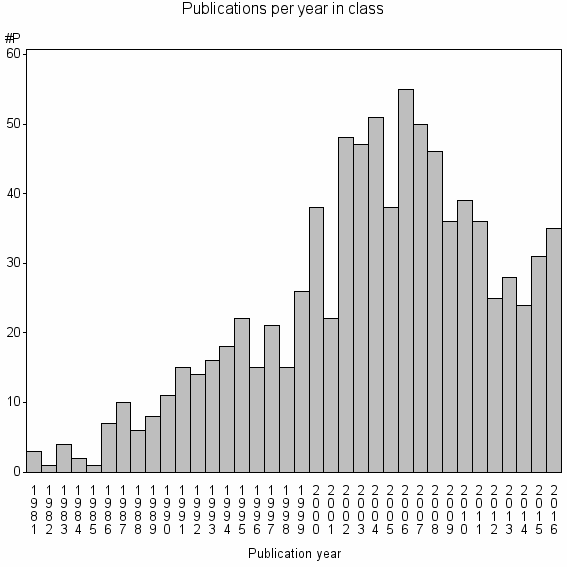
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 13045 | 864 | 43.3 | 83% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 126 | 2 | MOLECULAR BIOLOGY AND EVOLUTION//BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY | 23963 |
| 13045 | 1 | ISOCHORES//EVOLUZ MOL//MALE MUTATION BIAS | 864 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | ISOCHORES | authKW | 2418776 | 14% | 59% | 117 |
| 2 | EVOLUZ MOL | address | 883490 | 4% | 71% | 35 |
| 3 | MALE MUTATION BIAS | authKW | 351319 | 2% | 76% | 13 |
| 4 | BIASED GENE CONVERSION | authKW | 336554 | 2% | 48% | 20 |
| 5 | GC CONTENT | authKW | 321910 | 6% | 17% | 55 |
| 6 | BASE COMPOSITION | authKW | 241996 | 4% | 21% | 33 |
| 7 | MALE DRIVEN EVOLUTION | authKW | 235596 | 1% | 67% | 10 |
| 8 | CHROMOSOMAL BANDS | authKW | 216458 | 1% | 88% | 7 |
| 9 | GC BIASED GENE CONVERSION | authKW | 196327 | 1% | 56% | 10 |
| 10 | WINDOWLESS TECHNIQUE | authKW | 141362 | 0% | 100% | 4 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Genetics & Heredity | 26380 | 72% | 0% | 618 |
| 2 | Evolutionary Biology | 15172 | 28% | 0% | 240 |
| 3 | Biochemistry & Molecular Biology | 2347 | 44% | 0% | 382 |
| 4 | Mathematical & Computational Biology | 954 | 6% | 0% | 55 |
| 5 | Biotechnology & Applied Microbiology | 950 | 15% | 0% | 131 |
| 6 | Biology | 200 | 5% | 0% | 43 |
| 7 | Biochemical Research Methods | 80 | 4% | 0% | 36 |
| 8 | Biophysics | 51 | 4% | 0% | 34 |
| 9 | Cell Biology | 35 | 5% | 0% | 47 |
| 10 | Computer Science, Interdisciplinary Applications | 16 | 2% | 0% | 16 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | EVOLUZ MOL | 883490 | 4% | 71% | 35 |
| 2 | EVOL MOL | 70681 | 0% | 100% | 2 |
| 3 | MOL EVOLUT | 52572 | 3% | 6% | 26 |
| 4 | DIPARTIMENTO BIOL ANIM M LA GRECA | 39754 | 0% | 38% | 3 |
| 5 | SECC BIOMATEMAT | 38470 | 1% | 16% | 7 |
| 6 | ADV STUDIES IND PL MATH | 35340 | 0% | 100% | 1 |
| 7 | BASE LIFE SCI BIOTECHNOL EDUC | 35340 | 0% | 100% | 1 |
| 8 | BIOINFOMAT SYST BIOL MIPS | 35340 | 0% | 100% | 1 |
| 9 | BIOL MED VET SCI | 35340 | 0% | 100% | 1 |
| 10 | BIOMETRIE UA 243 | 35340 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF MOLECULAR EVOLUTION | 53569 | 9% | 2% | 76 |
| 2 | MOLECULAR BIOLOGY AND EVOLUTION | 41581 | 9% | 1% | 81 |
| 3 | GENE | 12531 | 10% | 0% | 83 |
| 4 | GENOME BIOLOGY AND EVOLUTION | 11520 | 2% | 2% | 21 |
| 5 | BIOTECHFORUM ( BTF ) | 7195 | 3% | 1% | 23 |
| 6 | TRENDS IN GENETICS | 5173 | 2% | 1% | 20 |
| 7 | BMC EVOLUTIONARY BIOLOGY | 2362 | 2% | 0% | 15 |
| 8 | GENOME BIOLOGY | 1980 | 2% | 0% | 13 |
| 9 | ANNUAL REVIEW OF GENOMICS AND HUMAN GENETICS | 1754 | 0% | 1% | 4 |
| 10 | GENOMICS | 1679 | 2% | 0% | 20 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ISOCHORES | 2418776 | 14% | 59% | 117 | Search ISOCHORES | Search ISOCHORES |
