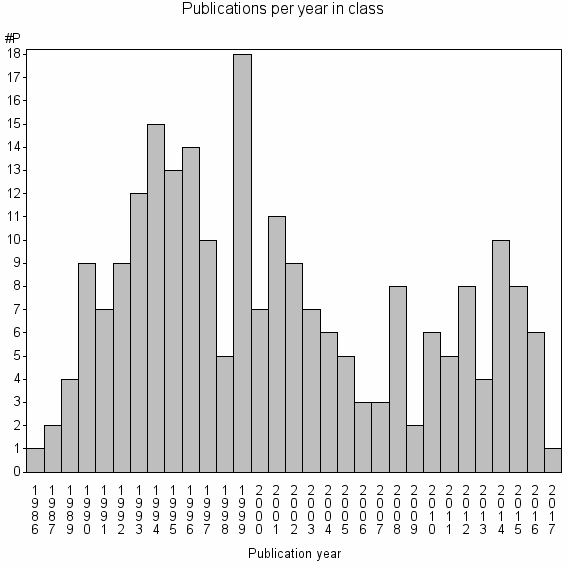
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 27495 | 228 | 40.5 | 86% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 219 | 3 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN FOLDING//PROTEIN STRUCTURE PREDICTION | 50409 |
| 2636 | 2 | COILED COIL//KRAB DOMAIN//PROTEIN DESIGN | 3515 |
| 27495 | 1 | UM SYLVESTER BRAMAN FAMILY BREAST CANC//GCN4P//DNA PROMOTER ELEMENTS TRE AND CRE | 228 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | UM SYLVESTER BRAMAN FAMILY BREAST CANC | address | 306116 | 2% | 57% | 4 |
| 2 | GCN4P | authKW | 304374 | 2% | 45% | 5 |
| 3 | DNA PROMOTER ELEMENTS TRE AND CRE | authKW | 267854 | 1% | 100% | 2 |
| 4 | TRANSCRIPTION FACTORS JUN AND FOS | authKW | 267854 | 1% | 100% | 2 |
| 5 | BASIC REGION | authKW | 238088 | 2% | 44% | 4 |
| 6 | BZIP DOMAIN | authKW | 214278 | 2% | 40% | 4 |
| 7 | CHANGE OF SPECIFICITY | authKW | 178568 | 1% | 67% | 2 |
| 8 | YEAST GCN4 | authKW | 178568 | 1% | 67% | 2 |
| 9 | GCN4 | authKW | 143990 | 4% | 11% | 10 |
| 10 | ALANINE SCANNING MUTANT | authKW | 133927 | 0% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 1374 | 63% | 0% | 144 |
| 2 | Chemistry, Multidisciplinary | 226 | 22% | 0% | 51 |
| 3 | Cell Biology | 176 | 16% | 0% | 36 |
| 4 | Biophysics | 82 | 8% | 0% | 18 |
| 5 | Chemistry, Medicinal | 76 | 6% | 0% | 14 |
| 6 | Chemistry, Organic | 58 | 9% | 0% | 20 |
| 7 | Biochemical Research Methods | 9 | 3% | 0% | 7 |
| 8 | Chemistry, Analytical | 0 | 2% | 0% | 5 |
| 9 | Mathematical & Computational Biology | 0 | 0% | 0% | 1 |
| 10 | Spectroscopy | 0 | 1% | 0% | 2 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | UM SYLVESTER BRAMAN FAMILY BREAST CANC | 306116 | 2% | 57% | 4 |
| 2 | BIOMATER BIOMED ENGN | 133927 | 0% | 100% | 1 |
| 3 | BIOMOL GRP QUIM | 133927 | 0% | 100% | 1 |
| 4 | BIOSCI NUTR BIONUT | 133927 | 0% | 100% | 1 |
| 5 | GENET MOL DESARROLLO EVOLUT PLANTAS | 133927 | 0% | 100% | 1 |
| 6 | SINGULAR INVEST QUIM BIOL MAT MOL CIQUS | 131172 | 5% | 8% | 12 |
| 7 | GRP INVEST BIOFARMA PLATAFORMA SCREENING USEF | 66963 | 0% | 50% | 1 |
| 8 | MIKROBIOL BIOL MED | 66963 | 0% | 50% | 1 |
| 9 | SCI BIOTENOL | 66963 | 0% | 50% | 1 |
