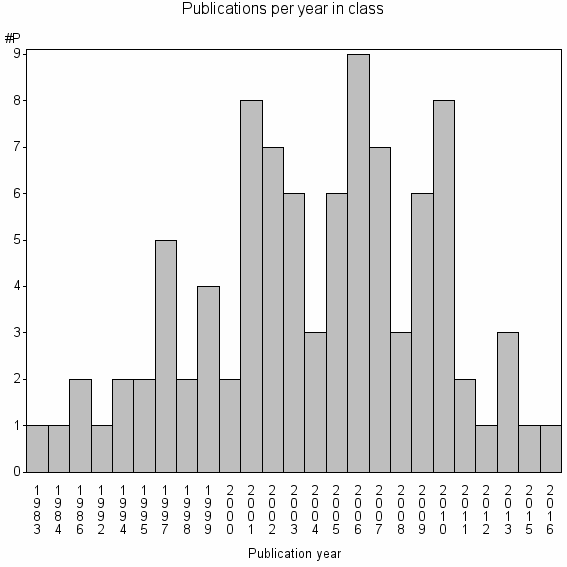
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 35727 | 93 | 27.0 | 71% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 219 | 3 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN FOLDING//PROTEIN STRUCTURE PREDICTION | 50409 |
| 365 | 2 | PROTEIN STRUCTURE PREDICTION//PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PSEUDO AMINO ACID COMPOSITION | 17331 |
| 35727 | 1 | CLOSED LOOPS//GENOME ERS//SKEW PAIRS | 93 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | CLOSED LOOPS | authKW | 3546060 | 19% | 60% | 18 |
| 2 | GENOME ERS | address | 1569580 | 30% | 17% | 28 |
| 3 | SKEW PAIRS | authKW | 1313361 | 4% | 100% | 4 |
| 4 | CONSERVED SEQUENCE STRUCTURE MODULES | authKW | 985021 | 3% | 100% | 3 |
| 5 | MAJOR FOLDS | authKW | 985021 | 3% | 100% | 3 |
| 6 | EARLIEST PROTEINS | authKW | 738764 | 3% | 75% | 3 |
| 7 | ANCIENT BINARY ALPHABET OF PROTEINS | authKW | 656680 | 2% | 100% | 2 |
| 8 | CLASSIFICATION OF LIFE | authKW | 656680 | 2% | 100% | 2 |
| 9 | CODE DEGENERACY | authKW | 656680 | 2% | 100% | 2 |
| 10 | EARLIEST MINI GENES | authKW | 656680 | 2% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biophysics | 585 | 29% | 0% | 27 |
| 2 | Biochemistry & Molecular Biology | 428 | 56% | 0% | 52 |
| 3 | Evolutionary Biology | 187 | 10% | 0% | 9 |
| 4 | Mathematical & Computational Biology | 146 | 8% | 0% | 7 |
| 5 | Genetics & Heredity | 89 | 14% | 0% | 13 |
| 6 | Biochemical Research Methods | 66 | 10% | 0% | 9 |
| 7 | Mathematics, Interdisciplinary Applications | 66 | 6% | 0% | 6 |
| 8 | Biology | 39 | 6% | 0% | 6 |
| 9 | Computer Science, Interdisciplinary Applications | 29 | 5% | 0% | 5 |
| 10 | Biotechnology & Applied Microbiology | 19 | 8% | 0% | 7 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | GENOME ERS | 1569580 | 30% | 17% | 28 |
| 2 | MATH CHEM GRP | 410418 | 5% | 25% | 5 |
| 3 | CNRSUMR 7590CASE 115 | 328340 | 1% | 100% | 1 |
| 4 | EQUIPE BIOL STRUCT | 328340 | 1% | 100% | 1 |
| 5 | EQUIPE SYST MOL BIOL STRUCTURALE | 328340 | 1% | 100% | 1 |
| 6 | STUCTURAL BIOL | 328340 | 1% | 100% | 1 |
| 7 | UMRC7590 | 328340 | 1% | 100% | 1 |
| 8 | EVOLUT | 318893 | 37% | 3% | 34 |
| 9 | EQUIPE SYST MOL BIOL STRUCT | 187620 | 2% | 29% | 2 |
| 10 | CNRS UMRC7590 | 164169 | 1% | 50% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS | 15096 | 13% | 0% | 12 |
| 2 | FOLDING & DESIGN | 7416 | 2% | 1% | 2 |
| 3 | JOURNAL OF MOLECULAR EVOLUTION | 3099 | 6% | 0% | 6 |
| 4 | PROTEIN ENGINEERING | 2320 | 4% | 0% | 4 |
| 5 | STUDIA UNIVERSITATIS BABES-BOLYAI CHEMIA | 1381 | 2% | 0% | 2 |
| 6 | ISRAEL JOURNAL OF ECOLOGY & EVOLUTION | 1333 | 1% | 0% | 1 |
| 7 | ORIGINS OF LIFE AND EVOLUTION OF BIOSPHERES | 1295 | 2% | 0% | 2 |
| 8 | SEMIGROUP FORUM | 1273 | 3% | 0% | 3 |
| 9 | JOURNAL OF MATHEMATICAL CHEMISTRY | 1132 | 3% | 0% | 3 |
| 10 | MATCH-COMMUNICATIONS IN MATHEMATICAL AND IN COMPUTER CHEMISTRY | 879 | 2% | 0% | 2 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CLOSED LOOPS | 3546060 | 19% | 60% | 18 | Search CLOSED+LOOPS | Search CLOSED+LOOPS |
