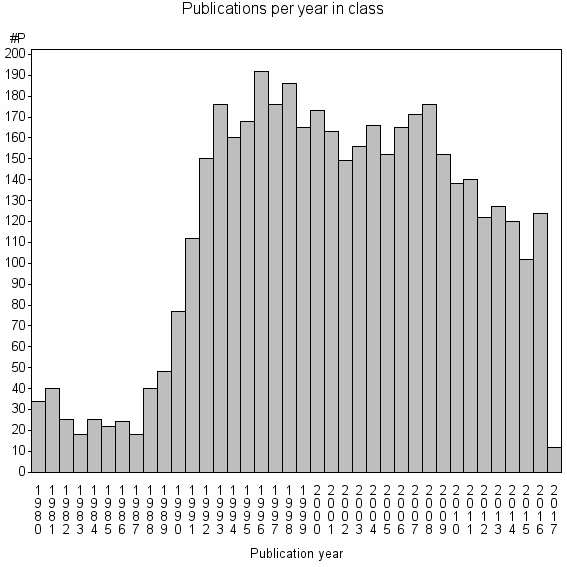
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 2356 | 4364 | 42.3 | 81% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 7 | 4 | INFECTIOUS DISEASES//MICROBIOLOGY//VIROLOGY | 1353914 |
| 29 | 3 | HIV//VIROLOGY//HIV 1 | 108954 |
| 2356 | 2 | NNRTIS//REVERSE TRANSCRIPTASE//HIV DRUG ISTANCE PROGRAM | 4364 |
| 1509 | 1 | NNRTIS//REVERSE TRANSCRIPTASE//HIV 1 REVERSE TRANSCRIPTASE | 2619 |
| 5490 | 1 | NCP7//RNA DIMERIZATION//NUCLEOCAPSID PROTEIN | 1617 |
| 33146 | 1 | 3 METHYL 1 6 METHYL 2 METHYLSULFANYLPYRIMIDIN 4 YL 4 PHENYLIMINOMETHYL 1H PYRAZOL 5 OL//5 ALKYL OR ARYLTHIO 6 VINYL URACILS//5 BENZYLPYRIMIDIN 2 4 6 TRIONES | 128 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | NNRTIS | authKW | 794278 | 5% | 53% | 215 |
| 2 | REVERSE TRANSCRIPTASE | authKW | 599364 | 8% | 25% | 346 |
| 3 | HIV DRUG ISTANCE PROGRAM | address | 360101 | 4% | 30% | 173 |
| 4 | HIV 1 REVERSE TRANSCRIPTASE | authKW | 267316 | 2% | 45% | 85 |
| 5 | NCP7 | authKW | 132151 | 1% | 82% | 23 |
| 6 | RNA DIMERIZATION | authKW | 128532 | 0% | 88% | 21 |
| 7 | NON NUCLEOSIDE REVERSE TRANSCRIPTASE INHIBITORS | authKW | 128257 | 2% | 27% | 68 |
| 8 | NUCLEOCAPSID PROTEIN | authKW | 127497 | 2% | 24% | 77 |
| 9 | HIV 1 | authKW | 123860 | 9% | 5% | 385 |
| 10 | REGA MED | address | 122213 | 5% | 8% | 232 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Virology | 51455 | 26% | 1% | 1146 |
| 2 | Chemistry, Medicinal | 11354 | 16% | 0% | 681 |
| 3 | Biochemistry & Molecular Biology | 10804 | 42% | 0% | 1853 |
| 4 | Chemistry, Organic | 1189 | 9% | 0% | 393 |
| 5 | Pharmacology & Pharmacy | 866 | 11% | 0% | 468 |
| 6 | Biophysics | 593 | 5% | 0% | 232 |
| 7 | Infectious Diseases | 249 | 3% | 0% | 133 |
| 8 | Biochemical Research Methods | 120 | 3% | 0% | 120 |
| 9 | Microbiology | 106 | 4% | 0% | 157 |
| 10 | Computer Science, Interdisciplinary Applications | 62 | 2% | 0% | 74 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | HIV DRUG ISTANCE PROGRAM | 360101 | 4% | 30% | 173 |
| 2 | REGA MED | 122213 | 5% | 8% | 232 |
| 3 | MCGILL AIDS | 99897 | 1% | 28% | 51 |
