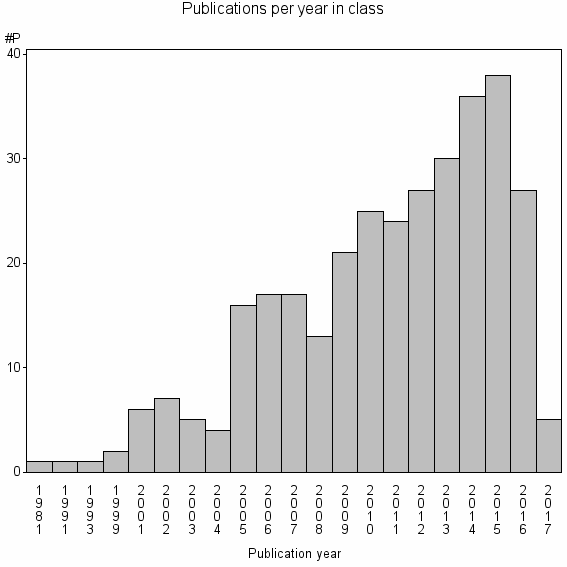
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 24117 | 323 | 44.8 | 85% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 126 | 2 | MOLECULAR BIOLOGY AND EVOLUTION//BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY | 23963 |
| 24117 | 1 | EVOLUTIONARY BIOINFORMAT//FOLD SUPERFAMILY//EVOLUTION OF DOMAIN ARCHITECTURE | 323 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | EVOLUTIONARY BIOINFORMAT | address | 1143868 | 7% | 55% | 22 |
| 2 | FOLD SUPERFAMILY | authKW | 302514 | 1% | 80% | 4 |
| 3 | EVOLUTION OF DOMAIN ARCHITECTURE | authKW | 283608 | 1% | 100% | 3 |
| 4 | MODULAR PROTEIN EVOLUTION | authKW | 283608 | 1% | 100% | 3 |
| 5 | DOMAIN ARCHITECTURE | authKW | 196330 | 3% | 23% | 9 |
| 6 | BIRTH DEATH STOCHASTIC PROCESSES | authKW | 189072 | 1% | 100% | 2 |
| 7 | DOMAIN GRAPH | authKW | 189072 | 1% | 100% | 2 |
| 8 | EPAKTOLOGS | authKW | 189072 | 1% | 100% | 2 |
| 9 | EXPRESSION QUANTITATIVE LOCI | authKW | 189072 | 1% | 100% | 2 |
| 10 | PHENOME GENOME ASSOCIATION | authKW | 189072 | 1% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 5554 | 24% | 0% | 78 |
| 2 | Biochemical Research Methods | 1510 | 23% | 0% | 75 |
| 3 | Biochemistry & Molecular Biology | 1335 | 53% | 0% | 172 |
| 4 | Genetics & Heredity | 1171 | 26% | 0% | 83 |
| 5 | Biotechnology & Applied Microbiology | 985 | 24% | 0% | 78 |
| 6 | Evolutionary Biology | 922 | 11% | 0% | 37 |
| 7 | Biophysics | 65 | 6% | 0% | 20 |
| 8 | Cell Biology | 36 | 7% | 0% | 24 |
| 9 | Biology | 23 | 3% | 0% | 10 |
| 10 | Mathematics, Interdisciplinary Applications | 4 | 1% | 0% | 4 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | EVOLUTIONARY BIOINFORMAT | 1143868 | 7% | 55% | 22 |
| 2 | PROGRAM COMPUTAT ORG SOC | 189072 | 1% | 100% | 2 |
| 3 | UMR 7238 BIOL COMPUTAT QUANTITAT | 189072 | 1% | 100% | 2 |
| 4 | EVOLUTIONARY BIOINFORMAT GRP | 162053 | 2% | 29% | 6 |
| 5 | CHAIR ACTUARIAL MATH | 94536 | 0% | 100% | 1 |
| 6 | CHAIR PROBABIL THEORY MATH STAT | 94536 | 0% | 100% | 1 |
| 7 | DOCTORAL STUDIES INFORMAT ITS PLICAT | 94536 | 0% | 100% | 1 |
| 8 | EVOLUT BIO ERSTAT | 94536 | 0% | 100% | 1 |
| 9 | INRIA RECH BORDEAUX SUD OUEST | 94536 | 0% | 100% | 1 |
| 10 | INSERMU1135ERL IMMUNOL MALAD INFECT CIMI | 94536 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOINFORMATICS | 9073 | 10% | 0% | 32 |
| 2 | CANCER GENOMICS & PROTEOMICS | 7195 | 1% | 2% | 4 |
| 3 | GENES | 5577 | 2% | 1% | 5 |
| 4 | DATABASE-THE JOURNAL OF BIOLOGICAL DATABASES AND CURATION | 5216 | 2% | 1% | 6 |
| 5 | NUCLEIC ACIDS RESEARCH | 4004 | 13% | 0% | 41 |
| 6 | MOLECULAR BIOLOGY AND EVOLUTION | 3308 | 4% | 0% | 14 |
| 7 | BMC BIOINFORMATICS | 3205 | 5% | 0% | 16 |
| 8 | BMC EVOLUTIONARY BIOLOGY | 2824 | 3% | 0% | 10 |
| 9 | GENOME BIOLOGY | 2554 | 3% | 0% | 9 |
| 10 | ARCHAEA-AN INTERNATIONAL MICROBIOLOGICAL JOURNAL | 2551 | 1% | 1% | 2 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | FOLD SUPERFAMILY | 302514 | 1% | 80% | 4 | Search FOLD+SUPERFAMILY | Search FOLD+SUPERFAMILY |
| 2 | EVOLUTION OF DOMAIN ARCHITECTURE | 283608 | 1% | 100% | 3 | Search EVOLUTION+OF+DOMAIN+ARCHITECTURE | Search EVOLUTION+OF+DOMAIN+ARCHITECTURE |
