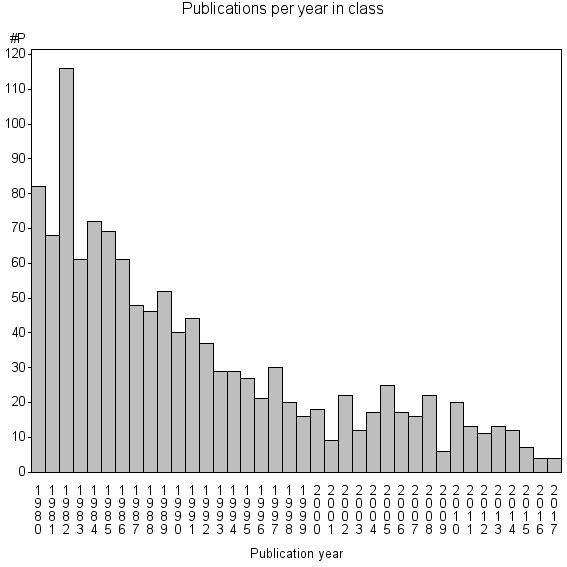
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 3633 | 1216 | 31.4 | 44% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 142 | 3 | DNA REPAIR//INTERNATIONAL JOURNAL OF RADIATION BIOLOGY//RADIATION RESEARCH | 64602 |
| 3633 | 2 | GENE POINT MUTATIONS//HETEROKARYON 12//AD 3A LOCUS | 1216 |
| 21237 | 1 | SNM1//PSORALEN SENSITIVITY//RNR3 LACZ | 427 |
| 26253 | 1 | GENE POINT MUTATIONS//HETEROKARYON 12//AD 3A LOCUS | 259 |
| 27930 | 1 | FLUOROVISUALIZATION//FLASH WAVE LIGHT//MOUSE OSTEOSARCOMA | 217 |
| 32541 | 1 | 3 DIMENSIONAL NEUROIMAGING//FUNCTIONAL STATE OF THE ORGANISM//FUNCTIONAL STATE OF THE ORGANISM FSO | 137 |
| 35601 | 1 | BASIC BLUE 141//INOSITOL DEHYDROGENASE//BASIC BLUE 54 | 94 |
| 36346 | 1 | METAPHENOXY BENZYL CHLORIDE//MUTATION RESEARCH//GENOTOXICITY ASSAY | 82 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | GENE POINT MUTATIONS | authKW | 406774 | 1% | 90% | 18 |
| 2 | HETEROKARYON 12 | authKW | 403148 | 1% | 94% | 17 |
| 3 | AD 3A LOCUS | authKW | 401755 | 1% | 100% | 16 |
| 4 | AD 3B LOCUS | authKW | 401755 | 1% | 100% | 16 |
| 5 | AD 3 REGION | authKW | 376645 | 1% | 100% | 15 |
| 6 | MULTIPLE LOCUS MUTATIONS | authKW | 376645 | 1% | 100% | 15 |
| 7 | MULTILOCUS DELETION MUTATIONS | authKW | 301316 | 1% | 100% | 12 |
| 8 | RECESSIVE LETHAL MUTATION | authKW | 249614 | 1% | 76% | 13 |
| 9 | SNM1 | authKW | 133913 | 1% | 67% | 8 |
| 10 | 2 COMPONENT HETEROKARYONS | authKW | 129134 | 0% | 86% | 6 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Genetics & Heredity | 8952 | 36% | 0% | 437 |
| 2 | Toxicology | 6095 | 22% | 0% | 262 |
| 3 | CYTOLOGY & HISTOLOGY | 1878 | 2% | 0% | 24 |
| 4 | Biochemistry & Molecular Biology | 816 | 25% | 0% | 301 |
| 5 | Biophysics | 643 | 9% | 0% | 113 |
| 6 | Biology | 598 | 7% | 0% | 84 |
| 7 | Microbiology | 344 | 8% | 0% | 102 |
| 8 | Mycology | 344 | 2% | 0% | 28 |
| 9 | Biotechnology & Applied Microbiology | 228 | 7% | 0% | 88 |
| 10 | Cell Biology | 159 | 8% | 0% | 96 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ABT BIOL MED MIKROBIOL | 25110 | 0% | 100% | 1 |
| 2 | AG MOL BIOENGN | 25110 | 0% | 100% | 1 |
| 3 | AUSENSTELLE BONN | 25110 | 0% | 100% | 1 |
| 4 | BACTERIAS LACTICAS PROBIOTICOS | 25110 | 0% | 100% | 1 |
