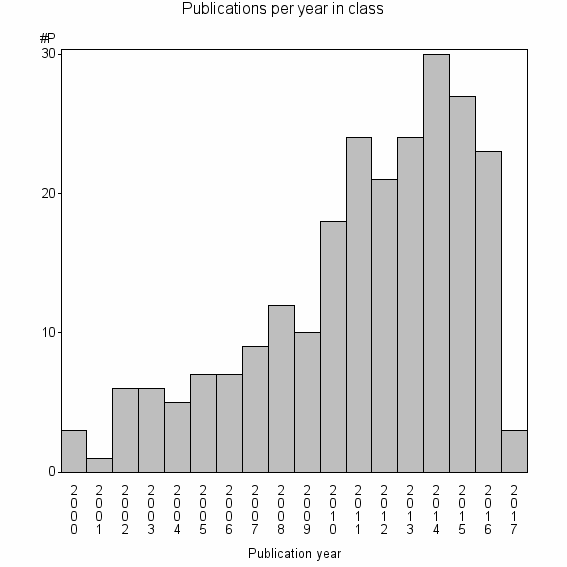
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 27163 | 236 | 48.0 | 93% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 563 | 3 | DNA METHYLATION//EPIGENETICS//METHYLATION | 17086 |
| 382 | 2 | DNA METHYLATION//EPIGENETICS//METHYLATION | 17086 |
| 27163 | 1 | MOL DIAGNOST RD//P16INK4A METHYLATION//TRIPLE COMBINATION ARRAY | 236 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | MOL DIAGNOST RD | address | 231042 | 2% | 36% | 5 |
| 2 | P16INK4A METHYLATION | authKW | 172515 | 1% | 67% | 2 |
| 3 | TRIPLE COMBINATION ARRAY | authKW | 172515 | 1% | 67% | 2 |
| 4 | 450K BEADCHIPS | authKW | 129387 | 0% | 100% | 1 |
| 5 | BACKGROUND LIVER FACTORS | authKW | 129387 | 0% | 100% | 1 |
| 6 | BARCELONA CLIN LIVER CANC GRPCIBEREHDLIVER UNIT | address | 129387 | 0% | 100% | 1 |
| 7 | BARCELONA CLINLIVER CANC GRPLIVER UNITCIBEREHD | address | 129387 | 0% | 100% | 1 |
| 8 | BUILD HEALTHY DIS GENOM | address | 129387 | 0% | 100% | 1 |
| 9 | CANC BRAIN KOREA STAGE 2 | address | 129387 | 0% | 100% | 1 |
| 10 | CANCER DNA METHYLATION | authKW | 129387 | 0% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Gastroenterology & Hepatology | 1683 | 26% | 0% | 61 |
| 2 | Oncology | 1398 | 40% | 0% | 95 |
| 3 | Pathology | 97 | 6% | 0% | 15 |
| 4 | Genetics & Heredity | 78 | 9% | 0% | 21 |
| 5 | Medical Laboratory Technology | 64 | 3% | 0% | 7 |
| 6 | Medicine, Research & Experimental | 20 | 5% | 0% | 11 |
| 7 | Cell Biology | 16 | 6% | 0% | 15 |
| 8 | Biotechnology & Applied Microbiology | 9 | 4% | 0% | 10 |
| 9 | Medicine, General & Internal | 6 | 4% | 0% | 10 |
| 10 | Biochemistry & Molecular Biology | 3 | 8% | 0% | 19 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOL DIAGNOST RD | 231042 | 2% | 36% | 5 |
| 2 | BARCELONA CLIN LIVER CANC GRPCIBEREHDLIVER UNIT | 129387 | 0% | 100% | 1 |
| 3 | BARCELONA CLINLIVER CANC GRPLIVER UNITCIBEREHD | 129387 | 0% | 100% | 1 |
| 4 | BUILD HEALTHY DIS GENOM | 129387 | 0% | 100% | 1 |
| 5 | CANC BRAIN KOREA STAGE 2 | 129387 | 0% | 100% | 1 |
| 6 | CLIN TRANSLAT C HELIOS MED WUPPE | 129387 | 0% | 100% | 1 |
| 7 | EX IMMUNOONCOL MED | 129387 | 0% | 100% | 1 |
| 8 | GASTROINTESTINAL CANC SAMMONS CANC | 129387 | 0% | 100% | 1 |
| 9 | GENET HERED SECT | 129387 | 0% | 100% | 1 |
| 10 | HLTH PAEDIAT | 129387 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CLINICAL EPIGENETICS | 11461 | 3% | 1% | 6 |
| 2 | DIGESTIVE DISEASES | 3634 | 3% | 0% | 7 |
| 3 | EPIGENOMICS | 2515 | 1% | 1% | 3 |
| 4 | EXPERIMENTAL AND MOLECULAR PATHOLOGY | 1740 | 3% | 0% | 6 |
| 5 | LIVER CANCER | 1360 | 0% | 1% | 1 |
| 6 | LIVER INTERNATIONAL | 1268 | 2% | 0% | 5 |
| 7 | HEPATOLOGY | 872 | 4% | 0% | 9 |
| 8 | HEPATOLOGY INTERNATIONAL | 795 | 1% | 0% | 2 |
| 9 | SCIENTIFIC DATA | 725 | 0% | 1% | 1 |