| 10 | SIBERIAN V | 66963 | 0% | 50% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOLECULAR BIOLOGY | 1389 | 3% | 0% | 7 |
| 2 | CHEMBIOCHEM | 1076 | 3% | 0% | 6 |
| 3 | CHEMICAL SCIENCE | 857 | 2% | 0% | 5 |
| 4 | BIOCHEMISTRY | 795 | 8% | 0% | 18 |
| 5 | NUCLEIC ACIDS RESEARCH | 556 | 6% | 0% | 13 |
| 6 | CURRENT OPINION IN STRUCTURAL BIOLOGY | 525 | 1% | 0% | 3 |
| 7 | GENE EXPRESSION | 398 | 0% | 0% | 1 |
| 8 | JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS | 379 | 1% | 0% | 3 |
| 9 | NATURE STRUCTURAL BIOLOGY | 336 | 1% | 0% | 2 |
| 10 | CHEMICAL COMMUNICATIONS | 300 | 4% | 0% | 10 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | GCN4P | 304374 | 2% | 45% | 5 | Search GCN4P | Search GCN4P |
| 2 | DNA PROMOTER ELEMENTS TRE AND CRE | 267854 | 1% | 100% | 2 | Search DNA+PROMOTER+ELEMENTS+TRE+AND+CRE | Search DNA+PROMOTER+ELEMENTS+TRE+AND+CRE |
| 3 | TRANSCRIPTION FACTORS JUN AND FOS | 267854 | 1% | 100% | 2 | Search TRANSCRIPTION+FACTORS+JUN+AND+FOS | Search TRANSCRIPTION+FACTORS+JUN+AND+FOS |
| 4 | BASIC REGION | 238088 | 2% | 44% | 4 | Search BASIC+REGION | Search BASIC+REGION |
| 5 | BZIP DOMAIN | 214278 | 2% | 40% | 4 | Search BZIP+DOMAIN | Search BZIP+DOMAIN |
| 6 | CHANGE OF SPECIFICITY | 178568 | 1% | 67% | 2 | Search CHANGE+OF+SPECIFICITY | Search CHANGE+OF+SPECIFICITY |
| 7 | YEAST GCN4 | 178568 | 1% | 67% | 2 | Search YEAST+GCN4 | Search YEAST+GCN4 |
| 8 | GCN4 | 143990 | 4% | 11% | 10 | Search GCN4 | Search GCN4 |
| 9 | ALANINE SCANNING MUTANT | 133927 | 0% | 100% | 1 | Search ALANINE+SCANNING+MUTANT | Search ALANINE+SCANNING+MUTANT |
| 10 | ALPHA AND BETA HAIRPINS | 133927 | 0% | 100% | 1 | Search ALPHA+AND+BETA+HAIRPINS | Search ALPHA+AND+BETA+HAIRPINS |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | RODRIGUEZ, J , MOSQUERA, J , VAZQUEZ, ME , MASCARENAS, JL , (2016) NICKEL-PROMOTED RECOGNITION OF LONG DNA SITES BY DESIGNED PEPTIDE DERIVATIVES.CHEMISTRY-A EUROPEAN JOURNAL. VOL. 22. ISSUE 38. P. 13474 -13477 | 35 | 69% | 1 |
| 2 | RODRIGUEZ, J , MOSQUERA, J , GARCIA-FANDINO, R , VAZQUEZ, ME , MASCARENAS, JL , (2016) A DESIGNED DNA BINDING MOTIF THAT RECOGNIZES EXTENDED SITES AND SPANS TWO ADJACENT MAJOR GROOVES.CHEMICAL SCIENCE. VOL. 7. ISSUE 5. P. 3298 -3303 | 31 | 55% | 3 |
