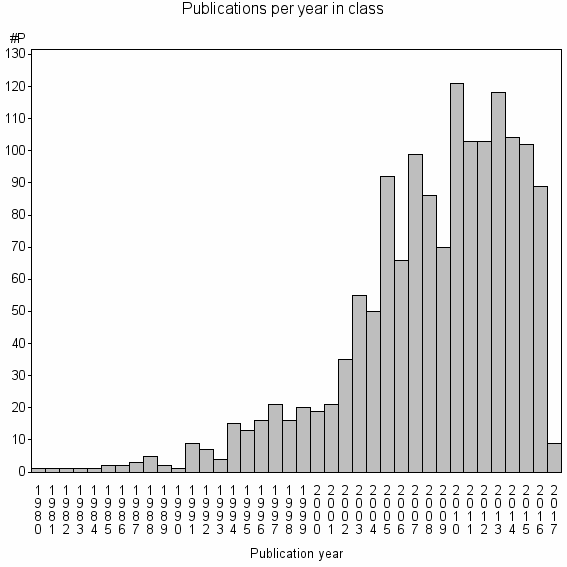
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 6443 | 1483 | 50.5 | 87% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 219 | 3 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN FOLDING//PROTEIN STRUCTURE PREDICTION | 50409 |
| 365 | 2 | PROTEIN STRUCTURE PREDICTION//PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PSEUDO AMINO ACID COMPOSITION | 17331 |
| 6443 | 1 | PROTEIN PROTEIN DOCKING//CAPRI//PROTEIN DOCKING | 1483 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | PROTEIN PROTEIN DOCKING | authKW | 931233 | 7% | 47% | 97 |
| 2 | CAPRI | authKW | 838139 | 4% | 77% | 53 |
| 3 | PROTEIN DOCKING | authKW | 605899 | 4% | 45% | 66 |
| 4 | PROTEIN PROTEIN INTERFACE | authKW | 471627 | 3% | 55% | 42 |
| 5 | PROTEIN INTERFACE | authKW | 298835 | 2% | 48% | 30 |
| 6 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS | journal | 292859 | 20% | 5% | 301 |
| 7 | SHAPE COMPLEMENTARITY | authKW | 245375 | 1% | 57% | 21 |
| 8 | INTERFACE PREDICTION | authKW | 210816 | 1% | 64% | 16 |
| 9 | PROTEIN PROTEIN INTERACTION | authKW | 209483 | 17% | 4% | 256 |
| 10 | ZDOCK | authKW | 185291 | 1% | 75% | 12 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biophysics | 12007 | 33% | 0% | 486 |
| 2 | Mathematical & Computational Biology | 11782 | 17% | 0% | 245 |
| 3 | Biochemistry & Molecular Biology | 9411 | 65% | 0% | 959 |
| 4 | Biochemical Research Methods | 4690 | 19% | 0% | 286 |
| 5 | Biotechnology & Applied Microbiology | 1113 | 13% | 0% | 190 |
| 6 | Computer Science, Interdisciplinary Applications | 674 | 6% | 0% | 94 |
| 7 | Cell Biology | 108 | 6% | 0% | 96 |
| 8 | Chemistry, Medicinal | 85 | 3% | 0% | 45 |
| 9 | Computer Science, Information Systems | 61 | 2% | 0% | 34 |
| 10 | Chemistry, Multidisciplinary | 32 | 6% | 0% | 94 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | SACKLER MOL MED | 178874 | 4% | 16% | 55 |
| 2 | HUMAN GENET MOL MED | 102036 | 3% | 10% | 50 |
| 3 | BIOMOL MODELLING | 91474 | 1% | 22% | 20 |
| 4 | BC6 | 61766 | 0% | 100% | 3 |
| 5 | CP 263 | 61766 | 0% | 100% | 3 |
| 6 | EXPT COMPUTAT BIOL | 58137 | 2% | 9% | 31 |
| 7 | IBBMC UMR 8619 | 54900 | 0% | 67% | 4 |
| 8 | JOINT BSC IRB PROGRAMME COMPUTAT BIOL | 54892 | 1% | 33% | 8 |
| 9 | JOINT BSC IRB PROGRAM COMPUTAT BIOL | 54166 | 1% | 26% | 10 |
| 10 | PROGRAM MOL COMPUTAT BIOPHYS | 45847 | 0% | 32% | 7 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS | 292859 | 20% | 5% | 301 |
| 2 | BIOINFORMATICS | 14202 | 6% | 1% | 86 |
| 3 | BMC BIOINFORMATICS | 14134 | 5% | 1% | 72 |
| 4 | PROTEIN SCIENCE | 12141 | 4% | 1% | 61 |
| 5 | CURRENT OPINION IN STRUCTURAL BIOLOGY | 9216 | 2% | 1% | 32 |
| 6 | BMC STRUCTURAL BIOLOGY | 8027 | 1% | 3% | 14 |
| 7 | JOURNAL OF MOLECULAR BIOLOGY | 6138 | 6% | 0% | 84 |
| 8 | STRUCTURE | 5508 | 2% | 1% | 32 |
| 9 | JOURNAL OF CHEMICAL INFORMATION AND MODELING | 4787 | 2% | 1% | 27 |
| 10 | PROTEIN ENGINEERING | 2920 | 1% | 1% | 18 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PROTEIN PROTEIN DOCKING | 931233 | 7% | 47% | 97 | Search PROTEIN+PROTEIN+DOCKING | Search PROTEIN+PROTEIN+DOCKING |
