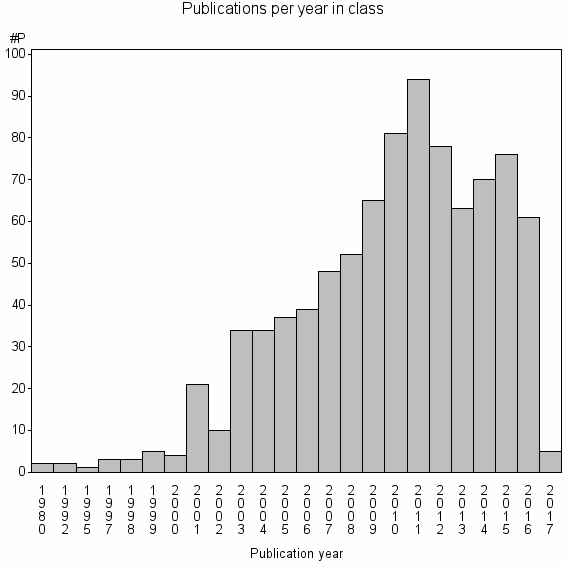
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 12684 | 888 | 36.3 | 76% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 447 | 3 | TCTP//TRANSLATIONALLY CONTROLLED TUMOR PROTEIN//GENETIC EPIDEMIOLOGY | 26092 |
| 257 | 2 | GENETIC EPIDEMIOLOGY//HUMAN HEREDITY//GENETICS & HEREDITY | 19642 |
| 12684 | 1 | GENE GENE INTERACTION//EPISTASIS//MULTIFACTOR DIMENSIONALITY REDUCTION | 888 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | GENE GENE INTERACTION | authKW | 760717 | 11% | 22% | 101 |
| 2 | EPISTASIS | authKW | 457316 | 16% | 9% | 141 |
| 3 | MULTIFACTOR DIMENSIONALITY REDUCTION | authKW | 258234 | 3% | 29% | 26 |
| 4 | SNP INTERACTION | authKW | 220062 | 1% | 80% | 8 |
| 5 | COMPUTAT GENET | address | 200342 | 3% | 22% | 26 |
| 6 | SNP SNP INTERACTIONS | authKW | 171926 | 1% | 100% | 5 |
| 7 | BIODATA MINING | journal | 157300 | 3% | 15% | 31 |
| 8 | GENOKEY S | address | 137541 | 0% | 100% | 4 |
| 9 | GRAMMATICAL EVOLUTION NEURAL NETWORKS | authKW | 137541 | 0% | 100% | 4 |
| 10 | SCIONDTU | address | 137541 | 0% | 100% | 4 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 34746 | 36% | 0% | 322 |
| 2 | Genetics & Heredity | 12039 | 48% | 0% | 428 |
| 3 | Biochemical Research Methods | 2150 | 17% | 0% | 151 |
| 4 | Biotechnology & Applied Microbiology | 1789 | 20% | 0% | 177 |
| 5 | Statistics & Probability | 678 | 7% | 0% | 64 |
| 6 | Medical Informatics | 493 | 3% | 0% | 27 |
| 7 | Computer Science, Interdisciplinary Applications | 163 | 4% | 0% | 38 |
| 8 | Computer Science, Artificial Intelligence | 69 | 3% | 0% | 26 |
| 9 | Computer Science, Theory & Methods | 42 | 3% | 0% | 23 |
| 10 | Medicine, Research & Experimental | 36 | 4% | 0% | 32 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | COMPUTAT GENET | 200342 | 3% | 22% | 26 |
| 2 | GENOKEY S | 137541 | 0% | 100% | 4 |
| 3 | SCIONDTU | 137541 | 0% | 100% | 4 |
| 4 | QUANTITAT BIOMED SCI | 109194 | 2% | 18% | 18 |
| 5 | SYST MODELING UNIT | 86662 | 1% | 23% | 11 |
| 6 | ABORAT 475 | 78143 | 1% | 45% | 5 |
| 7 | PARALLEL DISTRIBUTED ARCHITECTU GRP | 51575 | 0% | 50% | 3 |
| 8 | BIOTECHNOL CHEM ENGN | 38516 | 2% | 6% | 18 |
| 9 | ABIRISK CONSORTIUM | 34385 | 0% | 100% | 1 |
| 10 | BIOMED PATHOL MED | 34385 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIODATA MINING | 157300 | 3% | 15% | 31 |
| 2 | GENETIC EPIDEMIOLOGY | 117296 | 10% | 4% | 90 |
| 3 | HUMAN HEREDITY | 32292 | 5% | 2% | 44 |
