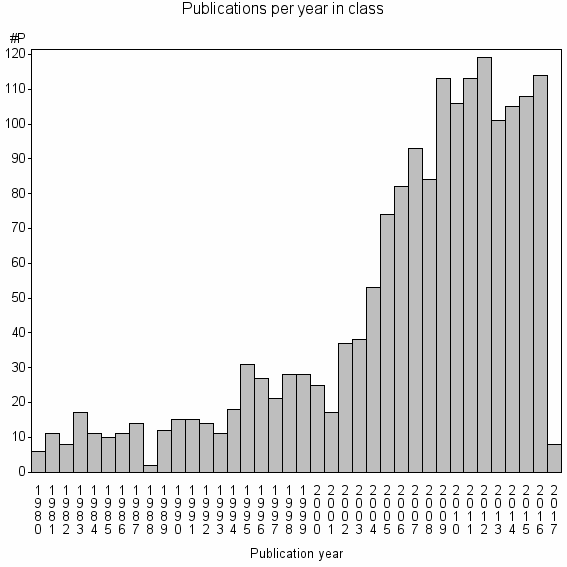
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 4939 | 1700 | 50.1 | 82% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 219 | 3 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN FOLDING//PROTEIN STRUCTURE PREDICTION | 50409 |
| 260 | 2 | PROTEIN FOLDING//JOURNAL OF CHEMICAL THEORY AND COMPUTATION//JOURNAL OF COMPUTATIONAL CHEMISTRY | 19582 |
| 4939 | 1 | ELASTIC NETWORK MODEL//NORMAL MODE ANALYSIS//GAUSSIAN NETWORK MODEL | 1700 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | ELASTIC NETWORK MODEL | authKW | 879085 | 4% | 64% | 76 |
| 2 | NORMAL MODE ANALYSIS | authKW | 552957 | 6% | 29% | 105 |
| 3 | GAUSSIAN NETWORK MODEL | authKW | 335239 | 2% | 67% | 28 |
| 4 | LH BAKER BIOINFORMAT BIOL STAT | address | 183989 | 1% | 41% | 25 |
| 5 | ELASTIC NETWORK | authKW | 146664 | 1% | 58% | 14 |
| 6 | PROTEIN STRUCTURE NETWORKS | authKW | 140796 | 1% | 56% | 14 |
| 7 | B FACTORS | authKW | 139312 | 1% | 48% | 16 |
| 8 | SACKLER MOL MED | address | 113925 | 3% | 14% | 47 |
| 9 | MOL STRUCT SECT | address | 97897 | 1% | 42% | 13 |
| 10 | PROTEIN DYNAMICS | authKW | 90102 | 5% | 6% | 79 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biophysics | 12407 | 31% | 0% | 530 |
| 2 | Mathematical & Computational Biology | 6530 | 12% | 0% | 197 |
| 3 | Biochemistry & Molecular Biology | 5473 | 48% | 0% | 810 |
| 4 | Biochemical Research Methods | 1980 | 12% | 0% | 206 |
| 5 | Physics, Atomic, Molecular & Chemical | 833 | 10% | 0% | 168 |
| 6 | Computer Science, Interdisciplinary Applications | 526 | 5% | 0% | 91 |
| 7 | Chemistry, Physical | 496 | 14% | 0% | 242 |
| 8 | Crystallography | 254 | 4% | 0% | 67 |
| 9 | Biotechnology & Applied Microbiology | 144 | 5% | 0% | 92 |
| 10 | Chemistry, Multidisciplinary | 115 | 8% | 0% | 144 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | LH BAKER BIOINFORMAT BIOL STAT | 183989 | 1% | 41% | 25 |
| 2 | SACKLER MOL MED | 113925 | 3% | 14% | 47 |
| 3 | MOL STRUCT SECT | 97897 | 1% | 42% | 13 |
| 4 | HUMAN GENET MOL MED | 72080 | 3% | 9% | 45 |
| 5 | BAKER BIOINFORMAT BIOL SCI | 71841 | 0% | 100% | 4 |
| 6 | BIOINFORMAT THEO BIOL GRP | 35921 | 0% | 100% | 2 |
| 7 | COMPUTAT PHARMACEUT CHEM GRP | 35921 | 0% | 100% | 2 |
| 8 | L H BAKER BIOINFORMAT BIOL STAT | 35921 | 0% | 100% | 2 |
| 9 | ITP SB | 35917 | 0% | 50% | 4 |
| 10 | PROGRAM BIOINFORMAT COMPUTAT BIOL | 35178 | 1% | 14% | 14 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS | 64125 | 9% | 2% | 151 |
| 2 | PHYSICAL BIOLOGY | 16931 | 2% | 3% | 28 |
| 3 | BIOPHYSICAL JOURNAL | 14996 | 8% | 1% | 128 |
| 4 | PROTEINS-STRUCTURE FUNCTION AND GENETICS | 11658 | 1% | 4% | 17 |
| 5 | PLOS COMPUTATIONAL BIOLOGY | 10837 | 3% | 1% | 53 |
| 6 | JOURNAL OF CHEMICAL THEORY AND COMPUTATION | 9323 | 3% | 1% | 49 |
| 7 | JOURNAL OF MOLECULAR GRAPHICS & MODELLING | 7499 | 2% | 2% | 28 |
| 8 | STRUCTURE | 5745 | 2% | 1% | 35 |
| 9 | JOURNAL OF COMPUTATIONAL CHEMISTRY | 5615 | 3% | 1% | 44 |
| 10 | BMC STRUCTURAL BIOLOGY | 5139 | 1% | 2% | 12 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ELASTIC NETWORK MODEL | 879085 | 4% | 64% | 76 | Search ELASTIC+NETWORK+MODEL | Search ELASTIC+NETWORK+MODEL |
