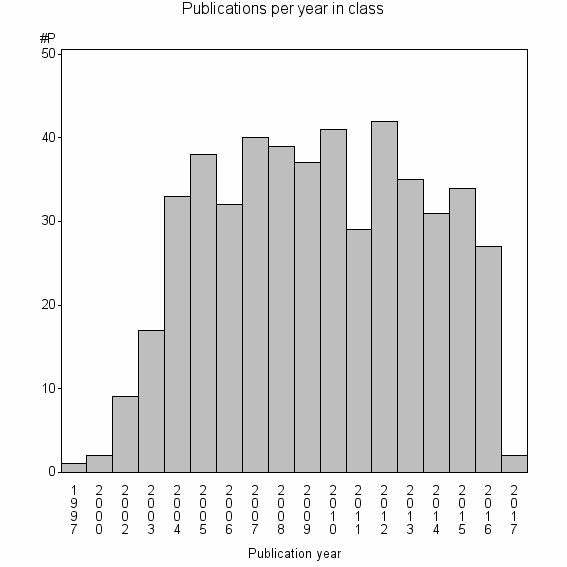
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 19783 | 489 | 51.2 | 87% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 126 | 2 | MOLECULAR BIOLOGY AND EVOLUTION//BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY | 23963 |
| 19783 | 1 | EXPRESSION EVOLUTION//TRANS REGULATION//CIS REGULATORY DIVERGENCE | 489 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | EXPRESSION EVOLUTION | authKW | 339967 | 1% | 78% | 7 |
| 2 | TRANS REGULATION | authKW | 190294 | 2% | 38% | 8 |
| 3 | CIS REGULATORY DIVERGENCE | authKW | 124887 | 0% | 100% | 2 |
| 4 | EXPRESSION CONSERVATION | authKW | 124887 | 0% | 100% | 2 |
| 5 | RHESUS MACAQUE GENOME ARRAY | authKW | 124887 | 0% | 100% | 2 |
| 6 | WE INFORMAT BIOINFORMAT | address | 124887 | 0% | 100% | 2 |
| 7 | CIS REGULATION | authKW | 92694 | 3% | 11% | 14 |
| 8 | EVOLUTION OF GENE EXPRESSION | authKW | 90822 | 1% | 36% | 4 |
| 9 | KRAB ZINC FINGER GENES | authKW | 83257 | 0% | 67% | 2 |
| 10 | EXPRESSION DIVERGENCE | authKW | 70239 | 1% | 19% | 6 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Genetics & Heredity | 10530 | 60% | 0% | 295 |
| 2 | Evolutionary Biology | 4416 | 20% | 0% | 98 |
| 3 | Biotechnology & Applied Microbiology | 1087 | 21% | 0% | 102 |
| 4 | Mathematical & Computational Biology | 650 | 7% | 0% | 34 |
| 5 | Biochemistry & Molecular Biology | 618 | 32% | 0% | 156 |
| 6 | Biochemical Research Methods | 185 | 7% | 0% | 36 |
| 7 | Biology | 184 | 6% | 0% | 30 |
| 8 | Ecology | 145 | 7% | 0% | 35 |
| 9 | Cell Biology | 40 | 7% | 0% | 33 |
| 10 | Developmental Biology | 34 | 2% | 0% | 10 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | WE INFORMAT BIOINFORMAT | 124887 | 0% | 100% | 2 |
| 2 | ARABIDOPSIS STOCK | 62443 | 0% | 100% | 1 |
| 3 | CHAIR SYSTEMAT ZOOL | 62443 | 0% | 100% | 1 |
| 4 | EPHEUMR 5175 | 62443 | 0% | 100% | 1 |
| 5 | EQUIPE RECH 7226 | 62443 | 0% | 100% | 1 |
| 6 | EUROPEAN PUBL HEATH GEN | 62443 | 0% | 100% | 1 |
| 7 | GODDARD S 203 | 62443 | 0% | 100% | 1 |
| 8 | GRP ANS IMMUNOREGULAT 5 | 62443 | 0% | 100% | 1 |
| 9 | HLTH SERV DEV BIOSTAT UNIT | 62443 | 0% | 100% | 1 |
| 10 | IKC | 62443 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOLECULAR BIOLOGY AND EVOLUTION | 13701 | 7% | 1% | 35 |
| 2 | GENOME RESEARCH | 10196 | 3% | 1% | 15 |
| 3 | GENOME BIOLOGY AND EVOLUTION | 9052 | 3% | 1% | 14 |
| 4 | MOLECULAR SYSTEMS BIOLOGY | 5270 | 2% | 1% | 8 |
