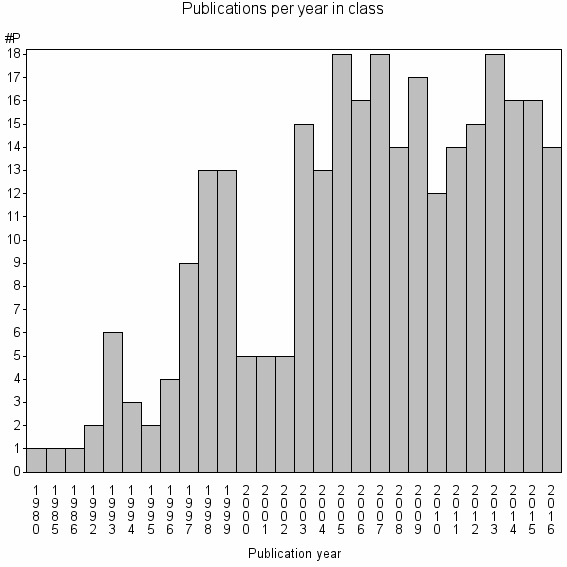
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 25281 | 286 | 30.4 | 61% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 219 | 3 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN FOLDING//PROTEIN STRUCTURE PREDICTION | 50409 |
| 260 | 2 | PROTEIN FOLDING//JOURNAL OF CHEMICAL THEORY AND COMPUTATION//JOURNAL OF COMPUTATIONAL CHEMISTRY | 19582 |
| 25281 | 1 | HP MODEL//AB OFF LATTICE MODEL//PROTEIN FOLDING PROBLEM | 286 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | HP MODEL | authKW | 2467695 | 12% | 66% | 35 |
| 2 | AB OFF LATTICE MODEL | authKW | 427066 | 1% | 100% | 4 |
| 3 | PROTEIN FOLDING PROBLEM | authKW | 427058 | 3% | 50% | 8 |
| 4 | HYDROPHOBIC POLAR MODEL | authKW | 333642 | 2% | 63% | 5 |
| 5 | FUNCTIONAL MODEL PROTEINS | authKW | 320300 | 1% | 100% | 3 |
| 6 | PULL MOVES | authKW | 320300 | 1% | 100% | 3 |
| 7 | HYDROPHOBIC POLAR HP MODEL | authKW | 213533 | 1% | 100% | 2 |
| 8 | MIYAZAWA JERNIGAN ENERGY FUNCTION | authKW | 213533 | 1% | 100% | 2 |
| 9 | OFF LATTICE AB MODEL | authKW | 213533 | 1% | 100% | 2 |
| 10 | OFF TRAP STRATEGY | authKW | 213533 | 1% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 1786 | 15% | 0% | 42 |
| 2 | Computer Science, Interdisciplinary Applications | 1404 | 19% | 0% | 55 |
| 3 | Computer Science, Artificial Intelligence | 1387 | 19% | 0% | 53 |
| 4 | Computer Science, Theory & Methods | 966 | 16% | 0% | 47 |
| 5 | Physics, Mathematical | 473 | 12% | 0% | 34 |
| 6 | Biochemical Research Methods | 384 | 13% | 0% | 37 |
| 7 | Physics, Fluids & Plasmas | 249 | 8% | 0% | 22 |
| 8 | Statistics & Probability | 204 | 7% | 0% | 20 |
| 9 | Biotechnology & Applied Microbiology | 145 | 11% | 0% | 31 |
| 10 | Physics, Atomic, Molecular & Chemical | 115 | 9% | 0% | 26 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | PHYS ENGN TANDOGAN | 213533 | 1% | 100% | 2 |
| 2 | GSIT | 160147 | 1% | 50% | 3 |
| 3 | ADM HOOFDGEBOUW | 106767 | 0% | 100% | 1 |
| 4 | GROVER ENGN | 106767 | 0% | 100% | 1 |
| 5 | GRUNDLAGEN INFORMATIONSVERARBEITUNG | 106767 | 0% | 100% | 1 |
| 6 | HUBEI INTELLIGENT INFORMAT PROC REAL TI | 106767 | 0% | 100% | 1 |