| 2 | CAPRI | 838139 | 4% | 77% | 53 | Search CAPRI | Search CAPRI |
| 3 | PROTEIN DOCKING | 605899 | 4% | 45% | 66 | Search PROTEIN+DOCKING | Search PROTEIN+DOCKING |
| 4 | PROTEIN PROTEIN INTERFACE | 471627 | 3% | 55% | 42 | Search PROTEIN+PROTEIN+INTERFACE | Search PROTEIN+PROTEIN+INTERFACE |
| 5 | PROTEIN INTERFACE | 298835 | 2% | 48% | 30 | Search PROTEIN+INTERFACE | Search PROTEIN+INTERFACE |
| 6 | SHAPE COMPLEMENTARITY | 245375 | 1% | 57% | 21 | Search SHAPE+COMPLEMENTARITY | Search SHAPE+COMPLEMENTARITY |
| 7 | INTERFACE PREDICTION | 210816 | 1% | 64% | 16 | Search INTERFACE+PREDICTION | Search INTERFACE+PREDICTION |
| 8 | PROTEIN PROTEIN INTERACTION | 209483 | 17% | 4% | 256 | Search PROTEIN+PROTEIN+INTERACTION | Search PROTEIN+PROTEIN+INTERACTION |
| 9 | ZDOCK | 185291 | 1% | 75% | 12 | Search ZDOCK | Search ZDOCK |
| 10 | UNBOUND DOCKING | 144120 | 0% | 100% | 7 | Search UNBOUND+DOCKING | Search UNBOUND+DOCKING |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | HUANG, SY , (2014) SEARCH STRATEGIES AND EVALUATION IN PROTEIN-PROTEIN DOCKING: PRINCIPLES, ADVANCES AND CHALLENGES.DRUG DISCOVERY TODAY. VOL. 19. ISSUE 8. P. 1081 -1096 | 143 | 89% | 21 |
| 2 | RITCHIE, DW , (2008) RECENT PROGRESS AND FUTURE DIRECTIONS IN PROTEIN-PROTEIN DOCKING.CURRENT PROTEIN & PEPTIDE SCIENCE. VOL. 9. ISSUE 1. P. 1 -15 | 138 | 68% | 150 |
| 3 | MOREIRA, IS , FERNANDES, PA , RAMOS, MJ , (2010) PROTEIN-PROTEIN DOCKING DEALING WITH THE UNKNOWN.JOURNAL OF COMPUTATIONAL CHEMISTRY. VOL. 31. ISSUE 2. P. 317 -342 | 133 | 73% | 52 |
| 4 | HUANG, SY , (2015) EXPLORING THE POTENTIAL OF GLOBAL PROTEIN-PROTEIN DOCKING: AN OVERVIEW AND CRITICAL ASSESSMENT OF CURRENT PROGRAMS FOR AUTOMATIC AB INITIO DOCKING.DRUG DISCOVERY TODAY. VOL. 20. ISSUE 8. P. 969 -977 | 77 | 89% | 10 |
| 5 | JANIN, J , (2010) PROTEIN-PROTEIN DOCKING TESTED IN BLIND PREDICTIONS: THE CAPRI EXPERIMENT.MOLECULAR BIOSYSTEMS. VOL. 6. ISSUE 12. P. 2351 -2362 | 92 | 75% | 78 |
| 6 | ESMAIELBEIKI, R , KRAWCZYK, K , KNAPP, B , NEBEL, JC , DEANE, CM , (2016) PROGRESS AND CHALLENGES IN PREDICTING PROTEIN INTERFACES.BRIEFINGS IN BIOINFORMATICS. VOL. 17. ISSUE 1. P. 117 -131 | 96 | 58% | 3 |
| 7 | RODRIGUES, JPGLM , BONVIN, AMJJ , (2014) INTEGRATIVE COMPUTATIONAL MODELING OF PROTEIN INTERACTIONS.FEBS JOURNAL. VOL. 281. ISSUE 8. P. 1988-2003 | 80 | 75% | 20 |
| 8 | KESKIN, O , TUNCBAG, N , GURSOY, A , (2016) PREDICTING PROTEIN-PROTEIN INTERACTIONS FROM THE MOLECULAR TO THE PROTEOME LEVEL.CHEMICAL REVIEWS. VOL. 116. ISSUE 8. P. 4884 -4909 | 97 | 49% | 6 |
| 9 | FERNANDEZ-RECIO, J , (2011) PREDICTION OF PROTEIN BINDING SITES AND HOT SPOTS.WILEY INTERDISCIPLINARY REVIEWS-COMPUTATIONAL MOLECULAR SCIENCE. VOL. 1. ISSUE 5. P. 680-698 | 96 | 72% | 20 |
| 10 | AUMENTADO-ARMSTRONG, TT , ISTRATE, B , MURGITA, RA , (2015) ALGORITHMIC APPROACHES TO PROTEIN-PROTEIN INTERACTION SITE PREDICTION.ALGORITHMS FOR MOLECULAR BIOLOGY. VOL. 10. ISSUE . P. - | 110 | 53% | 5 |
Classes with closest relation at Level 1 |