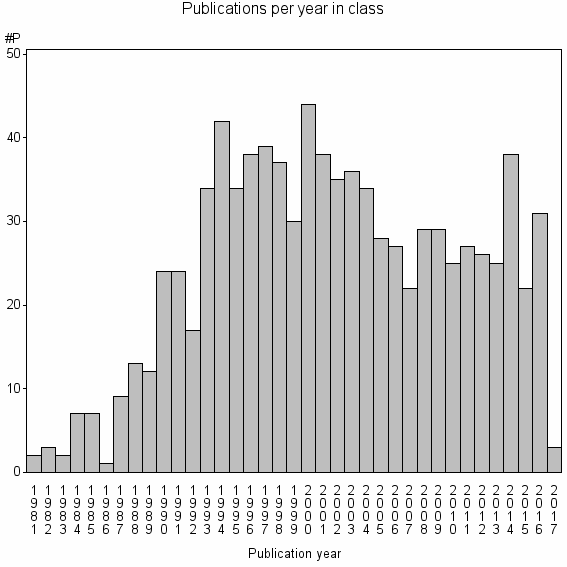
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 12602 | 894 | 45.6 | 86% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 142 | 3 | DNA REPAIR//INTERNATIONAL JOURNAL OF RADIATION BIOLOGY//RADIATION RESEARCH | 64602 |
| 160 | 2 | WERNER SYNDROME//DNA POLYMERASE//DNA REPAIR | 22798 |
| 12602 | 1 | HOLLIDAY JUNCTION//BRANCH MIGRATION//RUVAB | 894 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | HOLLIDAY JUNCTION | authKW | 2576675 | 15% | 56% | 134 |
| 2 | BRANCH MIGRATION | authKW | 719312 | 4% | 57% | 37 |
| 3 | RUVAB | authKW | 341544 | 1% | 100% | 10 |
| 4 | RESOLVING ENZYME | authKW | 273235 | 1% | 100% | 8 |
| 5 | MUS81 | authKW | 242858 | 2% | 44% | 16 |
| 6 | GEN1 | authKW | 239081 | 1% | 100% | 7 |
| 7 | RUVA | authKW | 204926 | 1% | 100% | 6 |
| 8 | FOUR WAY JUNCTION | authKW | 204914 | 1% | 50% | 12 |
| 9 | HOLLIDAY JUNCTION RESOLVASE | authKW | 185949 | 1% | 78% | 7 |
| 10 | RUVABC | authKW | 185949 | 1% | 78% | 7 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 8121 | 76% | 0% | 682 |
| 2 | Cell Biology | 1460 | 22% | 0% | 196 |
| 3 | Biophysics | 499 | 10% | 0% | 85 |
| 4 | Genetics & Heredity | 477 | 11% | 0% | 97 |
| 5 | Microbiology | 156 | 7% | 0% | 62 |
| 6 | Biochemical Research Methods | 29 | 3% | 0% | 26 |
| 7 | Toxicology | 27 | 2% | 0% | 22 |
| 8 | Developmental Biology | 20 | 1% | 0% | 12 |
| 9 | Biology | 15 | 2% | 0% | 17 |
| 10 | Crystallography | 5 | 1% | 0% | 12 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | NUCL ACID STRUCT GRP | 110649 | 1% | 36% | 9 |
| 2 | CLARE HALL S | 80883 | 4% | 6% | 40 |
| 3 | DNA REPLICAT GENOME INTEGR | 61476 | 0% | 60% | 3 |
| 4 | NORDEA HLTH AGING | 60714 | 0% | 44% | 4 |
| 5 | NUCLE ACID STRUCT GRP | 55880 | 1% | 27% | 6 |
| 6 | CANC UK NUCLE ACID STRUCT GRP | 54230 | 1% | 18% | 9 |
| 7 | CANC UK NUCL ACID STRUCT GRP | 49674 | 0% | 36% | 4 |
| 8 | PEDIAT PEDIAT INFECT DIS IMMUN | 49674 | 0% | 36% | 4 |
| 9 | CARCINOGEN SCREENING METHODS | 45538 | 0% | 67% | 2 |
| 10 | MICROBIAL DIS MOL MICROBIOL | 45538 | 0% | 67% | 2 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF MOLECULAR BIOLOGY | 14622 | 11% | 0% | 100 |
| 2 | NUCLEIC ACIDS RESEARCH | 5187 | 9% | 0% | 78 |
| 3 | DNA REPAIR | 3754 | 2% | 1% | 15 |
| 4 | EMBO JOURNAL | 3147 | 4% | 0% | 40 |
| 5 | BIOCHEMISTRY | 2011 | 6% | 0% | 57 |
| 6 | CELL | 1391 | 3% | 0% | 25 |
| 7 | MOLECULAR CELL | 1384 | 2% | 0% | 15 |
| 8 | JOURNAL OF BIOLOGICAL CHEMISTRY | 1207 | 9% | 0% | 76 |
| 9 | GENES TO CELLS | 1129 | 1% | 0% | 8 |
| 10 | MOLECULAR MICROBIOLOGY | 1118 | 2% | 0% | 20 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | HOLLIDAY JUNCTION | 2576675 | 15% | 56% | 134 | Search HOLLIDAY+JUNCTION | Search HOLLIDAY+JUNCTION |
| 2 | BRANCH MIGRATION | 719312 | 4% | 57% | 37 | Search BRANCH+MIGRATION | Search BRANCH+MIGRATION |
| 3 | RUVAB | 341544 | 1% | 100% | 10 | Search RUVAB | Search RUVAB |
| 4 | RESOLVING ENZYME | 273235 | 1% | 100% | 8 | Search RESOLVING+ENZYME | Search RESOLVING+ENZYME |
| 5 | MUS81 | 242858 | 2% | 44% | 16 | Search MUS81 | Search MUS81 |