| 3 | HIRATA, A , UENO, M , AIZAWA, Y , OHKUBO, K , MORII, T , YOSHIKAWA, S , (2005) DUAL DNA RECOGNITION CODES OF A SHORT PEPTIDE DERIVED FROM THE BASIC LEUCINE ZIPPER PROTEIN EMBP1.BIOORGANIC & MEDICINAL CHEMISTRY. VOL. 13. ISSUE 9. P. 3107 -3116 | 37 | 67% | 1 |
| 4 | MOSQUERA, J , SANCHEZ, MI , VALERO, J , DE MENDOZA, J , VAZQUEZ, ME , MASCARENAS, JL , (2015) SEQUENCE-SELECTIVE DNA BINDING WITH CELL-PERMEABLE OLIGOGUANIDINIUM-PEPTIDE CONJUGATES.CHEMICAL COMMUNICATIONS. VOL. 51. ISSUE 23. P. 4811 -4814 | 31 | 55% | 1 |
| 5 | MOSQUERA, J , JIMENEZ-BALSA, A , DODERO, VI , VAZQUEZ, ME , MASCARENAS, JL , (2013) STIMULI-RESPONSIVE SELECTION OF TARGET DNA SEQUENCES BY SYNTHETIC BZIP PEPTIDES.NATURE COMMUNICATIONS. VOL. 4. ISSUE . P. - | 25 | 69% | 7 |
| 6 | RODRIGUEZ, J , MOSQUERA, J , COUCEIRO, JR , VAZQUEZ, ME , MASCARENAS, JL , (2015) THE AT-HOOK MOTIF AS A VERSATILE MINOR GROOVE ANCHOR FOR PROMOTING DNA BINDING OF TRANSCRIPTION FACTOR FRAGMENTS.CHEMICAL SCIENCE. VOL. 6. ISSUE 8. P. 4767 -4771 | 27 | 57% | 2 |
| 7 | SANCHEZ, MI , MOSQUERA, J , VAZQUEZ, ME , MASCARENAS, JL , (2014) REVERSIBLE SUPRAMOLECULAR ASSEMBLY AT SPECIFIC DNA SITES: NICKEL-PROMOTED BIVALENT DNA BINDING WITH DESIGNED PEPTIDE AND BIPYRIDYL-BIS(BENZAMIDINE) COMPONENTS.ANGEWANDTE CHEMIE-INTERNATIONAL EDITION. VOL. 53. ISSUE 37. P. 9917 -9921 | 25 | 56% | 5 |
| 8 | SANCHEZ, MI , VAZQUEZ, O , VAZQUEZ, ME , MASCARENAS, JL , (2013) SEQUENCE-SELECTIVE DNA RECOGNITION WITH PEPTIDE-BISBENZAMIDINE CONJUGATES.CHEMISTRY-A EUROPEAN JOURNAL. VOL. 19. ISSUE 30. P. 9923 -9929 | 23 | 53% | 7 |
| 9 | BLANCO, JB , DODERO, VL , VAZQUEZ, ME , MOSQUERA, M , CASTEDO, L , MASCARENAS, JL , (2006) SEQUENCE-SPECIFIC DNA BINDING BY NONCOVALENT PEPTIDE-TRIPYRROLE CONJUGATES.ANGEWANDTE CHEMIE-INTERNATIONAL EDITION. VOL. 45. ISSUE 48. P. 8210 -8214 | 19 | 79% | 6 |
| 10 | PELLEGRINI, M , EBRIGHT, RH , (1996) ARTIFICIAL SEQUENCE-SPECIFIC DNA BINDING PEPTIDES: BRANCHED-CHAIN BASIC REGIONS.JOURNAL OF THE AMERICAN CHEMICAL SOCIETY. VOL. 118. ISSUE 25. P. 5831-5835 | 28 | 65% | 17 |
Classes with closest relation at Level 1 |