| 4 | RT BIOCHEM SECT | 78140 | 0% | 62% | 18 |
| 5 | ISTANCE MECH | 69535 | 0% | 76% | 13 |
| 6 | AIDS VACCINE PROGRAM | 68062 | 1% | 19% | 51 |
| 7 | ORETRO | 63773 | 1% | 40% | 23 |
| 8 | REVERSE TRANSCRIPTASE BIOCHEM SECT | 51507 | 0% | 82% | 9 |
| 9 | VIROL DRUG DISCOVERY | 41971 | 0% | 100% | 6 |
| 10 | PHARMACOCHIM MOL STRUCT | 36056 | 1% | 20% | 26 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF VIROLOGY | 50992 | 12% | 1% | 539 |
| 2 | ANTIVIRAL CHEMISTRY & CHEMOTHERAPY | 27279 | 1% | 9% | 44 |
| 3 | JOURNAL OF MEDICINAL CHEMISTRY | 11275 | 4% | 1% | 188 |
| 4 | RETROVIROLOGY | 10563 | 1% | 4% | 43 |
| 5 | JOURNAL OF MOLECULAR BIOLOGY | 8952 | 4% | 1% | 175 |
| 6 | VIROLOGY | 6365 | 3% | 1% | 132 |
| 7 | BIOCHEMISTRY | 5712 | 5% | 0% | 214 |
| 8 | NUCLEIC ACIDS RESEARCH | 5600 | 4% | 0% | 182 |
| 9 | ANTIVIRAL RESEARCH | 4380 | 1% | 1% | 47 |
| 10 | BIOORGANIC & MEDICINAL CHEMISTRY | 3936 | 2% | 1% | 87 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | NNRTIS | 794278 | 5% | 53% | 215 | Search NNRTIS | Search NNRTIS |
| 2 | REVERSE TRANSCRIPTASE | 599364 | 8% | 25% | 346 | Search REVERSE+TRANSCRIPTASE | Search REVERSE+TRANSCRIPTASE |
| 3 | HIV 1 REVERSE TRANSCRIPTASE | 267316 | 2% | 45% | 85 | Search HIV+1+REVERSE+TRANSCRIPTASE | Search HIV+1+REVERSE+TRANSCRIPTASE |
| 4 | NCP7 | 132151 | 1% | 82% | 23 | Search NCP7 | Search NCP7 |
| 5 | RNA DIMERIZATION | 128532 | 0% | 88% | 21 | Search RNA+DIMERIZATION | Search RNA+DIMERIZATION |
| 6 | NON NUCLEOSIDE REVERSE TRANSCRIPTASE INHIBITORS | 128257 | 2% | 27% | 68 | Search NON+NUCLEOSIDE+REVERSE+TRANSCRIPTASE+INHIBITORS | Search NON+NUCLEOSIDE+REVERSE+TRANSCRIPTASE+INHIBITORS |
| 7 | NUCLEOCAPSID PROTEIN | 127497 | 2% | 24% | 77 | Search NUCLEOCAPSID+PROTEIN | Search NUCLEOCAPSID+PROTEIN |
| 8 | HIV 1 | 123860 | 9% | 5% | 385 | Search HIV+1 | Search HIV+1 |
| 9 | REVERSE TRANSCRIPTION | 120984 | 2% | 17% | 101 | Search REVERSE+TRANSCRIPTION | Search REVERSE+TRANSCRIPTION |