| 3 | MODULAR PROTEIN EVOLUTION | 283608 | 1% | 100% | 3 | Search MODULAR+PROTEIN+EVOLUTION | Search MODULAR+PROTEIN+EVOLUTION |
| 4 | DOMAIN ARCHITECTURE | 196330 | 3% | 23% | 9 | Search DOMAIN+ARCHITECTURE | Search DOMAIN+ARCHITECTURE |
| 5 | BIRTH DEATH STOCHASTIC PROCESSES | 189072 | 1% | 100% | 2 | Search BIRTH+DEATH+STOCHASTIC+PROCESSES | Search BIRTH+DEATH+STOCHASTIC+PROCESSES |
| 6 | DOMAIN GRAPH | 189072 | 1% | 100% | 2 | Search DOMAIN+GRAPH | Search DOMAIN+GRAPH |
| 7 | EPAKTOLOGS | 189072 | 1% | 100% | 2 | Search EPAKTOLOGS | Search EPAKTOLOGS |
| 8 | EXPRESSION QUANTITATIVE LOCI | 189072 | 1% | 100% | 2 | Search EXPRESSION+QUANTITATIVE+LOCI | Search EXPRESSION+QUANTITATIVE+LOCI |
| 9 | PHENOME GENOME ASSOCIATION | 189072 | 1% | 100% | 2 | Search PHENOME+GENOME+ASSOCIATION | Search PHENOME+GENOME+ASSOCIATION |
| 10 | UNCHARACTERIZED PROTEINS | 189072 | 1% | 100% | 2 | Search UNCHARACTERIZED+PROTEINS | Search UNCHARACTERIZED+PROTEINS |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | LINKEVICIUTE, V , RACKHAM, OJL , GOUGH, J , OATES, ME , FANG, H , (2015) FUNCTION-SELECTIVE DOMAIN ARCHITECTURE PLASTICITY POTENTIALS IN EUKARYOTIC GENOME EVOLUTION.BIOCHIMIE. VOL. 119. ISSUE . P. 269 -277 | 27 | 79% | 2 |
| 2 | COHEN-GIHON, I , FONG, JH , SHARAN, R , NUSSINOV, R , PRZYTYCKA, TM , PANCHENKO, AR , (2011) EVOLUTION OF DOMAIN PROMISCUITY IN EUKARYOTIC GENOMES-A PERSPECTIVE FROM THE INFERRED ANCESTRAL DOMAIN ARCHITECTURES.MOLECULAR BIOSYSTEMS. VOL. 7. ISSUE 3. P. 784 -792 | 30 | 73% | 8 |
| 3 | SCAIEWICZ, A , LEVITT, M , (2015) THE LANGUAGE OF THE PROTEIN UNIVERSE.CURRENT OPINION IN GENETICS & DEVELOPMENT. VOL. 35. ISSUE . P. 50 -56 | 32 | 52% | 3 |
| 4 | MOORE, AD , GRATH, S , SCHULER, A , HUYLMANS, AK , BORNBERG-BAUER, E , (2013) QUANTIFICATION AND FUNCTIONAL ANALYSIS OF MODULAR PROTEIN EVOLUTION IN A DENSE PHYLOGENETIC TREE.BIOCHIMICA ET BIOPHYSICA ACTA-PROTEINS AND PROTEOMICS. VOL. 1834. ISSUE 5. P. 898 -907 | 34 | 41% | 10 |
| 5 | NASIR, A , KIM, KM , CAETANO-ANOLLES, G , (2014) GLOBAL PATTERNS OF PROTEIN DOMAIN GAIN AND LOSS IN SUPERKINGDOMS.PLOS COMPUTATIONAL BIOLOGY. VOL. 10. ISSUE 1. P. - | 31 | 38% | 10 |
| 6 | BORNBERG-BAUER, E , ALBA, MM , (2013) DYNAMICS AND ADAPTIVE BENEFITS OF MODULAR PROTEIN EVOLUTION.CURRENT OPINION IN STRUCTURAL BIOLOGY. VOL. 23. ISSUE 3. P. 459-466 | 28 | 41% | 15 |
| 7 | LEES, JG , DAWSON, NL , SILLITOE, I , ORENGO, CA , (2016) FUNCTIONAL INNOVATION FROM CHANGES IN PROTEIN DOMAINS AND THEIR COMBINATIONS.CURRENT OPINION IN STRUCTURAL BIOLOGY. VOL. 38. ISSUE . P. 44 -52 | 25 | 35% | 3 |
| 8 | WANG, ML , CAETANO-ANOLLES, G , (2009) THE EVOLUTIONARY MECHANICS OF DOMAIN ORGANIZATION IN PROTEOMES AND THE RISE OF MODULARITY IN THE PROTEIN WORLD.STRUCTURE. VOL. 17. ISSUE 1. P. 66-78 | 25 | 43% | 32 |
| 9 | HSU, CH , CHEN, CK , HWANG, MJ , (2013) THE ARCHITECTURAL DESIGN OF NETWORKS OF PROTEIN DOMAIN ARCHITECTURES.BIOLOGY LETTERS. VOL. 9. ISSUE 4. P. - | 13 | 81% | 1 |
| 10 | ZMASEK, CM , GODZIK, A , (2012) THIS DEJA VU FEELING-ANALYSIS OF MULTIDOMAIN PROTEIN EVOLUTION IN EUKARYOTIC GENOMES.PLOS COMPUTATIONAL BIOLOGY. VOL. 8. ISSUE 11. P. - | 22 | 47% | 8 |
Classes with closest relation at Level 1 |