| 5 | BMC GENOMICS | 4729 | 6% | 0% | 27 |
| 6 | NATURE REVIEWS GENETICS | 3362 | 2% | 1% | 8 |
| 7 | PLOS GENETICS | 3337 | 4% | 0% | 18 |
| 8 | GENOME BIOLOGY | 2997 | 2% | 0% | 12 |
| 9 | GENETICS | 2560 | 5% | 0% | 23 |
| 10 | BIOTECHFORUM ( BTF ) | 1941 | 2% | 0% | 9 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | COOLON, JD , MCMANUS, CJ , STEVENSON, KR , GRAVELEY, BR , WITTKOPP, PJ , (2014) TEMPO AND MODE OF REGULATORY EVOLUTION IN DROSOPHILA.GENOME RESEARCH. VOL. 24. ISSUE 5. P. 797 -808 | 39 | 50% | 20 |
| 2 | MACK, KL , CAMPBELL, P , NACHMAN, MW , (2016) GENE REGULATION AND SPECIATION IN HOUSE MICE.GENOME RESEARCH. VOL. 26. ISSUE 4. P. 451 -461 | 27 | 44% | 4 |
| 3 | THOMPSON, DA , CUBILLOS, FA , (2017) NATURAL GENE EXPRESSION VARIATION STUDIES IN YEAST.YEAST. VOL. 34. ISSUE 1. P. 3 -17 | 44 | 31% | 0 |
| 4 | SOMEL, M , ROHLFS, R , LIU, XL , (2014) TRANSCRIPTOMIC INSIGHTS INTO HUMAN BRAIN EVOLUTION: ACCELERATION, NEUTRALITY, HETEROCHRONY.CURRENT OPINION IN GENETICS & DEVELOPMENT. VOL. 29. ISSUE . P. 110 -119 | 36 | 34% | 6 |
| 5 | MCMANUS, CJ , COOLON, JD , DUFF, MO , EIPPER-MAINS, J , GRAVELEY, BR , WITTKOPP, PJ , (2010) REGULATORY DIVERGENCE IN DROSOPHILA REVEALED BY MRNA-SEQ.GENOME RESEARCH. VOL. 20. ISSUE 6. P. 816 -825 | 21 | 46% | 121 |
| 6 | TIROSH, I , BARKAI, N , (2011) INFERRING REGULATORY MECHANISMS FROM PATTERNS OF EVOLUTIONARY DIVERGENCE.MOLECULAR SYSTEMS BIOLOGY. VOL. 7. ISSUE . P. - | 33 | 35% | 16 |
| 7 | STAUBACH, F , TESCHKE, M , VOOLSTRA, CR , WOLF, JBW , TAUTZ, D , (2010) A TEST OF THE NEUTRAL MODEL OF EXPRESSION CHANGE IN NATURAL POPULATIONS OF HOUSE MOUSE SUBSPECIES.EVOLUTION. VOL. 64. ISSUE 2. P. 549-560 | 22 | 55% | 5 |
| 8 | SUVOROV, A , NOLTE, V , PANDEY, RV , FRANSSEN, SU , FUTSCHIK, A , SCHLOTTERER, C , (2013) INTRA-SPECIFIC REGULATORY VARIATION IN DROSOPHILA PSEUDOOBSCURA.PLOS ONE. VOL. 8. ISSUE 12. P. - | 18 | 58% | 8 |
| 9 | SCHAEFKE, B , EMERSON, JJ , WANG, TY , LU, MYJ , HSIEH, LC , LI, WH , (2013) INHERITANCE OF GENE EXPRESSION LEVEL AND SELECTIVE CONSTRAINTS ON TRANS- AND CIS-REGULATORY CHANGES IN YEAST.MOLECULAR BIOLOGY AND EVOLUTION. VOL. 30. ISSUE 9. P. 2121 -2133 | 21 | 45% | 21 |
| 10 | WARNEFORS, M , EYRE-WALKER, A , (2012) A SELECTION INDEX FOR GENE EXPRESSION EVOLUTION AND ITS APPLICATION TO THE DIVERGENCE BETWEEN HUMANS AND CHIMPANZEES.PLOS ONE. VOL. 7. ISSUE 4. P. - | 25 | 41% | 6 |
Classes with closest relation at Level 1 |