| 2 | MALE MUTATION BIAS | 351319 | 2% | 76% | 13 | Search MALE+MUTATION+BIAS | Search MALE+MUTATION+BIAS |
| 3 | BIASED GENE CONVERSION | 336554 | 2% | 48% | 20 | Search BIASED+GENE+CONVERSION | Search BIASED+GENE+CONVERSION |
| 4 | GC CONTENT | 321910 | 6% | 17% | 55 | Search GC+CONTENT | Search GC+CONTENT |
| 5 | BASE COMPOSITION | 241996 | 4% | 21% | 33 | Search BASE+COMPOSITION | Search BASE+COMPOSITION |
| 6 | MALE DRIVEN EVOLUTION | 235596 | 1% | 67% | 10 | Search MALE+DRIVEN+EVOLUTION | Search MALE+DRIVEN+EVOLUTION |
| 7 | CHROMOSOMAL BANDS | 216458 | 1% | 88% | 7 | Search CHROMOSOMAL+BANDS | Search CHROMOSOMAL+BANDS |
| 8 | GC BIASED GENE CONVERSION | 196327 | 1% | 56% | 10 | Search GC+BIASED+GENE+CONVERSION | Search GC+BIASED+GENE+CONVERSION |
| 9 | WINDOWLESS TECHNIQUE | 141362 | 0% | 100% | 4 | Search WINDOWLESS+TECHNIQUE | Search WINDOWLESS+TECHNIQUE |
| 10 | COMPOSITIONAL HOMOGENEITY | 133200 | 1% | 54% | 7 | Search COMPOSITIONAL+HOMOGENEITY | Search COMPOSITIONAL+HOMOGENEITY |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | BERNARDI, G , (2000) ISOCHORES AND THE EVOLUTIONARY GENOMICS OF VERTEBRATES.GENE. VOL. 241. ISSUE 1. P. 3 -17 | 61 | 79% | 339 |
| 2 | HODGKINSON, A , EYRE-WALKER, A , (2011) VARIATION IN THE MUTATION RATE ACROSS MAMMALIAN GENOMES.NATURE REVIEWS GENETICS. VOL. 12. ISSUE 11. P. 756 -766 | 58 | 54% | 113 |
| 3 | DURET, L , GALTIER, N , (2009) BIASED GENE CONVERSION AND THE EVOLUTION OF MAMMALIAN GENOMIC LANDSCAPES.ANNUAL REVIEW OF GENOMICS AND HUMAN GENETICS. VOL. 10. ISSUE . P. 285 -311 | 61 | 45% | 230 |
| 4 | BERNARDI, G , (2007) THE NEOSELECTIONIST THEORY OF GENOME EVOLUTION.PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA. VOL. 104. ISSUE 20. P. 8385 -8390 | 45 | 66% | 56 |
| 5 | BERNARDI, G , (2000) THE COMPOSITIONAL EVOLUTION OF VERTEBRATE GENOMES.GENE. VOL. 259. ISSUE 1-2. P. 31 -43 | 46 | 82% | 51 |
| 6 | SCHMEGNER, C , HOEGEL, J , VOGEL, W , ASSUM, G , (2007) THE RATE, NOT THE SPECTRUM, OF BASE PAIR SUBSTITUTIONS CHANGES AT A GC-CONTENT TRANSITION IN THE HUMAN NF1 GENE REGION: IMPLICATIONS FOR THE EVOLUTION OF THE MAMMALIAN GENOME STRUCTURE.GENETICS. VOL. 175. ISSUE 1. P. 421 -428 | 40 | 80% | 4 |
| 7 | COZZI, P , MILANESI, L , BERNARDI, G , (2015) SEGMENTING THE HUMAN GENOME INTO ISOCHORES.EVOLUTIONARY BIOINFORMATICS. VOL. 11. ISSUE . P. 253 -261 | 30 | 86% | 1 |
| 8 | BERNARDI, G , (1995) THE HUMAN GENOME: ORGANIZATION AND EVOLUTIONARY HISTORY.ANNUAL REVIEW OF GENETICS. VOL. 29. ISSUE . P. 445-476 | 57 | 55% | 244 |
| 9 | FUJITA, MK , EDWARDS, SV , PONTING, CP , (2011) THE ANOLIS LIZARD GENOME: AN AMNIOTE GENOME WITHOUT ISOCHORES.GENOME BIOLOGY AND EVOLUTION. VOL. 3. ISSUE . P. 974-984 | 35 | 69% | 15 |
| 10 | COSTANTINI, M , (2015) AN OVERVIEW ON GENOME ORGANIZATION OF MARINE ORGANISMS.MARINE GENOMICS. VOL. 24. ISSUE 1. P. 3 -9 | 27 | 82% | 0 |
Classes with closest relation at Level 1 |