| 7 | INT INTEGRATED INTELLIGENT SYST | 106767 | 0% | 100% | 1 |
| 8 | JOHN VO NEUMANN COMP | 106767 | 0% | 100% | 1 |
| 9 | PHYS ENGN BEYTEPE | 106767 | 0% | 100% | 1 |
| 10 | SOFT TECH | 106767 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF COMPUTATIONAL BIOLOGY | 19784 | 6% | 1% | 17 |
| 2 | COMPUTATIONAL BIOLOGY AND CHEMISTRY | 5965 | 2% | 1% | 7 |
| 3 | BULLETIN OF THE BELGIAN MATHEMATICAL SOCIETY-SIMON STEVIN | 3319 | 2% | 1% | 6 |
| 4 | IEEE TRANSACTIONS ON EVOLUTIONARY COMPUTATION | 3108 | 2% | 1% | 5 |
| 5 | NATURAL COMPUTING | 2656 | 1% | 1% | 3 |
| 6 | CONSTRAINTS | 1614 | 1% | 1% | 2 |
| 7 | INTERDISCIPLINARY SCIENCES-COMPUTATIONAL LIFE SCIENCES | 1378 | 1% | 1% | 2 |
| 8 | LECTURE NOTES IN COMPUTER SCIENCE | 1273 | 12% | 0% | 33 |
| 9 | BMC BIOINFORMATICS | 1139 | 3% | 0% | 9 |
| 10 | JOURNAL OF MOLECULAR MODELING | 1060 | 2% | 0% | 6 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | HP MODEL | 2467695 | 12% | 66% | 35 | Search HP+MODEL | Search HP+MODEL |
| 2 | AB OFF LATTICE MODEL | 427066 | 1% | 100% | 4 | Search AB+OFF+LATTICE+MODEL | Search AB+OFF+LATTICE+MODEL |
| 3 | PROTEIN FOLDING PROBLEM | 427058 | 3% | 50% | 8 | Search PROTEIN+FOLDING+PROBLEM | Search PROTEIN+FOLDING+PROBLEM |
| 4 | HYDROPHOBIC POLAR MODEL | 333642 | 2% | 63% | 5 | Search HYDROPHOBIC+POLAR+MODEL | Search HYDROPHOBIC+POLAR+MODEL |
| 5 | FUNCTIONAL MODEL PROTEINS | 320300 | 1% | 100% | 3 | Search FUNCTIONAL+MODEL+PROTEINS | Search FUNCTIONAL+MODEL+PROTEINS |
| 6 | PULL MOVES | 320300 | 1% | 100% | 3 | Search PULL+MOVES | Search PULL+MOVES |
| 7 | HYDROPHOBIC POLAR HP MODEL | 213533 | 1% | 100% | 2 | Search HYDROPHOBIC+POLAR+HP+MODEL | Search HYDROPHOBIC+POLAR+HP+MODEL |
| 8 | MIYAZAWA JERNIGAN ENERGY FUNCTION | 213533 | 1% | 100% | 2 | Search MIYAZAWA+JERNIGAN+ENERGY+FUNCTION | Search MIYAZAWA+JERNIGAN+ENERGY+FUNCTION |
| 9 | OFF LATTICE AB MODEL | 213533 | 1% | 100% | 2 | Search OFF+LATTICE+AB+MODEL | Search OFF+LATTICE+AB+MODEL |
| 10 | OFF TRAP STRATEGY | 213533 | 1% | 100% | 2 | Search OFF+TRAP+STRATEGY | Search OFF+TRAP+STRATEGY |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | GARZA-FABRE, M , RODRIGUEZ-TELLO, E , TOSCANO-PULIDO, G , (2013) COMPARATIVE ANALYSIS OF DIFFERENT EVALUATION FUNCTIONS FOR PROTEIN STRUCTURE PREDICTION UNDER THE HP MODEL.JOURNAL OF COMPUTER SCIENCE AND TECHNOLOGY. VOL. 28. ISSUE 5. P. 868 -889 | 20 | 80% | 2 |
