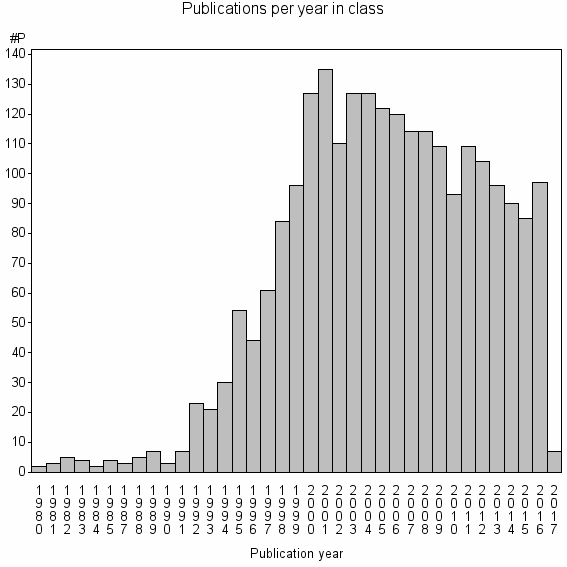
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 2138 | 2344 | 49.9 | 85% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 5 | 4 | CHEMISTRY, ORGANIC//CHEMISTRY, INORGANIC & NUCLEAR//CHEMISTRY, MULTIDISCIPLINARY | 1745167 |
| 189 | 3 | JOURNAL OF MAGNETIC RESONANCE//MAGNETIC RESONANCE IN MEDICINE//JOURNAL OF BIOMOLECULAR NMR | 54843 |
| 866 | 2 | JOURNAL OF BIOMOLECULAR NMR//JOURNAL OF MAGNETIC RESONANCE//RESIDUAL DIPOLAR COUPLINGS | 11581 |
| 2138 | 1 | RESIDUAL DIPOLAR COUPLINGS//JOURNAL OF BIOMOLECULAR NMR//PSEUDOCONTACT SHIFTS | 2344 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | RESIDUAL DIPOLAR COUPLINGS | authKW | 1329603 | 8% | 52% | 198 |
| 2 | JOURNAL OF BIOMOLECULAR NMR | journal | 914151 | 18% | 16% | 432 |
| 3 | PSEUDOCONTACT SHIFTS | authKW | 435531 | 2% | 63% | 53 |
| 4 | N 15 RELAXATION | authKW | 344478 | 2% | 58% | 46 |
| 5 | CROSS CORRELATED RELAXATION | authKW | 272603 | 1% | 70% | 30 |
| 6 | PROTEIN DYNAMICS | authKW | 235701 | 6% | 12% | 150 |
| 7 | DIPOLAR COUPLING | authKW | 228906 | 3% | 26% | 68 |
| 8 | RELAXATION DISPERSION | authKW | 209908 | 1% | 46% | 35 |
| 9 | RDC | authKW | 188175 | 1% | 43% | 34 |
| 10 | ALIGNMENT TENSOR | authKW | 188075 | 1% | 76% | 19 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Spectroscopy | 29306 | 31% | 0% | 721 |
| 2 | Biochemistry & Molecular Biology | 6617 | 45% | 0% | 1054 |
| 3 | Physics, Atomic, Molecular & Chemical | 4025 | 17% | 0% | 404 |
| 4 | Chemistry, Multidisciplinary | 3974 | 28% | 0% | 660 |
| 5 | Biochemical Research Methods | 2210 | 11% | 0% | 258 |
| 6 | Biophysics | 752 | 8% | 0% | 176 |
| 7 | Chemistry, Physical | 471 | 12% | 0% | 292 |
| 8 | Mathematical & Computational Biology | 48 | 1% | 0% | 27 |
| 9 | Cell Biology | 9 | 3% | 0% | 77 |
| 10 | Chemistry, Medicinal | -0 | 1% | 0% | 17 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | PROT ENGN NETWORK EXCELLENCE | 110613 | 2% | 18% | 46 |
| 2 | NMR BASED STRUCT BIOL | 102561 | 2% | 15% | 54 |
| 3 | STRUCT MOL BIOL UNIT | 83716 | 1% | 43% | 15 |
| 4 | CHEM BIOPHYS | 82983 | 1% | 28% | 23 |
| 5 | BIOMOL STRUCT ORG | 44674 | 1% | 16% | 22 |
| 6 | BIOL STRUCT JEAN PIERRE EBEL | 43069 | 2% | 9% | 38 |
| 7 | PROGRAM MOL STRUCT FUNCT | 34894 | 1% | 10% | 26 |
| 8 | UJF UMR 5075 | 34731 | 0% | 67% | 4 |
| 9 | COMPLEX CARBOHYDRATE | 28628 | 2% | 4% | 52 |
| 10 | MED GENET CHEM | 26050 | 0% | 100% | 2 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF BIOMOLECULAR NMR | 914151 | 18% | 16% | 432 |
| 2 | JOURNAL OF MAGNETIC RESONANCE | 57986 | 9% | 2% | 200 |
| 3 | JOURNAL OF THE AMERICAN CHEMICAL SOCIETY | 30888 | 20% | 1% | 468 |
| 4 | PROGRESS IN NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY | 18031 | 1% | 5% | 26 |
| 5 | JOURNAL OF MAGNETIC RESONANCE SERIES B | 13475 | 1% | 4% | 25 |
| 6 | PROTEIN SCIENCE | 7637 | 3% | 1% | 61 |
| 7 | BIOCHEMISTRY | 5759 | 7% | 0% | 156 |
| 8 | JOURNAL OF MOLECULAR BIOLOGY | 4401 | 4% | 0% | 90 |
| 9 | NATURE STRUCTURAL BIOLOGY | 3962 | 1% | 1% | 22 |
| 10 | CONCEPTS IN MAGNETIC RESONANCE PART A | 3724 | 0% | 3% | 9 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | RESIDUAL DIPOLAR COUPLINGS | 1329603 | 8% | 52% | 198 | Search RESIDUAL+DIPOLAR+COUPLINGS | Search RESIDUAL+DIPOLAR+COUPLINGS |