| 10 | BMC MEDICAL GENOMICS | 705 | 1% | 0% | 2 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | XU, BY , NIE, YF , LIU, XX , FENG, SQ , YANG, ZL , WANG, ZG , ZHENG, Q , LUO, XY , (2014) QUANTITATIVE ANALYSIS OF APC PROMOTER METHYLATION IN HEPATOCELLULAR CARCINOMA AND ITS PROGNOSTIC IMPLICATIONS.ONCOLOGY LETTERS. VOL. 7. ISSUE 5. P. 1683-1688 | 16 | 80% | 5 |
| 2 | SHEN, J , WANG, S , ZHANG, YJ , KAPPIL, M , WU, HC , KIBRIYA, MG , WANG, Q , JASMINE, F , AHSAN, H , LEE, PH , ET AL (2012) GENOME-WIDE DNA METHYLATION PROFILES IN HEPATOCELLULAR CARCINOMA.HEPATOLOGY. VOL. 55. ISSUE 6. P. 1799-1808 | 19 | 53% | 64 |
| 3 | NISHIDA, N , KUDO, M , (2014) ALTERATION OF EPIGENETIC PROFILE IN HUMAN HEPATOCELLULAR CARCINOMA AND ITS CLINICAL IMPLICATIONS.LIVER CANCER. VOL. 3. ISSUE 3-4. P. 417 -427 | 20 | 56% | 7 |
| 4 | JAIN, S , CHEN, ST , CHANG, KC , LIN, YJ , HU, CT , BOLDBAATAR, B , HAMILTON, JP , LIN, SY , CHANG, TT , CHEN, SH , ET AL (2012) IMPACT OF THE LOCATION OF CPG METHYLATION WITHIN THE GSTP1 GENE ON ITS SPECIFICITY AS A DNA MARKER FOR HEPATOCELLULAR CARCINOMA.PLOS ONE. VOL. 7. ISSUE 4. P. - | 21 | 55% | 11 |
| 5 | NISHIDA, N , IWANISHI, M , MINAMI, T , CHISHINA, H , ARIZUMI, T , TAKITA, M , KITAI, S , YADA, N , IDA, H , HAGIWARA, S , ET AL (2015) HEPATIC DNA METHYLATION IS AFFECTED BY HEPATOCELLULAR CARCINOMA RISK IN PATIENTS WITH AND WITHOUT HEPATITIS VIRUS.DIGESTIVE DISEASES. VOL. 33. ISSUE 6. P. 745 -750 | 17 | 57% | 1 |
| 6 | HUANG, ZH , HU, Y , HUA, D , WU, YY , SONG, MX , CHENG, ZH , (2011) QUANTITATIVE ANALYSIS OF MULTIPLE METHYLATED GENES IN PLASMA FOR THE DIAGNOSIS AND PROGNOSIS OF HEPATOCELLULAR CARCINOMA.EXPERIMENTAL AND MOLECULAR PATHOLOGY. VOL. 91. ISSUE 3. P. 702-707 | 14 | 64% | 23 |
| 7 | LIU, M , CUI, LH , LI, CC , ZHANG, L , (2015) ASSOCIATION OF APC, GSTP1 AND SOCS1 PROMOTER METHYLATION WITH THE RISK OF HEPATOCELLULAR CARCINOMA: A META-ANALYSIS.EUROPEAN JOURNAL OF CANCER PREVENTION. VOL. 24. ISSUE 6. P. 470 -483 | 17 | 45% | 1 |
| 8 | MAH, WC , THURNHERR, T , CHOW, PKH , CHUNG, AYF , OOI, LLPJ , TOH, HC , TEH, BT , SAUNTHARARAJAH, Y , LEE, CGL , (2014) METHYLATION PROFILES REVEAL DISTINCT SUBGROUP OF HEPATOCELLULAR CARCINOMA PATIENTS WITH POOR PROGNOSIS.PLOS ONE. VOL. 9. ISSUE 8. P. - | 18 | 38% | 10 |
| 9 | HUANG, WQ , LI, T , YANG, WL , CHAI, XJ , CHEN, KF , WEI, L , DUAN, SW , LI, B , QIN, Y , (2015) ANALYSIS OF DNA METHYLATION IN PLASMA FOR MONITORING HEPATOCARCINOGENESIS.GENETIC TESTING AND MOLECULAR BIOMARKERS. VOL. 19. ISSUE 6. P. 295 -302 | 17 | 43% | 1 |
| 10 | XU, BY , DI, JZ , WANG, ZG , HAN, XD , LI, ZH , LUO, XY , ZHENG, Q , (2013) QUANTITATIVE ANALYSIS OF RASSF1A PROMOTER METHYLATION IN HEPATOCELLULAR CARCINOMA AND ITS PROGNOSTIC IMPLICATIONS.BIOCHEMICAL AND BIOPHYSICAL RESEARCH COMMUNICATIONS. VOL. 438. ISSUE 2. P. 324-328 | 12 | 60% | 9 |
Classes with closest relation at Level 1 |