| 2 | NORMAL MODE ANALYSIS | 552957 | 6% | 29% | 105 | Search NORMAL+MODE+ANALYSIS | Search NORMAL+MODE+ANALYSIS |
| 3 | GAUSSIAN NETWORK MODEL | 335239 | 2% | 67% | 28 | Search GAUSSIAN+NETWORK+MODEL | Search GAUSSIAN+NETWORK+MODEL |
| 4 | ELASTIC NETWORK | 146664 | 1% | 58% | 14 | Search ELASTIC+NETWORK | Search ELASTIC+NETWORK |
| 5 | PROTEIN STRUCTURE NETWORKS | 140796 | 1% | 56% | 14 | Search PROTEIN+STRUCTURE+NETWORKS | Search PROTEIN+STRUCTURE+NETWORKS |
| 6 | B FACTORS | 139312 | 1% | 48% | 16 | Search B+FACTORS | Search B+FACTORS |
| 7 | PROTEIN DYNAMICS | 90102 | 5% | 6% | 79 | Search PROTEIN+DYNAMICS | Search PROTEIN+DYNAMICS |
| 8 | HINGE BENDING | 84297 | 1% | 36% | 13 | Search HINGE+BENDING | Search HINGE+BENDING |
| 9 | NETWORK RIGIDITY | 80818 | 0% | 75% | 6 | Search NETWORK+RIGIDITY | Search NETWORK+RIGIDITY |
| 10 | MEAN SQUARE FLUCTUATIONS | 74833 | 0% | 83% | 5 | Search MEAN+SQUARE+FLUCTUATIONS | Search MEAN+SQUARE+FLUCTUATIONS |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | FUGLEBAKK, E , TIWARI, SP , REUTER, N , (2015) COMPARING THE INTRINSIC DYNAMICS OF MULTIPLE PROTEIN STRUCTURES USING ELASTIC NETWORK MODELS.BIOCHIMICA ET BIOPHYSICA ACTA-GENERAL SUBJECTS. VOL. 1850. ISSUE 5. P. 911 -922 | 97 | 82% | 13 |
| 2 | MAHAJAN, S , SANEJOUAND, YH , (2015) ON THE RELATIONSHIP BETWEEN LOW-FREQUENCY NORMAL MODES AND THE LARGE-SCALE CONFORMATIONAL CHANGES OF PROTEINS.ARCHIVES OF BIOCHEMISTRY AND BIOPHYSICS. VOL. 567. ISSUE . P. 59 -65 | 111 | 70% | 8 |
| 3 | BAHAR, I , LEZON, TR , YANG, LW , EYAL, E , (2010) GLOBAL DYNAMICS OF PROTEINS: BRIDGING BETWEEN STRUCTURE AND FUNCTION.ANNUAL REVIEW OF BIOPHYSICS, VOL 39. VOL. 39. ISSUE . P. 23-42 | 60 | 85% | 213 |
| 4 | BASTOLLA, U , (2014) COMPUTING PROTEIN DYNAMICS FROM PROTEIN STRUCTURE WITH ELASTIC NETWORK MODELS.WILEY INTERDISCIPLINARY REVIEWS-COMPUTATIONAL MOLECULAR SCIENCE. VOL. 4. ISSUE 5. P. 488 -503 | 79 | 80% | 6 |
| 5 | SKJAERVEN, L , HOLLUP, SM , REUTER, N , (2009) NORMAL MODE ANALYSIS FOR PROTEINS.JOURNAL OF MOLECULAR STRUCTURE-THEOCHEM. VOL. 898. ISSUE 1-3. P. 42-48 | 79 | 83% | 37 |
| 6 | BAHAR, I , RADER, AJ , (2005) COARSE-GRAINED NORMAL MODE ANALYSIS IN STRUCTURAL BIOLOGY.CURRENT OPINION IN STRUCTURAL BIOLOGY. VOL. 15. ISSUE 5. P. 586-592 | 52 | 83% | 405 |
| 7 | LOPEZ-BLANCO, JR , CHACON, P , (2016) NEW GENERATION OF ELASTIC NETWORK MODELS.CURRENT OPINION IN STRUCTURAL BIOLOGY. VOL. 37. ISSUE . P. 46 -53 | 61 | 71% | 2 |
| 8 | TIWARI, SP , FUGLEBAKK, E , HOLLUP, SM , SKJAERVEN, L , CRAGNOLINI, T , GRINDHAUG, SH , TEKLE, KM , REUTER, N , (2014) WEBNM@ V2.0: WEB SERVER AND SERVICES FOR COMPARING PROTEIN FLEXIBILITY.BMC BIOINFORMATICS. VOL. 15. ISSUE . P. - | 58 | 81% | 6 |
| 9 | MICHELETTI, C , (2013) COMPARING PROTEINS BY THEIR INTERNAL DYNAMICS: EXPLORING STRUCTURE-FUNCTION RELATIONSHIPS BEYOND STATIC STRUCTURAL ALIGNMENTS.PHYSICS OF LIFE REVIEWS. VOL. 10. ISSUE 1. P. 1 -26 | 81 | 52% | 31 |
| 10 | BAHAR, I , LEZON, TR , BAKAN, A , SHRIVASTAVA, IH , (2010) NORMAL MODE ANALYSIS OF BIOMOLECULAR STRUCTURES: FUNCTIONAL MECHANISMS OF MEMBRANE PROTEINS.CHEMICAL REVIEWS. VOL. 110. ISSUE 3. P. 1463 -1497 | 114 | 31% | 170 |
Classes with closest relation at Level 1 |