| 2 | SKEW PAIRS | 1313361 | 4% | 100% | 4 | Search SKEW+PAIRS | Search SKEW+PAIRS |
| 3 | CONSERVED SEQUENCE STRUCTURE MODULES | 985021 | 3% | 100% | 3 | Search CONSERVED+SEQUENCE+STRUCTURE+MODULES | Search CONSERVED+SEQUENCE+STRUCTURE+MODULES |
| 4 | MAJOR FOLDS | 985021 | 3% | 100% | 3 | Search MAJOR+FOLDS | Search MAJOR+FOLDS |
| 5 | EARLIEST PROTEINS | 738764 | 3% | 75% | 3 | Search EARLIEST+PROTEINS | Search EARLIEST+PROTEINS |
| 6 | ANCIENT BINARY ALPHABET OF PROTEINS | 656680 | 2% | 100% | 2 | Search ANCIENT+BINARY+ALPHABET+OF+PROTEINS | Search ANCIENT+BINARY+ALPHABET+OF+PROTEINS |
| 7 | CLASSIFICATION OF LIFE | 656680 | 2% | 100% | 2 | Search CLASSIFICATION+OF+LIFE | Search CLASSIFICATION+OF+LIFE |
| 8 | CODE DEGENERACY | 656680 | 2% | 100% | 2 | Search CODE+DEGENERACY | Search CODE+DEGENERACY |
| 9 | EARLIEST MINI GENES | 656680 | 2% | 100% | 2 | Search EARLIEST+MINI+GENES | Search EARLIEST+MINI+GENES |
| 10 | EARLIEST MRNA | 656680 | 2% | 100% | 2 | Search EARLIEST+MRNA | Search EARLIEST+MRNA |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | SOBOLEVSKY, Y , FRENKEL, ZM , TRIFONOV, EN , (2007) COMBINATIONS OF ANCESTRAL MODULES IN PROTEINS.JOURNAL OF MOLECULAR EVOLUTION. VOL. 65. ISSUE 6. P. 640-650 | 15 | 88% | 5 |
| 2 | SOBOLEVSKY, Y , GUIMARAES, RC , TRIFONOV, EN , (2013) TOWARDS FUNCTIONAL REPERTOIRE OF THE EARLIEST PROTEINS.JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS. VOL. 31. ISSUE 11. P. 1293-1300 | 18 | 50% | 3 |
| 3 | TRIFONOV, EN , (2006) EARLY MOLECULAR EVOLUTION.ISRAEL JOURNAL OF ECOLOGY & EVOLUTION. VOL. 52. ISSUE 3-4. P. 375-387 | 16 | 59% | 3 |
| 4 | AHARONOVSKY, E , TRIFONOV, EN , (2005) SEQUENCE STRUCTURE OF VAN DER WAALS LOCKS IN PROTEINS.JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS. VOL. 22. ISSUE 5. P. 545-553 | 9 | 82% | 5 |
| 5 | ROSENFELD, VR , (2007) EMULATING THE FUNCTION OF INTRONS IN PRE-MRNA.MATCH-COMMUNICATIONS IN MATHEMATICAL AND IN COMPUTER CHEMISTRY. VOL. 57. ISSUE 1. P. 135-142 | 6 | 100% | 0 |
| 6 | TRIFONOV, EN , (2009) THE ORIGIN OF THE GENETIC CODE AND OF THE EARLIEST OLIGOPEPTIDES.RESEARCH IN MICROBIOLOGY. VOL. 160. ISSUE 7. P. 481-486 | 13 | 41% | 9 |
| 7 | BEREZOVSKY, IN , TRIFONOV, EN , (2002) FLOWERING BUDS OF GLOBULAR PROTEINS: TRANSPIRING SIMPLICITY OF PROTEIN ORGANIZATION.COMPARATIVE AND FUNCTIONAL GENOMICS. VOL. 3. ISSUE 6. P. 525-534 | 10 | 63% | 13 |
| 8 | FRENKEL, ZM , TRIFONOV, EN , (2007) EVOLUTIONARY NETWORKS IN THE FORMATTED PROTEIN SEQUENCE SPACE.JOURNAL OF COMPUTATIONAL BIOLOGY. VOL. 14. ISSUE 8. P. 1044-1057 | 8 | 67% | 1 |
| 9 | BEREZOVSKY, IN , KIRZHNER, VM , KIRZHNER, A , ROSENFELD, VR , TRIFONOV, EN , (2002) CLOSED LOOPS: PERSISTENCE OF THE PROTEIN CHAIN RETURNS.PROTEIN ENGINEERING. VOL. 15. ISSUE 12. P. 955-957 | 7 | 88% | 5 |
| 10 | FRENKEL, ZM , TRIFONOV, EN , (2008) FROM PROTEIN SEQUENCE SPACE TO ELEMENTARY PROTEIN MODULES.GENE. VOL. 408. ISSUE 1-2. P. 64 -71 | 12 | 41% | 5 |
Classes with closest relation at Level 1 |