| 2 | PSEUDOCONTACT SHIFTS | 435531 | 2% | 63% | 53 | Search PSEUDOCONTACT+SHIFTS | Search PSEUDOCONTACT+SHIFTS |
| 3 | N 15 RELAXATION | 344478 | 2% | 58% | 46 | Search N+15+RELAXATION | Search N+15+RELAXATION |
| 4 | CROSS CORRELATED RELAXATION | 272603 | 1% | 70% | 30 | Search CROSS+CORRELATED+RELAXATION | Search CROSS+CORRELATED+RELAXATION |
| 5 | PROTEIN DYNAMICS | 235701 | 6% | 12% | 150 | Search PROTEIN+DYNAMICS | Search PROTEIN+DYNAMICS |
| 6 | DIPOLAR COUPLING | 228906 | 3% | 26% | 68 | Search DIPOLAR+COUPLING | Search DIPOLAR+COUPLING |
| 7 | RELAXATION DISPERSION | 209908 | 1% | 46% | 35 | Search RELAXATION+DISPERSION | Search RELAXATION+DISPERSION |
| 8 | RDC | 188175 | 1% | 43% | 34 | Search RDC | Search RDC |
| 9 | ALIGNMENT TENSOR | 188075 | 1% | 76% | 19 | Search ALIGNMENT+TENSOR | Search ALIGNMENT+TENSOR |
| 10 | CHEMICAL EXCHANGE | 167837 | 3% | 22% | 59 | Search CHEMICAL+EXCHANGE | Search CHEMICAL+EXCHANGE |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | SALMON, L , BLACKLEDGE, M , (2015) INVESTIGATING PROTEIN CONFORMATIONAL ENERGY LANDSCAPES AND ATOMIC RESOLUTION DYNAMICS FROM NMR DIPOLAR COUPLINGS: A REVIEW.REPORTS ON PROGRESS IN PHYSICS. VOL. 78. ISSUE 12. P. - | 192 | 65% | 4 |
| 2 | PALMER, AG , (2004) NMR CHARACTERIZATION OF THE DYNAMICS OF BIOMACROMOLECULES.CHEMICAL REVIEWS. VOL. 104. ISSUE 8. P. 3623-3640 | 145 | 76% | 451 |
| 3 | PRESTEGARD, JH , BOUGAULT, CM , KISHORE, AI , (2004) RESIDUAL DIPOLAR COUPLINGS IN STRUCTURE DETERMINATION OF BIOMOLECULES.CHEMICAL REVIEWS. VOL. 104. ISSUE 8. P. 3519-3540 | 158 | 75% | 227 |
| 4 | BAN, D , SABO, TM , GRIESINGER, C , LEE, D , (2013) MEASURING DYNAMIC AND KINETIC INFORMATION IN THE PREVIOUSLY INACCESSIBLE SUPRA-TAU(C) WINDOW OF NANOSECONDS TO MICROSECONDS BY SOLUTION NMR SPECTROSCOPY.MOLECULES. VOL. 18. ISSUE 10. P. 11904-11937 | 140 | 76% | 7 |
| 5 | BLACKLEDGE, M , (2005) RECENT PROGRESS IN THE STUDY OF BIOMOLECULAR STRUCTURE AND DYNAMICS IN SOLUTION FROM RESIDUAL DIPOLAR COUPLINGS.PROGRESS IN NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY. VOL. 46. ISSUE 1. P. 23-61 | 135 | 78% | 180 |
| 6 | CARLON, A , RAVERA, E , ANDRALOJC, W , PARIGI, G , MURSHUDOV, GN , LUCHINAT, C , (2016) HOW TO TACKLE PROTEIN STRUCTURAL DATA FROM SOLUTION AND SOLID STATE: AN INTEGRATED APPROACH.PROGRESS IN NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY. VOL. 92-93. ISSUE . P. 54 -70 | 119 | 70% | 2 |
| 7 | SHAPIRO, YE , (2013) NMR SPECTROSCOPY ON DOMAIN DYNAMICS IN BIOMACROMOLECULES.PROGRESS IN BIOPHYSICS & MOLECULAR BIOLOGY. VOL. 112. ISSUE 3. P. 58 -117 | 214 | 45% | 8 |
| 8 | SHAHKHATUNI, AA , SHAHKHATUNI, AG , (2002) 3D STRUCTURE DETERMINATION OF WEAKLY ORIENTED BIOMOLECULES BY NMR SPECTROSCOPY.USPEKHI KHIMII. VOL. 71. ISSUE 12. P. 1132-1172 | 248 | 53% | 0 |
| 9 | PALMER, AG , (2014) CHEMICAL EXCHANGE IN BIOMACROMOLECULES: PAST, PRESENT, AND FUTURE.JOURNAL OF MAGNETIC RESONANCE. VOL. 241. ISSUE . P. 3-17 | 107 | 69% | 29 |
| 10 | TORCHIA, DA , (2015) NMR STUDIES OF DYNAMIC BIOMOLECULAR CONFORMATIONAL ENSEMBLES.PROGRESS IN NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY. VOL. 84. ISSUE . P. 14 -32 | 105 | 72% | 11 |
Classes with closest relation at Level 1 |