| 4 | BMC GENETICS | 28082 | 4% | 2% | 37 |
| 5 | ANNALS OF HUMAN GENETICS | 15528 | 3% | 2% | 28 |
| 6 | BMC BIOINFORMATICS | 14313 | 6% | 1% | 56 |
| 7 | STATISTICAL APPLICATIONS IN GENETICS AND MOLECULAR BIOLOGY | 9781 | 1% | 2% | 12 |
| 8 | BIOINFORMATICS | 8672 | 6% | 0% | 52 |
| 9 | BRIEFINGS IN BIOINFORMATICS | 7101 | 1% | 2% | 12 |
| 10 | ADVANCES IN GENETICS | 4842 | 1% | 2% | 7 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | CORDELL, HJ , (2009) DETECTING GENE-GENE INTERACTIONS THAT UNDERLIE HUMAN DISEASES.NATURE REVIEWS GENETICS. VOL. 10. ISSUE 6. P. 392 -404 | 56 | 60% | 578 |
| 2 | WEI, WH , HEMANI, G , HALEY, CS , (2014) DETECTING EPISTASIS IN HUMAN COMPLEX TRAITS.NATURE REVIEWS GENETICS. VOL. 15. ISSUE 11. P. 722 -733 | 80 | 56% | 43 |
| 3 | MOORE, JH , ASSELBERGS, FW , WILLIAMS, SM , (2010) BIOINFORMATICS CHALLENGES FOR GENOME-WIDE ASSOCIATION STUDIES.BIOINFORMATICS. VOL. 26. ISSUE 4. P. 445 -455 | 49 | 54% | 189 |
| 4 | GOLA, D , JOHN, JMM , VAN STEEN, K , KONIG, IR , (2016) A ROADMAP TO MULTIFACTOR DIMENSIONALITY REDUCTION METHODS.BRIEFINGS IN BIOINFORMATICS. VOL. 17. ISSUE 2. P. 293 -308 | 48 | 65% | 1 |
| 5 | VAN STEEN, K , (2012) TRAVELLING THE WORLD OF GENE-GENE INTERACTIONS.BRIEFINGS IN BIOINFORMATICS. VOL. 13. ISSUE 1. P. 1 -19 | 63 | 51% | 26 |
| 6 | GOUDEY, B , RAWLINSON, D , WANG, Q , SHI, F , FERRA, H , CAMPBELL, RM , STERN, L , INOUYE, MT , ONG, CS , KOWALCZYK, A , (2013) GWIS - MODEL-FREE, FAST AND EXHAUSTIVE SEARCH FOR EPISTATIC INTERACTIONS IN CASE-CONTROL GWAS.BMC GENOMICS. VOL. 14. ISSUE . P. - | 35 | 88% | 16 |
| 7 | MOORE, JH , (2010) DETECTING, CHARACTERIZING, AND INTERPRETING NONLINEAR GENE-GENE INTERACTIONS USING MULTIFACTOR DIMENSIONALITY REDUCTION.COMPUTATIONAL METHODS FOR GENETICS OF COMPLEX TRAITS. VOL. 72. ISSUE . P. 101-116 | 37 | 90% | 25 |
| 8 | GUO, X , YU, N , GU, F , DING, XJ , WANG, JX , PAN, Y , (2014) GENOME-WIDE INTERACTION-BASED ASSOCIATION OF HUMAN DISEASES - A SURVEY.TSINGHUA SCIENCE AND TECHNOLOGY. VOL. 19. ISSUE 6. P. 596 -616 | 44 | 70% | 1 |
| 9 | MOORE, JH , WILLIAMS, SM , (2009) EPISTASIS AND ITS IMPLICATIONS FOR PERSONAL GENETICS.AMERICAN JOURNAL OF HUMAN GENETICS. VOL. 85. ISSUE 3. P. 309-320 | 46 | 57% | 137 |
| 10 | UPSTILL-GODDARD, R , ECCLES, D , FLIEGE, J , COLLINS, A , (2013) MACHINE LEARNING APPROACHES FOR THE DISCOVERY OF GENE-GENE INTERACTIONS IN DISEASE DATA.BRIEFINGS IN BIOINFORMATICS. VOL. 14. ISSUE 2. P. 251-260 | 33 | 87% | 16 |
Classes with closest relation at Level 1 |