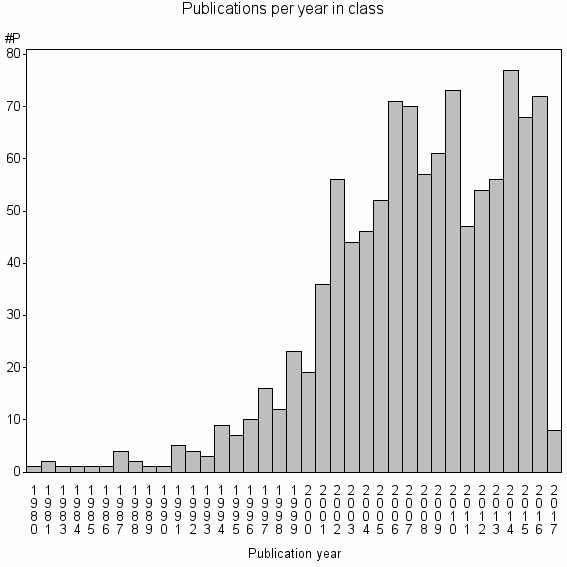
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 10356 | 1071 | 39.1 | 89% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 563 | 3 | DNA METHYLATION//EPIGENETICS//METHYLATION | 17086 |
| 382 | 2 | DNA METHYLATION//EPIGENETICS//METHYLATION | 17086 |
| 10356 | 1 | METHYLTRANSFERASE ACTIVITY//DNA METHYLATION//METHYL BINDING DOMAIN PROTEIN | 1071 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | METHYLTRANSFERASE ACTIVITY | authKW | 215596 | 1% | 69% | 11 |
| 2 | DNA METHYLATION | authKW | 205650 | 25% | 3% | 264 |
| 3 | METHYL BINDING DOMAIN PROTEIN | authKW | 114038 | 0% | 100% | 4 |
| 4 | MS HRM | authKW | 107321 | 1% | 47% | 8 |
| 5 | 5 METHYLCYTOSINE | authKW | 103101 | 3% | 11% | 32 |
| 6 | BISULFITE MODIFICATION | authKW | 100392 | 1% | 39% | 9 |
| 7 | DNA METHYLAT GENOME FUNCT PROJECT | address | 93124 | 1% | 47% | 7 |
| 8 | AMPLIFICATION OF INTERMETHYLATED SITES | authKW | 85529 | 0% | 100% | 3 |
| 9 | DAM MTASE | authKW | 85529 | 0% | 100% | 3 |
| 10 | SEGREGATION OF PARTLY MELTED MOLECULES | authKW | 85529 | 0% | 100% | 3 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemical Research Methods | 2838 | 18% | 0% | 190 |
| 2 | Chemistry, Analytical | 2809 | 23% | 0% | 244 |
| 3 | Biochemistry & Molecular Biology | 1216 | 31% | 0% | 327 |
| 4 | Oncology | 991 | 17% | 0% | 186 |
| 5 | Hematology | 804 | 9% | 0% | 97 |
| 6 | Genetics & Heredity | 629 | 11% | 0% | 121 |
| 7 | Electrochemistry | 549 | 7% | 0% | 71 |
| 8 | Biotechnology & Applied Microbiology | 327 | 9% | 0% | 94 |
| 9 | Medical Laboratory Technology | 254 | 3% | 0% | 30 |
| 10 | Nanoscience & Nanotechnology | 238 | 6% | 0% | 64 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | DNA METHYLAT GENOME FUNCT PROJECT | 93124 | 1% | 47% | 7 |
| 2 | MOL PATHOL DEV | 71259 | 1% | 25% | 10 |
| 3 | BIOCHEM MOL BIOLUROL CANC | 42761 | 0% | 50% | 3 |
| 4 | ELLIS FI EL CANC | 38587 | 1% | 8% | 16 |
| 5 | KH MANAGEMENT | 38011 | 0% | 67% | 2 |
| 6 | PERSONALISED NANOMED | 35083 | 0% | 31% | 4 |
| 7 | ANIM SCI NUTRIT SCI HLTH | 28510 | 0% | 100% | 1 |
| 8 | BIOMED MASS SPE OMETR | 28510 | 0% | 100% | 1 |
| 9 | BIOMOL INTEGRATED MED SCI | 28510 | 0% | 100% | 1 |
| 10 | CANERCOL MARSEILLE | 28510 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOTECHNIQUES | 9056 | 4% | 1% | 44 |
| 2 | BIOSENSORS & BIOELECTRONICS | 6513 | 4% | 0% | 47 |
| 3 | EPIGENOMICS | 6151 | 1% | 2% | 10 |
| 4 | NUCLEIC ACIDS RESEARCH | 3453 | 7% | 0% | 70 |
| 5 | ANALYTICAL BIOCHEMISTRY | 3073 | 4% | 0% | 45 |
| 6 | EXPERT REVIEW OF MOLECULAR DIAGNOSTICS | 2784 | 1% | 1% | 10 |
| 7 | METHODS | 2687 | 2% | 1% | 17 |
| 8 | CIBA FOUNDATION SYMPOSIA | 2030 | 1% | 1% | 14 |
| 9 | LEUKEMIA RESEARCH | 1673 | 2% | 0% | 19 |
| 10 | ANALYTICAL CHEMISTRY | 1407 | 4% | 0% | 43 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | METHYLTRANSFERASE ACTIVITY | 215596 | 1% | 69% | 11 | Search METHYLTRANSFERASE+ACTIVITY | Search METHYLTRANSFERASE+ACTIVITY |
| 2 | DNA METHYLATION | 205650 | 25% | 3% | 264 | Search DNA+METHYLATION | Search DNA+METHYLATION |
| 3 | METHYL BINDING DOMAIN PROTEIN | 114038 | 0% | 100% | 4 | Search METHYL+BINDING+DOMAIN+PROTEIN | Search METHYL+BINDING+DOMAIN+PROTEIN |