| 2 | WUST, T , LANDAU, DP , (2012) OPTIMIZED WANG-LANDAU SAMPLING OF LATTICE POLYMERS: GROUND STATE SEARCH AND FOLDING THERMODYNAMICS OF HP MODEL PROTEINS.JOURNAL OF CHEMICAL PHYSICS. VOL. 137. ISSUE 6. P. - | 30 | 49% | 16 |
| 3 | TSAY, JJ , SU, SC , (2013) AN EFFECTIVE EVOLUTIONARY ALGORITHM FOR PROTEIN FOLDING ON 3D FCC HP MODEL BY LATTICE ROTATION AND GENERALIZED MOVE SETS.PROTEOME SCIENCE. VOL. 11. ISSUE . P. - | 20 | 77% | 0 |
| 4 | LIU, JF , LI, G , YU, J , (2011) PROTEIN-FOLDING SIMULATIONS OF THE HYDROPHOBIC-HYDROPHILIC MODEL BY COMBINING PULL MOVES WITH ENERGY LANDSCAPE PAVING.PHYSICAL REVIEW E. VOL. 84. ISSUE 3. P. - | 20 | 74% | 5 |
| 5 | LJU, JF , SUN, YY , LI, G , SONG, BB , HUANG, WB , (2013) HEURISTIC-BASED TABU SEARCH ALGORITHM FOR FOLDING TWO-DIMENSIONAL AB OFF-LATTICE MODEL PROTEINS.COMPUTATIONAL BIOLOGY AND CHEMISTRY. VOL. 47. ISSUE . P. 142-148 | 15 | 94% | 0 |
| 6 | SHI, GJ , VOGEL, T , WUST, T , LI, YW , LANDAU, DP , (2014) EFFECT OF SINGLE-SITE MUTATIONS ON HYDROPHOBIC-POLAR LATTICE PROTEINS.PHYSICAL REVIEW E. VOL. 90. ISSUE 3. P. - | 26 | 47% | 7 |
| 7 | GARZA-FABRE, M , RODRIGUEZ-TELLO, E , TOSCANO-PULIDO, G , (2015) CONSTRAINT-HANDLING THROUGH MULTI-OBJECTIVE OPTIMIZATION: THE HYDROPHOBIC-POLAR MODEL FOR PROTEIN STRUCTURE PREDICTION.COMPUTERS & OPERATIONS RESEARCH. VOL. 53. ISSUE . P. 128 -153 | 21 | 51% | 4 |
| 8 | LU, ZP , HUANG, WQ , SHI, H , (2007) QUASI-PHYSICAL ALGORITHM FOR PROTEIN FOLDING IN AN OFF-LATTICE MODEL.COMMUNICATIONS IN THEORETICAL PHYSICS. VOL. 47. ISSUE 1. P. 181-185 | 16 | 84% | 1 |
| 9 | HUANG, WQ , LU, ZP , SHI, H , (2005) GROWTH ALGORITHM FOR FINDING LOW ENERGY CONFIGURATIONS OF SIMPLE LATTICE PROTEINS.PHYSICAL REVIEW E. VOL. 72. ISSUE 1. P. - | 18 | 75% | 9 |
| 10 | SANTOS, J , VILLOT, P , DIEGUEZ, M , (2014) EMERGENT PROTEIN FOLDING MODELED WITH EVOLVED NEURAL CELLULAR AUTOMATA USING THE 3D HP MODEL.JOURNAL OF COMPUTATIONAL BIOLOGY. VOL. 21. ISSUE 11. P. 823 -845 | 16 | 67% | 2 |
Classes with closest relation at Level 1 |