| 10 | S DABOS | 111923 | 0% | 100% | 16 | Search S+DABOS | Search S+DABOS |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | LEVIN, JG , GUO, JH , ROUZINA, I , MUSIER-FORSYTH, K , (2005) NUCLEIC ACID CHAPERONE ACTIVITY OF HIV-1 NUCLEOCAPSID PROTEIN: CRITICAL ROLE IN REVERSE TRANSCRIPTION AND MOLECULAR MECHANISM.PROGRESS IN NUCLEIC ACID RESEARCH AND MOLECULAR BIOLOGY, VOL 80. VOL. 80. ISSUE . P. 217 -286 | 308 | 83% | 228 |
| 2 | ZHAN, P , CHEN, XW , LI, DY , FANG, ZJ , DE CLERCQ, E , LIU, XY , (2013) HIV-1 NNRTIS: STRUCTURAL DIVERSITY, PHARMACOPHORE SIMILARITY, AND IMPLIATIONS FOR DRUG DESIGN.MEDICINAL RESEARCH REVIEWS. VOL. 33. ISSUE . P. E1-E72 | 216 | 67% | 32 |
| 3 | LU, K , HENG, X , SUMMERS, MF , (2011) STRUCTURAL DETERMINANTS AND MECHANISM OF HIV-1 GENOME PACKAGING.JOURNAL OF MOLECULAR BIOLOGY. VOL. 410. ISSUE 4. P. 609-633 | 185 | 73% | 89 |
| 4 | LEVIN, JG , MITRA, M , MASCARENHAS, A , MUSIER-FORSYTH, K , (2010) ROLE OF HIV-1 NUCLEOCAPSID PROTEIN IN HIV-1 REVERSE TRANSCRIPTION.RNA BIOLOGY. VOL. 7. ISSUE 6. P. 754 -774 | 178 | 75% | 49 |
| 5 | BASU, VP , SONG, M , GAO, L , RIGBY, ST , HANSON, MN , BAMBARA, RA , (2008) STRAND TRANSFER EVENTS DURING HIV-1 REVERSE TRANSCRIPTION.VIRUS RESEARCH. VOL. 134. ISSUE 1-2. P. 19-38 | 148 | 94% | 61 |
| 6 | DARLIX, JL , DE ROCQUIGNY, H , MAUFFRET, O , MELY, Y , (2014) RETROSPECTIVE ON THE ALL-IN-ONE RETROVIRAL NUCLEOCAPSID PROTEIN.VIRUS RESEARCH. VOL. 193. ISSUE . P. 2 -15 | 176 | 68% | 6 |
| 7 | MORI, M , KOVALENKO, L , LYONNAIS, S , ANTAKI, D , TORBETT, BE , BOTTA, M , MIRAMBEAU, G , MELY, Y , (2015) NUCLEOCAPSID PROTEIN: A DESIRABLE TARGET FOR FUTURE THERAPIES AGAINST HIV-1.FUTURE OF HIV-1 THERAPEUTICS: RESISTANCE IS FUTILE?. VOL. 389. ISSUE . P. 53 -92 | 134 | 78% | 8 |
| 8 | D'SOUZA, V , SUMMERS, MF , (2005) HOW RETROVIRUSES SELECT THEIR GENOMES.NATURE REVIEWS MICROBIOLOGY. VOL. 3. ISSUE 8. P. 643-655 | 133 | 86% | 180 |
| 9 | ALI, LM , RIZVI, TA , MUSTAFA, F , (2016) CROSS- AND CO-PACKAGING OF RETROVIRAL RNAS AND THEIR CONSEQUENCES.VIRUSES-BASEL. VOL. 8. ISSUE 10. P. - | 150 | 71% | 0 |
| 10 | SCHULTZ, SJ , CHAMPOUX, JJ , (2008) RNASE H ACTIVITY: STRUCTURE, SPECIFICITY, AND FUNCTION IN REVERSE TRANSCRIPTION.VIRUS RESEARCH. VOL. 134. ISSUE 1-2. P. 86-103 | 131 | 86% | 53 |
Classes with closest relation at Level 2 |