| 4 | MS HRM | 107321 | 1% | 47% | 8 | Search MS+HRM | Search MS+HRM |
| 5 | 5 METHYLCYTOSINE | 103101 | 3% | 11% | 32 | Search 5+METHYLCYTOSINE | Search 5+METHYLCYTOSINE |
| 6 | BISULFITE MODIFICATION | 100392 | 1% | 39% | 9 | Search BISULFITE+MODIFICATION | Search BISULFITE+MODIFICATION |
| 7 | AMPLIFICATION OF INTERMETHYLATED SITES | 85529 | 0% | 100% | 3 | Search AMPLIFICATION+OF+INTERMETHYLATED+SITES | Search AMPLIFICATION+OF+INTERMETHYLATED+SITES |
| 8 | DAM MTASE | 85529 | 0% | 100% | 3 | Search DAM+MTASE | Search DAM+MTASE |
| 9 | SEGREGATION OF PARTLY MELTED MOLECULES | 85529 | 0% | 100% | 3 | Search SEGREGATION+OF+PARTLY+MELTED+MOLECULES | Search SEGREGATION+OF+PARTLY+MELTED+MOLECULES |
| 10 | BISULFITE TREATMENT | 82166 | 1% | 41% | 7 | Search BISULFITE+TREATMENT | Search BISULFITE+TREATMENT |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | KURITA, R , NIWA, O , (2016) MICROFLUIDIC PLATFORMS FOR DNA METHYLATION ANALYSIS.LAB ON A CHIP. VOL. 16. ISSUE 19. P. 3631 -3644 | 72 | 51% | 1 |
| 2 | POH, WJ , WEE, CPP , GAO, ZQ , (2016) DNA METHYLTRANSFERASE ACTIVITY ASSAYS: ADVANCES AND CHALLENGES.THERANOSTICS. VOL. 6. ISSUE 3. P. 369 -391 | 69 | 43% | 1 |
| 3 | CLARK, SJ , STATHAM, A , STIRZAKER, C , MOLLOY, PL , FROMMER, M , (2006) DNA METHYLATION: BISULPHITE MODIFICATION AND ANALYSIS.NATURE PROTOCOLS. VOL. 1. ISSUE 5. P. 2353 -2364 | 36 | 84% | 148 |
| 4 | TALEAT, Z , MATHWIG, K , SUDHOLTER, EJR , RASSAEI, L , (2015) DETECTION STRATEGIES FOR METHYLATED AND HYPERMETHYLATED DNA.TRAC-TRENDS IN ANALYTICAL CHEMISTRY. VOL. 66. ISSUE . P. 80 -89 | 50 | 56% | 4 |
| 5 | LI, W , LIU, XJ , HOU, T , LI, HY , LI, F , (2015) ULTRASENSITIVE HOMOGENEOUS ELECTROCHEMICAL STRATEGY FOR DNA METHYLTRANSFERASE ACTIVITY ASSAY BASED ON AUTONOMOUS EXONUCLEASE III-ASSISTED ISOTHERMAL CYCLING SIGNAL AMPLIFICATION.BIOSENSORS & BIOELECTRONICS. VOL. 70. ISSUE . P. 304 -309 | 29 | 71% | 17 |
| 6 | YANG, Y , FAN, MX , GUO, ZH , ZHANG, H , WU, P , CAI, CX , (2014) ELECTROCHEMICAL ANALYSIS FOR DNA METHYLATION.PROGRESS IN CHEMISTRY. VOL. 26. ISSUE 12. P. 1977 -1986 | 48 | 55% | 0 |
| 7 | ZHANG, L , XU, YZ , XIAO, XF , CHEN, J , ZHOU, XQ , ZHU, WY , DAI, Z , ZOU, XY , (2015) DEVELOPMENT OF TECHNIQUES FOR DNA-METHYLATION ANALYSIS.TRAC-TRENDS IN ANALYTICAL CHEMISTRY. VOL. 72. ISSUE . P. 114 -122 | 42 | 59% | 1 |
| 8 | HERNANDEZ, HG , TSE, MY , PANG, SC , ARBOLEDA, H , FORERO, DA , (2013) OPTIMIZING METHODOLOGIES FOR PCR-BASED DNA METHYLATION ANALYSIS.BIOTECHNIQUES. VOL. 55. ISSUE 4. P. 181-197 | 38 | 59% | 28 |
| 9 | WANG, P , CHEN, HB , TIAN, JY , DAI, Z , ZOU, XY , (2013) ELECTROCHEMICAL EVALUATION OF DNA METHYLATION LEVEL BASED ON THE STOICHIOMETRIC RELATIONSHIP BETWEEN PURINE AND PYRIMIDINE BASES.BIOSENSORS & BIOELECTRONICS. VOL. 45. ISSUE . P. 34 -39 | 36 | 65% | 15 |
| 10 | GHABKARI, A , BODOOR, K , HADDAD, Y , ALKHATEEB, A , AL-ABBADI, A , DOWAIRI, M , MAGABLEH, A , BSOUL, N , (2014) DNA HYPERMETHYLATION OF CELL CYCLE (P15 AND P16) AND APOPTOTIC (P14, P53, DAPK AND TMS1) GENES IN PERIPHERAL BLOOD OF LEUKEMIA PATIENTS.ASIAN PACIFIC JOURNAL OF CANCER PREVENTION. VOL. 15. ISSUE 1. P. 75 -84 | 42 | 51% | 17 |
Classes with closest relation at Level 1 |