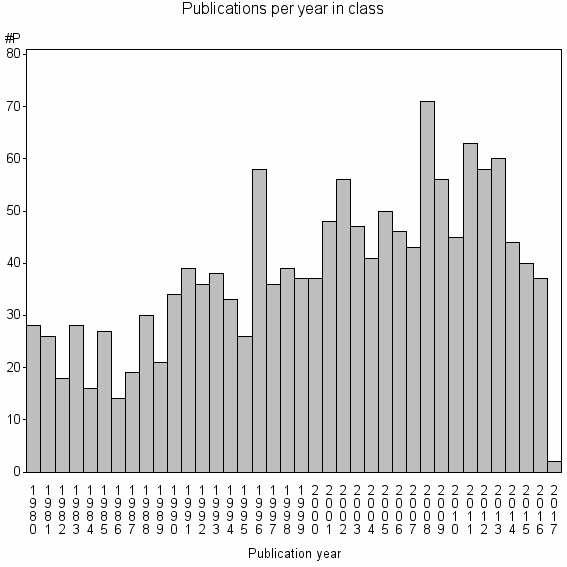
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 6760 | 1447 | 45.0 | 76% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 7 | 4 | INFECTIOUS DISEASES//MICROBIOLOGY//VIROLOGY | 1353914 |
| 33 | 3 | JOURNAL OF BACTERIOLOGY//MICROBIOLOGY//MOLECULAR MICROBIOLOGY | 107033 |
| 371 | 2 | RNA POLYMERASE//HFQ//JOURNAL OF BACTERIOLOGY | 17271 |
| 6760 | 1 | RNASE E//POLYNUCLEOTIDE PHOSPHORYLASE//RNA DEGRADOSOME | 1447 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | RNASE E | authKW | 1259897 | 6% | 70% | 85 |
| 2 | POLYNUCLEOTIDE PHOSPHORYLASE | authKW | 498026 | 3% | 64% | 37 |
| 3 | RNA DEGRADOSOME | authKW | 419098 | 2% | 83% | 24 |
| 4 | DEGRADOSOME | authKW | 415729 | 2% | 73% | 27 |
| 5 | PNPASE | authKW | 379791 | 2% | 60% | 30 |
| 6 | RNASE G | authKW | 338784 | 1% | 94% | 17 |
| 7 | RNA DEGRADATION | authKW | 307452 | 4% | 24% | 60 |
| 8 | RNASE II | authKW | 304687 | 1% | 76% | 19 |
| 9 | RNASE III | authKW | 290347 | 2% | 40% | 34 |
| 10 | UPR 9073 | address | 163643 | 2% | 24% | 32 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 10067 | 67% | 0% | 976 |
| 2 | Microbiology | 7733 | 31% | 0% | 449 |
| 3 | Genetics & Heredity | 520 | 9% | 0% | 133 |
| 4 | Cell Biology | 468 | 11% | 0% | 160 |
| 5 | Biophysics | 310 | 6% | 0% | 92 |
| 6 | Biochemical Research Methods | 104 | 4% | 0% | 55 |
| 7 | Biotechnology & Applied Microbiology | 63 | 4% | 0% | 63 |
| 8 | Biology | 20 | 2% | 0% | 26 |
| 9 | Developmental Biology | 15 | 1% | 0% | 15 |
| 10 | Virology | 3 | 1% | 0% | 13 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | UPR 9073 | 163643 | 2% | 24% | 32 |
| 2 | CNRS UPR 9073 | 90023 | 1% | 53% | 8 |
| 3 | UMR 5100 | 51913 | 1% | 15% | 16 |
| 4 | ICSN RMN | 48227 | 0% | 57% | 4 |
| 5 | MIKRO MOL BIOL | 42194 | 0% | 33% | 6 |
| 6 | BIOL PHYSICOCHIM | 41157 | 2% | 6% | 31 |
| 7 | GRP ARN | 34520 | 0% | 27% | 6 |
| 8 | MICROBIOL GENET MOL | 29656 | 2% | 6% | 25 |
| 9 | FRC550 | 27126 | 0% | 43% | 3 |
| 10 | UPR9073 | 25494 | 1% | 13% | 9 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOLECULAR MICROBIOLOGY | 29503 | 9% | 1% | 129 |
| 2 | JOURNAL OF BACTERIOLOGY | 17872 | 12% | 1% | 168 |
| 3 | RNA | 7951 | 2% | 1% | 31 |
| 4 | NUCLEIC ACIDS RESEARCH | 6703 | 8% | 0% | 113 |
| 5 | RNA BIOLOGY | 5148 | 1% | 2% | 15 |
| 6 | JOURNAL OF MOLECULAR BIOLOGY | 4348 | 5% | 0% | 70 |
| 7 | WILEY INTERDISCIPLINARY REVIEWS-RNA | 3664 | 1% | 2% | 8 |
| 8 | PROGRESS IN MOLECULAR BIOLOGY AND TRANSLATIONAL SCIENCE | 3153 | 1% | 2% | 10 |
| 9 | RNA-A PUBLICATION OF THE RNA SOCIETY | 2780 | 1% | 1% | 13 |
| 10 | BIOCHIMIE | 2527 | 2% | 0% | 27 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | RNASE E | 1259897 | 6% | 70% | 85 | Search RNASE+E | Search RNASE+E |
| 2 | POLYNUCLEOTIDE PHOSPHORYLASE | 498026 | 3% | 64% | 37 | Search POLYNUCLEOTIDE+PHOSPHORYLASE | Search POLYNUCLEOTIDE+PHOSPHORYLASE |
| 3 | RNA DEGRADOSOME | 419098 | 2% | 83% | 24 | Search RNA+DEGRADOSOME | Search RNA+DEGRADOSOME |
| 4 | DEGRADOSOME | 415729 | 2% | 73% | 27 | Search DEGRADOSOME | Search DEGRADOSOME |
| 5 | PNPASE | 379791 | 2% | 60% | 30 | Search PNPASE | Search PNPASE |
| 6 | RNASE G | 338784 | 1% | 94% | 17 | Search RNASE+G | Search RNASE+G |
