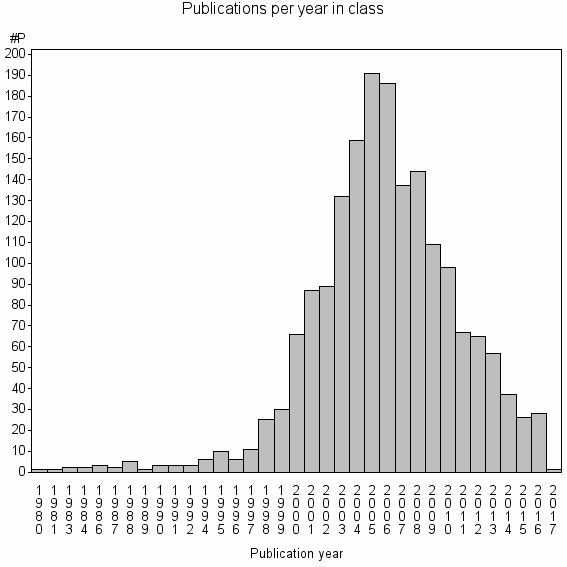
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 4394 | 1793 | 31.7 | 80% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 447 | 3 | TCTP//TRANSLATIONALLY CONTROLLED TUMOR PROTEIN//GENETIC EPIDEMIOLOGY | 26092 |
| 257 | 2 | GENETIC EPIDEMIOLOGY//HUMAN HEREDITY//GENETICS & HEREDITY | 19642 |
| 4394 | 1 | HAPLOTYPE INFERENCE//LINKAGE DISEQUILIBRIUM//HAPLOTYPE ASSEMBLY | 1793 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | HAPLOTYPE INFERENCE | authKW | 700591 | 3% | 86% | 48 |
| 2 | LINKAGE DISEQUILIBRIUM | authKW | 384383 | 13% | 10% | 230 |
| 3 | HAPLOTYPE ASSEMBLY | authKW | 204343 | 1% | 100% | 12 |
| 4 | HAPLOTYPE BLOCK | authKW | 174809 | 2% | 38% | 27 |
| 5 | HAPMAP | authKW | 155620 | 2% | 26% | 35 |
| 6 | PURE PARSIMONY | authKW | 153257 | 1% | 100% | 9 |
| 7 | HAPLOTYPE | authKW | 152651 | 12% | 4% | 209 |
| 8 | GENETIC EPIDEMIOLOGY | journal | 142521 | 8% | 6% | 141 |
| 9 | MINIMUM ERROR CORRECTION | authKW | 102171 | 0% | 100% | 6 |
| 10 | HAPLOTYPING | authKW | 99039 | 1% | 24% | 24 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Genetics & Heredity | 38123 | 60% | 0% | 1075 |
| 2 | Mathematical & Computational Biology | 28565 | 23% | 0% | 417 |
| 3 | Biotechnology & Applied Microbiology | 3053 | 18% | 0% | 331 |
| 4 | Biochemical Research Methods | 2738 | 14% | 0% | 246 |
| 5 | Statistics & Probability | 1434 | 7% | 0% | 132 |
| 6 | Computer Science, Interdisciplinary Applications | 726 | 6% | 0% | 108 |
| 7 | Biochemistry & Molecular Biology | 213 | 14% | 0% | 244 |
| 8 | Evolutionary Biology | 155 | 2% | 0% | 42 |
| 9 | Computer Science, Theory & Methods | 88 | 3% | 0% | 47 |
| 10 | Mathematics, Interdisciplinary Applications | 88 | 2% | 0% | 37 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | AKOREN VOCAT | 34057 | 0% | 100% | 2 |
| 2 | CANC ADV TECHNOL NIH | 34057 | 0% | 100% | 2 |
| 3 | UK MOL GENET | 34057 | 0% | 100% | 2 |
| 4 | ALGORITHM TEAM | 30649 | 0% | 60% | 3 |
| 5 | STAT PROGRAM STAT GENET | 30649 | 0% | 60% | 3 |
| 6 | ERABLE TEAM | 24764 | 0% | 36% | 4 |
| 7 | U535 | 23479 | 1% | 10% | 14 |
| 8 | ANHUI PROV MOST CO HIGH PERFORMANCE COMP | 22703 | 0% | 67% | 2 |
| 9 | US DISCOVERY GENET | 22703 | 0% | 67% | 2 |
| 10 | SECT GENOM VARIAT | 20168 | 0% | 15% | 8 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | GENETIC EPIDEMIOLOGY | 142521 | 8% | 6% | 141 |
| 2 | ANNALS OF HUMAN GENETICS | 42750 | 4% | 4% | 66 |
| 3 | HUMAN HEREDITY | 38166 | 4% | 3% | 68 |
| 4 | AMERICAN JOURNAL OF HUMAN GENETICS | 29440 | 7% | 1% | 118 |
| 5 | BMC GENETICS | 25364 | 3% | 3% | 50 |
| 6 | LECTURE NOTES IN BIOINFORMATICS | 17668 | 1% | 12% | 9 |
| 7 | IEEE-ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS | 14422 | 2% | 3% | 31 |
| 8 | JOURNAL OF COMPUTATIONAL BIOLOGY | 12577 | 2% | 2% | 34 |
| 9 | BIOINFORMATICS | 11444 | 5% | 1% | 85 |
| 10 | EUROPEAN JOURNAL OF HUMAN GENETICS | 8970 | 2% | 1% | 44 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | HAPLOTYPE INFERENCE | 700591 | 3% | 86% | 48 | Search HAPLOTYPE+INFERENCE | Search HAPLOTYPE+INFERENCE |
