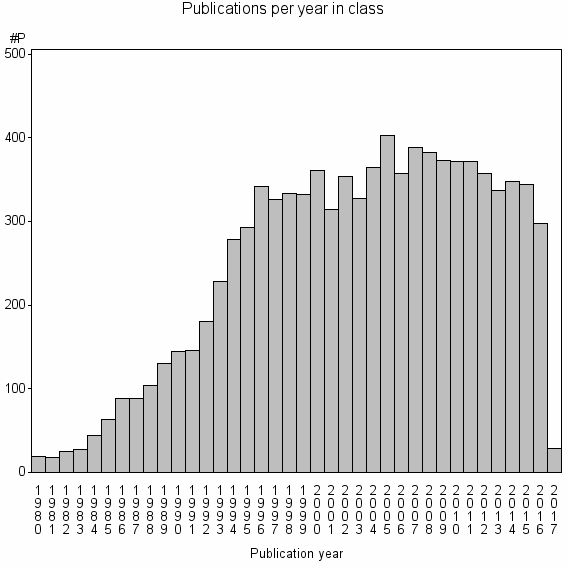
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 1185 | 9289 | 42.8 | 84% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 5 | 4 | CHEMISTRY, ORGANIC//CHEMISTRY, INORGANIC & NUCLEAR//CHEMISTRY, MULTIDISCIPLINARY | 1745167 |
| 183 | 3 | APTAMER//G QUADRUPLEX//RIBOZYME | 56307 |
| 1185 | 2 | RIBOZYME//RNASE P//CATALYTIC RNA | 9289 |
| 2673 | 1 | RNA SECONDARY STRUCTURE//RNA SECONDARY STRUCTURE PREDICTION//PSEUDOKNOT | 2177 |
| 2916 | 1 | RIBOZYME//HAMMERHEAD RIBOZYME//CATALYTIC RNA | 2115 |
| 4185 | 1 | GROUP I INTRON//RIBOZYME//SELF SPLICING | 1832 |
| 10611 | 1 | RNASE P//RIBONUCLEASE P//CARTILAGE HAIR HYPOPLASIA | 1048 |
| 11256 | 1 | DEOXYRIBOZYME//DNAZYME//CATALYTIC DNA | 996 |
| 12082 | 1 | RIBOSWITCH//CANC UK NUCLE ACID STRUCT GRP//RNA SWITCH | 933 |
| 29317 | 1 | AOGEN//BRADYTROPHIC PHENOCOPY//CONTROL OF TRANSGENE EXPRESSION | 188 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | RIBOZYME | authKW | 1863746 | 9% | 65% | 873 |
| 2 | RNASE P | authKW | 617922 | 2% | 85% | 222 |
| 3 | CATALYTIC RNA | authKW | 485439 | 2% | 83% | 177 |
| 4 | RIBOSWITCH | authKW | 461510 | 2% | 68% | 206 |
| 5 | GROUP I INTRON | authKW | 398440 | 2% | 59% | 204 |
| 6 | RNA FOLDING | authKW | 388378 | 2% | 56% | 212 |
| 7 | RNA CATALYSIS | authKW | 325893 | 1% | 87% | 114 |
| 8 | HAMMERHEAD RIBOZYME | authKW | 306536 | 2% | 67% | 140 |
| 9 | RNA STRUCTURE | authKW | 301462 | 3% | 34% | 267 |
| 10 | RIBONUCLEASE P | authKW | 207397 | 1% | 96% | 66 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 53259 | 62% | 0% | 5735 |
| 2 | Mathematical & Computational Biology | 10208 | 6% | 1% | 590 |
| 3 | Biochemical Research Methods | 5124 | 9% | 0% | 805 |
| 4 | Biotechnology & Applied Microbiology | 4104 | 10% | 0% | 949 |
| 5 | Biophysics | 2386 | 7% | 0% | 636 |
| 6 | Cell Biology | 2127 | 10% | 0% | 899 |
| 7 | Genetics & Heredity | 1576 | 7% | 0% | 633 |
| 8 | Biology | 354 | 2% | 0% | 230 |
| 9 | Computer Science, Interdisciplinary Applications | 297 | 2% | 0% | 204 |
| 10 | Virology | 232 | 2% | 0% | 162 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CANC UK NUCLE ACID STRUCT GRP | 74444 | 0% | 67% | 34 |
| 2 | PROGRAM COMPARAT BIOCHEM | 68744 | 0% | 70% | 30 |
| 3 | GRP ECOL GENET | 52651 | 0% | 70% | 23 |
| 4 | SINGLE MOL ANAL GRP | 42740 | 0% | 48% | 27 |
| 5 | MOL BIOPHYS BIOCHEM | 38954 | 2% | 6% | 187 |
| 6 | RNA GRP | 32450 | 0% | 27% | 36 |
| 7 | NONCODING RNA TECHNOL HLTH | 31391 | 0% | 27% | 35 |
| 8 | MOL CELLULAR DEV BIOL | 31318 | 3% | 4% | 242 |
| 9 | GRP BIOINFORMAT COMPUTAT BIOL | 27821 | 0% | 71% | 12 |
| 10 | RNA BIOL | 26987 | 1% | 10% | 80 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | RNA | 167608 | 4% | 14% | 360 |
| 2 | RNA-A PUBLICATION OF THE RNA SOCIETY | 113640 | 2% | 17% | 210 |
| 3 | NUCLEIC ACIDS RESEARCH | 79706 | 11% | 3% | 983 |
| 4 | JOURNAL OF MOLECULAR BIOLOGY | 22481 | 4% | 2% | 404 |
| 5 | ANTISENSE & NUCLEIC ACID DRUG DEVELOPMENT | 18607 | 0% | 12% | 46 |
| 6 | BIOCHEMISTRY | 18013 | 6% | 1% | 551 |