| 7 | RNA DEGRADATION | 307452 | 4% | 24% | 60 | Search RNA+DEGRADATION | Search RNA+DEGRADATION |
| 8 | RNASE II | 304687 | 1% | 76% | 19 | Search RNASE+II | Search RNASE+II |
| 9 | RNASE III | 290347 | 2% | 40% | 34 | Search RNASE+III | Search RNASE+III |
| 10 | RNASE R | 159913 | 1% | 63% | 12 | Search RNASE+R | Search RNASE+R |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | ARRAIANO, CM , ANDRADE, JM , DOMINGUES, S , GUINOTE, IB , MALECKI, M , MATOS, RG , MOREIRA, RN , POBRE, V , REIS, FP , SARAMAGO, M , ET AL (2010) THE CRITICAL ROLE OF RNA PROCESSING AND DEGRADATION IN THE CONTROL OF GENE EXPRESSION.FEMS MICROBIOLOGY REVIEWS. VOL. 34. ISSUE 5. P. 883 -923 | 284 | 64% | 116 |
| 2 | LAALAMI, S , ZIG, L , PUTZER, H , (2014) INITIATION OF MRNA DECAY IN BACTERIA.CELLULAR AND MOLECULAR LIFE SCIENCES. VOL. 71. ISSUE 10. P. 1799 -1828 | 184 | 63% | 23 |
| 3 | CARPOUSIS, AJ , LUISI, BF , MCDOWALL, KJ , (2009) ENDONUCLEOLYTIC INITIATION OF MRNA DECAY IN ESCHERICHIA COLI.MOLECULAR BIOLOGY OF RNA PROCESSING AND DECAY IN PROKARYOTES. VOL. 85. ISSUE . P. 91 -135 | 155 | 78% | 68 |
| 4 | HUI, MP , FOLEY, PL , BELASCO, JG , (2014) MESSENGER RNA DEGRADATION IN BACTERIAL CELLS.ANNUAL REVIEW OF GENETICS, VOL 48. VOL. 48. ISSUE . P. 537 -559 | 121 | 80% | 18 |
| 5 | MACKIE, GA , (2013) RNASE E: AT THE INTERFACE OF BACTERIAL RNA PROCESSING AND DECAY.NATURE REVIEWS MICROBIOLOGY. VOL. 11. ISSUE 1. P. 45-57 | 96 | 82% | 67 |
| 6 | MOHANTY, BK , KUSHNER, SR , (2016) REGULATION OF MRNA DECAY IN BACTERIA.ANNUAL REVIEW OF MICROBIOLOGY, VOL 70. VOL. 70. ISSUE . P. 25 -44 | 108 | 81% | 2 |
| 7 | GRUNBERG-MANAGO, M , (1999) MESSENGER RNA STABILITY AND ITS ROLE IN CONTROL OF GENE EXPRESSION IN BACTERIA AND PHAGES.ANNUAL REVIEW OF GENETICS. VOL. 33. ISSUE . P. 193-227 | 144 | 84% | 207 |
| 8 | ANDRADE, JM , POBRE, V , SILVA, IJ , DOMINGUES, S , ARRAIANO, CM , (2009) THE ROLE OF 3 '-5 ' EXORIBONUCLEASES IN RNA DEGRADATION.MOLECULAR BIOLOGY OF RNA PROCESSING AND DECAY IN PROKARYOTES. VOL. 85. ISSUE . P. 187 -229 | 144 | 72% | 43 |
| 9 | ARRAIANO, CM , MAUXION, F , VIEGAS, SC , MATOS, RG , SERAPHIN, B , (2013) INTRACELLULAR RIBONUCLEASES INVOLVED IN TRANSCRIPT PROCESSING AND DECAY: PRECISION TOOLS FOR RNA.BIOCHIMICA ET BIOPHYSICA ACTA-GENE REGULATORY MECHANISMS. VOL. 1829. ISSUE 6-7. P. 491-513 | 179 | 51% | 15 |
| 10 | BRIANI, F , CARZANIGA, T , DEHO, G , (2016) REGULATION AND FUNCTIONS OF BACTERIAL PNPASE.WILEY INTERDISCIPLINARY REVIEWS-RNA. VOL. 7. ISSUE 2. P. 241 -258 | 104 | 76% | 2 |
Classes with closest relation at Level 1 |
| Rank | Class id | link |
|---|---|---|
| 1 | 17891 | RRP6//MRNP BIOGENESIS METAB//DIS3 |
| 2 | 7121 | HFQ//SRNA//RYHB |
| 3 | 19084 | TRNASE Z//TRNA NUCLEOTIDYLTRANSFERASE//CCA ADDING ENZYME |
| 4 | 32149 | MINIATURE INSERTION SEQUENCE//REPEATED DNA FAMILIES//AVENTIS UMR 1889 |
| 5 | 23884 | RIBOSOMAL PROTEIN L1//RPMG//L1 CONFORMATIONS |
| 6 | 18779 | TMRNA//TRANS TRANSLATION//SMPB |
| 7 | 9543 | SHINE DALGARNO SEQUENCE//LEADERLESS MRNA//TRANSLATION INITIATION |
| 8 | 18097 | GENE UVSX//BACTERIOPHAGE T4//UVSX |
| 9 | 21689 | OBG//RBFA//DRG2 |
| 10 | 10611 | RNASE P//RIBONUCLEASE P//CARTILAGE HAIR HYPOPLASIA |