| 6 | GEN1 | 239081 | 1% | 100% | 7 | Search GEN1 | Search GEN1 |
| 7 | RUVA | 204926 | 1% | 100% | 6 | Search RUVA | Search RUVA |
| 8 | FOUR WAY JUNCTION | 204914 | 1% | 50% | 12 | Search FOUR+WAY+JUNCTION | Search FOUR+WAY+JUNCTION |
| 9 | HOLLIDAY JUNCTION RESOLVASE | 185949 | 1% | 78% | 7 | Search HOLLIDAY+JUNCTION+RESOLVASE | Search HOLLIDAY+JUNCTION+RESOLVASE |
| 10 | RUVABC | 185949 | 1% | 78% | 7 | Search RUVABC | Search RUVABC |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | WYATT, HDM , WEST, SC , (2014) HOLLIDAY JUNCTION RESOLVASES.COLD SPRING HARBOR PERSPECTIVES IN BIOLOGY. VOL. 6. ISSUE 9. P. - | 174 | 84% | 19 |
| 2 | LILLEY, DMJ , (2000) STRUCTURES OF HELICAL JUNCTIONS IN NUCLEIC ACIDS.QUARTERLY REVIEWS OF BIOPHYSICS. VOL. 33. ISSUE 2. P. 109 -159 | 122 | 61% | 203 |
| 3 | WEST, SC , (1997) PROCESSING OF RECOMBINATION INTERMEDIATES BY THE RUVABC PROTEINS.ANNUAL REVIEW OF GENETICS. VOL. 31. ISSUE . P. 213 -244 | 110 | 60% | 314 |
| 4 | LILLEY, DMJ , WHITE, MF , (2001) THE JUNCTION-RESOLVING ENZYMES.NATURE REVIEWS MOLECULAR CELL BIOLOGY. VOL. 2. ISSUE 6. P. 433 -443 | 70 | 84% | 95 |
| 5 | WHITE, MF , GIRAUDPANIS, MJE , POHLER, JRG , LILLEY, DMJ , (1997) RECOGNITION AND MANIPULATION OF BRANCHED DNA STRUCTURE BY JUNCTION-RESOLVING ENZYMES.JOURNAL OF MOLECULAR BIOLOGY. VOL. 269. ISSUE 5. P. 647-664 | 93 | 66% | 67 |
| 6 | PRIVEZENTZEV, CV , KEELEY, A , SIGALA, B , TSANEVA, IR , (2005) THE ROLE OF RUVA OCTAMERIZATION FOR RUVAB FUNCTION IN VITRO AND IN VIVO.JOURNAL OF BIOLOGICAL CHEMISTRY. VOL. 280. ISSUE 5. P. 3365 -3375 | 53 | 95% | 19 |
| 7 | CHAN, YW , WEST, S , (2015) GEN1 PROMOTES HOLLIDAY JUNCTION RESOLUTION BY A COORDINATED NICK AND COUNTER-NICK MECHANISM.NUCLEIC ACIDS RESEARCH. VOL. 43. ISSUE 22. P. - | 43 | 93% | 0 |
| 8 | BLANCO, MG , MATOS, J , (2015) HOLD YOUR HORSSES: CONTROLLING STRUCTURE-SELECTIVE ENDONUCLEASES MUS81 AND YEN1/GEN1.FRONTIERS IN GENETICS. VOL. 6. ISSUE . P. - | 58 | 63% | 4 |
| 9 | PUNATAR, RS , MARTIN, MJ , WYATT, HDM , CHAN, YW , WEST, SC , (2017) RESOLUTION OF SINGLE AND DOUBLE HOLLIDAY JUNCTION RECOMBINATION INTERMEDIATES BY GEN1.PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA. VOL. 114. ISSUE 3. P. 443 -450 | 47 | 76% | 0 |
| 10 | ABD WAHAB, S , CHOI, M , BIANCO, PR , (2013) CHARACTERIZATION OF THE ATPASE ACTIVITY OF RECG AND RUVAB PROTEINS ON MODEL FORK STRUCTURES REVEALS INSIGHT INTO STALLED DNA REPLICATION FORK REPAIR.JOURNAL OF BIOLOGICAL CHEMISTRY. VOL. 288. ISSUE 37. P. 26397-26409 | 48 | 72% | 12 |
Classes with closest relation at Level 1 |
| Rank | Class id | link |
|---|---|---|
| 1 | 8496 | RECBCD//RECBCD ENZYME//RECOMBINEERING |
| 2 | 18097 | GENE UVSX//BACTERIOPHAGE T4//UVSX |
| 3 | 4671 | RAD51//RAD52//HOMOLOGOUS RECOMBINATION |
| 4 | 19522 | REPLICATION FORK BARRIER//FORK ARREST//REPLICATION TERMINATOR PROTEIN |
| 5 | 21108 | FEN1//BRAUN S//CREAT INITIAT CELL CYCLE CONTROL |
| 6 | 4991 | RECA PROTEIN//RECA//LEXA REPRESSOR |
| 7 | 27003 | REPTIN//R2TP//TIP49 |
| 8 | 13276 | UVRB//UVRD//UVRABC |
| 9 | 3755 | WERNER SYNDROME//BLOOM SYNDROME//RECQ HELICASE |
| 10 | 4863 | CLAMP LOADER//REPLICATION FACTOR C//PRIMOSOME |