| 7 | RNA BIOLOGY | 12361 | 1% | 6% | 59 |
| 8 | BIOINFORMATICS | 10593 | 2% | 2% | 188 |
| 9 | JOURNAL OF COMPUTATIONAL BIOLOGY | 9914 | 1% | 4% | 69 |
| 10 | CHEMISTRY & BIOLOGY | 9399 | 1% | 3% | 88 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | RIBOZYME | 1863746 | 9% | 65% | 873 | Search RIBOZYME | Search RIBOZYME |
| 2 | RNASE P | 617922 | 2% | 85% | 222 | Search RNASE+P | Search RNASE+P |
| 3 | CATALYTIC RNA | 485439 | 2% | 83% | 177 | Search CATALYTIC+RNA | Search CATALYTIC+RNA |
| 4 | RIBOSWITCH | 461510 | 2% | 68% | 206 | Search RIBOSWITCH | Search RIBOSWITCH |
| 5 | GROUP I INTRON | 398440 | 2% | 59% | 204 | Search GROUP+I+INTRON | Search GROUP+I+INTRON |
| 6 | RNA FOLDING | 388378 | 2% | 56% | 212 | Search RNA+FOLDING | Search RNA+FOLDING |
| 7 | RNA CATALYSIS | 325893 | 1% | 87% | 114 | Search RNA+CATALYSIS | Search RNA+CATALYSIS |
| 8 | HAMMERHEAD RIBOZYME | 306536 | 2% | 67% | 140 | Search HAMMERHEAD+RIBOZYME | Search HAMMERHEAD+RIBOZYME |
| 9 | RNA STRUCTURE | 301462 | 3% | 34% | 267 | Search RNA+STRUCTURE | Search RNA+STRUCTURE |
| 10 | RIBONUCLEASE P | 207397 | 1% | 96% | 66 | Search RIBONUCLEASE+P | Search RIBONUCLEASE+P |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | KRASILNIKOV, AS , ESAKOVA, O , (2010) OF PROTEINS AND RNA: THE RNASE P/MRP FAMILY.RNA. VOL. 16. ISSUE 9. P. 1725 -1747 | 234 | 92% | 75 |
| 2 | PUERTA-FERNANDEZ, E , ROMERO-LOPEZ, C , BARROSO-DELJESUS, A , BERZAL-HERRANZ, A , (2003) RIBOZYMES: RECENT ADVANCES IN THE DEVELOPMENT OF RNA TOOLS.FEMS MICROBIOLOGY REVIEWS. VOL. 27. ISSUE 1. P. 75-97 | 262 | 90% | 64 |
| 3 | FEDOR, MJ , (2009) COMPARATIVE ENZYMOLOGY AND STRUCTURAL BIOLOGY OF RNA SELF-CLEAVAGE.ANNUAL REVIEW OF BIOPHYSICS. VOL. 38. ISSUE . P. 271-299 | 163 | 95% | 77 |
| 4 | WARD, WL , PLAKOS, K , DEROSE, VJ , (2014) NUCLEIC ACID CATALYSIS: METALS, NUCLEOBASES, AND OTHER COFACTORS.CHEMICAL REVIEWS. VOL. 114. ISSUE 8. P. 4318 -4342 | 142 | 88% | 35 |
| 5 | LONNBERG, T , (2011) UNDERSTANDING CATALYSIS OF PHOSPHATE-TRANSFER REACTIONS BY THE LARGE RIBOZYMES.CHEMISTRY-A EUROPEAN JOURNAL. VOL. 17. ISSUE 26. P. 7140 -7153 | 187 | 89% | 3 |
| 6 | PESELIS, A , SERGANOV, A , (2014) THEMES AND VARIATIONS IN RIBOSWITCH STRUCTURE AND FUNCTION.BIOCHIMICA ET BIOPHYSICA ACTA-GENE REGULATORY MECHANISMS. VOL. 1839. ISSUE 10. P. 908 -918 | 123 | 87% | 28 |
| 7 | WALKER, SC , ENGELKE, DR , (2006) RIBONUCLEASE P: THE EVOLUTION OF AN ANCIENT RNA ENZYME.CRITICAL REVIEWS IN BIOCHEMISTRY AND MOLECULAR BIOLOGY. VOL. 41. ISSUE 2. P. 77-102 | 161 | 84% | 108 |
| 8 | SERGANOV, A , PATEL, DJ , (2012) METABOLITE RECOGNITION PRINCIPLES AND MOLECULAR MECHANISMS UNDERLYING RIBOSWITCH FUNCTION.ANNUAL REVIEW OF BIOPHYSICS, VOL 41. VOL. 41. ISSUE . P. 343-370 | 122 | 92% | 50 |
| 9 | FEDOR, MJ , (2000) STRUCTURE AND FUNCTION OF THE HAIRPIN RIBOZYME.JOURNAL OF MOLECULAR BIOLOGY. VOL. 297. ISSUE 2. P. 269 -291 | 159 | 95% | 147 |
| 10 | MONDRAGON, A , (2013) STRUCTURAL STUDIES OF RNASE P.ANNUAL REVIEW OF BIOPHYSICS, VOL 42. VOL. 42. ISSUE . P. 537-557 | 115 | 94% | 17 |
Classes with closest relation at Level 2 |