| 2 | LINKAGE DISEQUILIBRIUM | 384383 | 13% | 10% | 230 | Search LINKAGE+DISEQUILIBRIUM | Search LINKAGE+DISEQUILIBRIUM |
| 3 | HAPLOTYPE ASSEMBLY | 204343 | 1% | 100% | 12 | Search HAPLOTYPE+ASSEMBLY | Search HAPLOTYPE+ASSEMBLY |
| 4 | HAPLOTYPE BLOCK | 174809 | 2% | 38% | 27 | Search HAPLOTYPE+BLOCK | Search HAPLOTYPE+BLOCK |
| 5 | HAPMAP | 155620 | 2% | 26% | 35 | Search HAPMAP | Search HAPMAP |
| 6 | PURE PARSIMONY | 153257 | 1% | 100% | 9 | Search PURE+PARSIMONY | Search PURE+PARSIMONY |
| 7 | HAPLOTYPE | 152651 | 12% | 4% | 209 | Search HAPLOTYPE | Search HAPLOTYPE |
| 8 | MINIMUM ERROR CORRECTION | 102171 | 0% | 100% | 6 | Search MINIMUM+ERROR+CORRECTION | Search MINIMUM+ERROR+CORRECTION |
| 9 | HAPLOTYPING | 99039 | 1% | 24% | 24 | Search HAPLOTYPING | Search HAPLOTYPING |
| 10 | LINKAGE DISEQUILIBRIUM MAP | 87574 | 0% | 86% | 6 | Search LINKAGE+DISEQUILIBRIUM+MAP | Search LINKAGE+DISEQUILIBRIUM+MAP |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | LIU, NJ , ZHANG, K , ZHAO, HY , (2008) HAPLOTYPE-ASSOCIATION ANALYSIS.GENETIC DISSECTION OF COMPLEX TRAITS, 2ND EDITION. VOL. 60. ISSUE . P. 335-405 | 129 | 64% | 40 |
| 2 | GU, CC , YU, K , RAO, DC , (2008) CHARACTERIZATION OF LD STRUCTURES AND THE UTILITY OF HAPMAP IN GENETIC ASSOCIATION STUDIES.GENETIC DISSECTION OF COMPLEX TRAITS, 2ND EDITION. VOL. 60. ISSUE . P. 407-435 | 76 | 75% | 8 |
| 3 | RHEE, JK , LI, H , JOUNG, JG , HWANG, KB , ZHANG, BT , SHIN, SY , (2016) SURVEY OF COMPUTATIONAL HAPLOTYPE DETERMINATION METHODS FOR SINGLE INDIVIDUAL.GENES & GENOMICS. VOL. 38. ISSUE 1. P. 1 -12 | 54 | 83% | 0 |
| 4 | SCHAID, DJ , (2004) EVALUATING ASSOCIATIONS OF HAPLOTYPES WITH TRAITS.GENETIC EPIDEMIOLOGY. VOL. 27. ISSUE 4. P. 348-364 | 55 | 75% | 211 |
| 5 | DE BAKKER, PIW , YELENSKY, R , PE'ER, I , GABRIEL, SB , DALY, MJ , ALTSHULER, D , (2005) EFFICIENCY AND POWER IN GENETIC ASSOCIATION STUDIES.NATURE GENETICS. VOL. 37. ISSUE 11. P. 1217-1223 | 27 | 82% | 1257 |
| 6 | GU, CC , YU, K , KETKAR, S , TEMPLETON, AR , RAO, DC , (2008) ON TRANSFERABILITY OF GENOME-WIDE TAGSNPS.GENETIC EPIDEMIOLOGY. VOL. 32. ISSUE 2. P. 89-97 | 41 | 95% | 8 |
| 7 | CRAWFORD, DC , NICKERSON, DA , (2005) DEFINITION AND CLINICAL IMPORTANCE OF HAPLOTYPES.ANNUAL REVIEW OF MEDICINE. VOL. 56. ISSUE . P. 303 -+ | 58 | 57% | 179 |
| 8 | PATTARO, C , RUCZINSKI, I , FALLIN, DM , PARMIGIANI, G , (2008) HAPLOTYPE BLOCK PARTITIONING AS A TOOL FOR DIMENSIONALITY REDUCTION IN SNP ASSOCIATION STUDIES.BMC GENOMICS. VOL. 9. ISSUE . P. - | 45 | 85% | 5 |
| 9 | HOWIE, BN , CARLSON, CS , RIEDER, MJ , NICKERSON, DA , (2006) EFFICIENT SELECTION OF TAGGING SINGLE-NUCLEOTIDE POLYMORPHISMS IN MULTIPLE POPULATIONS.HUMAN GENETICS. VOL. 120. ISSUE 1. P. 58 -68 | 45 | 83% | 39 |
| 10 | HALLDORSSON, BV , ISTRAIL, S , DE LA VEGA, FM , (2004) OPTIMAL SELECTION OF SNP MARKERS FOR DISEASE ASSOCIATION STUDIES.HUMAN HEREDITY. VOL. 58. ISSUE 3-4. P. 190-202 | 41 | 89% | 43 |
Classes with closest relation at Level 1 |