| 5 | BIOCIENCIAS FISIOL FARMACOL BIOFIS | 25110 | 0% | 100% | 1 |
| 6 | CAMPUS VALE 210 | 25110 | 0% | 100% | 1 |
| 7 | CANC BIOL RADIAL ONCOL | 25110 | 0% | 100% | 1 |
| 8 | CELL DIFFERECTIAT | 25110 | 0% | 100% | 1 |
| 9 | CIENCIAS FARMACUET | 25110 | 0% | 100% | 1 |
| 10 | DIPARTAMENTO BIOFIS | 25110 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MUTATION RESEARCH | 92751 | 14% | 2% | 173 |
| 2 | CURRENT GENETICS | 19125 | 4% | 1% | 53 |
| 3 | MUTATION RESEARCH-DNA REPAIR | 8688 | 1% | 3% | 13 |
| 4 | MOLECULAR AND GENERAL GENETICS | 8313 | 3% | 1% | 40 |
| 5 | MOLECULAR & GENERAL GENETICS | 4529 | 2% | 1% | 21 |
| 6 | TSITOLOGIYA | 3046 | 1% | 1% | 17 |
| 7 | REVISTA BRASILEIRA DE GENETICA | 2554 | 1% | 1% | 9 |
| 8 | PHOTOCHEMISTRY AND PHOTOBIOLOGY | 2481 | 2% | 0% | 29 |
| 9 | BIOLOGISKE SKRIFTER KONGELIGE DANSKE VIDENSKABERNES SELSKAB | 2091 | 0% | 8% | 1 |
| 10 | MUTATION RESEARCH-REVIEWS IN MUTATION RESEARCH | 1997 | 1% | 1% | 7 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | GENE POINT MUTATIONS | 406774 | 1% | 90% | 18 | Search GENE+POINT+MUTATIONS | Search GENE+POINT+MUTATIONS |
| 2 | HETEROKARYON 12 | 403148 | 1% | 94% | 17 | Search HETEROKARYON+12 | Search HETEROKARYON+12 |
| 3 | AD 3A LOCUS | 401755 | 1% | 100% | 16 | Search AD+3A+LOCUS | Search AD+3A+LOCUS |
| 4 | AD 3B LOCUS | 401755 | 1% | 100% | 16 | Search AD+3B+LOCUS | Search AD+3B+LOCUS |
| 5 | AD 3 REGION | 376645 | 1% | 100% | 15 | Search AD+3+REGION | Search AD+3+REGION |
| 6 | MULTIPLE LOCUS MUTATIONS | 376645 | 1% | 100% | 15 | Search MULTIPLE+LOCUS+MUTATIONS | Search MULTIPLE+LOCUS+MUTATIONS |
| 7 | MULTILOCUS DELETION MUTATIONS | 301316 | 1% | 100% | 12 | Search MULTILOCUS+DELETION+MUTATIONS | Search MULTILOCUS+DELETION+MUTATIONS |
| 8 | RECESSIVE LETHAL MUTATION | 249614 | 1% | 76% | 13 | Search RECESSIVE+LETHAL+MUTATION | Search RECESSIVE+LETHAL+MUTATION |
| 9 | SNM1 | 133913 | 1% | 67% | 8 | Search SNM1 | Search SNM1 |
| 10 | 2 COMPONENT HETEROKARYONS | 129134 | 0% | 86% | 6 | Search 2+COMPONENT+HETEROKARYONS | Search 2+COMPONENT+HETEROKARYONS |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | INOUE, H , (1999) DNA REPAIR AND SPECIFIC-LOCUS MUTAGENESIS IN NEUROSPORA CRASSA.MUTATION RESEARCH-REVIEWS IN MUTATION RESEARCH. VOL. 437. ISSUE 2. P. 121-133 | 29 | 85% | 7 |
| 2 | KUSUZAKI, K , HOSOGI, S , ASHIHARA, E , MATSUBARA, T , SATONAKA, H , NAKAMURA, T , MATSUMINE, A , SUDO, A , UCHIDA, A , MURATA, H , ET AL (2012) TRANSLATIONAL RESEARCH OF PHOTODYNAMIC THERAPY WITH ACRIDINE ORANGE WHICH TARGETS CANCER ACIDITY.CURRENT PHARMACEUTICAL DESIGN. VOL. 18. ISSUE 10. P. 1414 -1420 | 26 | 63% | 1 |
| 3 | HORI, H , TERAMOTO, Y , FUKUYAMA, Y , MARUO, T , (2014) MARGINAL RESECTION AND ACRIDINE ORANGE PHOTODYNAMIC THERAPY IN A CAT WITH RECURRENT CUTANEOUS MALIGNANT MELANOMA.INTERNATIONAL JOURNAL OF APPLIED RESEARCH IN VETERINARY MEDICINE. VOL. 12. ISSUE 3. P. 181 -185 | 15 | 100% | 0 |
| 4 | FRIEDBERG, EC , (1988) DEOXYRIBONUCLEIC-ACID REPAIR IN THE YEAST SACCHAROMYCES-CEREVISIAE.MICROBIOLOGICAL REVIEWS. VOL. 52. ISSUE 1. P. 70-102 | 64 | 38% | 308 |
| 5 | MUNARI, FM , REVERS, LF , CARDONE, JM , IMMICH, BF , MOURA, DJ , GUECHEVA, TN , BONATTO, D , LAURINO, JP , SAFFI, J , BRENDEL, M , ET AL (2014) SAK1 KINASE INTERACTS WITH PSO2 NUCLEASE IN RESPONSE TO DNA DAMAGE INDUCED BY INTERSTRAND CROSSLINK-INDUCING AGENTS IN SACCHAROMYCES CEREVISIAE.JOURNAL OF PHOTOCHEMISTRY AND PHOTOBIOLOGY B-BIOLOGY. VOL. 130. ISSUE . P. 241-253 | 32 | 43% | 1 |
| 6 | MUNARI, FM , GUECHEVA, TN , BONATTO, D , HENRIQUES, JAP , (2013) NEW FEATURES ON PSO2 PROTEIN FAMILY IN DNA INTERSTRAND CROSS-LINK REPAIR AND IN THE MAINTENANCE OF GENOMIC INTEGRITY IN SACCHAROMYCES CEREVISIAE.FUNGAL GENETICS AND BIOLOGY. VOL. 60. ISSUE . P. 122-132 | 41 | 34% | 2 |
| 7 | MOUSTACCHI, E , (1987) DNA-REPAIR IN YEAST - GENETIC-CONTROL AND BIOLOGICAL CONSEQUENCES.ADVANCES IN RADIATION BIOLOGY. VOL. 13. ISSUE . P. 1-30 | 56 | 53% | 22 |
| 8 | CATTELL, E , SENGEROVA, B , MCHUGH, PJ , (2010) THE SNM1/PSO2 FAMILY OF ICL REPAIR NUCLEASES: FROM YEAST TO MAN.ENVIRONMENTAL AND MOLECULAR MUTAGENESIS. VOL. 51. ISSUE 6. P. 635-645 | 34 | 37% | 20 |
| 9 | WOLTER, R , SIEDE, W , BRENDEL, M , (1996) REGULATION OF SNM1, AN INDUCIBLE SACCHAROMYCES CEREVISIAE GENE REQUIRED FOR REPAIR OF DNA CROSS-LINKS.MOLECULAR AND GENERAL GENETICS. VOL. 250. ISSUE 2. P. 162 -168 | 27 | 66% | 35 |
| 10 | HENRIQUES, JAP , BRENDEL, M , (1990) THE ROLE OF PSO AND SNM GENES IN DNA-REPAIR OF THE YEAST SACCHAROMYCES-CEREVISIAE.CURRENT GENETICS. VOL. 18. ISSUE 5. P. 387-393 | 34 | 61% | 16 |
Classes with closest relation at Level 2 |