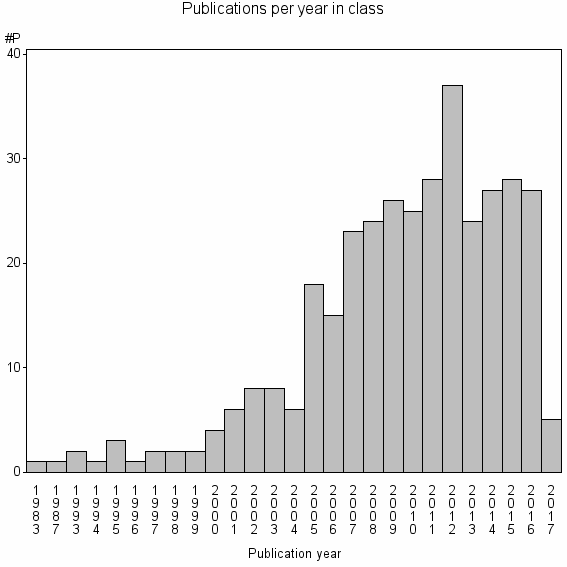
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 23215 | 354 | 46.5 | 88% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 157 | 3 | JOURNAL OF THE AMERICAN SOCIETY FOR MASS SPECTROMETRY//BIOCHEMICAL RESEARCH METHODS//INTERNATIONAL JOURNAL OF MASS SPECTROMETRY | 61558 |
| 370 | 2 | PROTEOMICS//JOURNAL OF PROTEOME RESEARCH//BIOCHEMICAL RESEARCH METHODS | 17272 |
| 23215 | 1 | DEGRADOMICS//LIFE SCI 4 401//N TERMINOMICS | 354 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | DEGRADOMICS | authKW | 745848 | 6% | 41% | 21 |
| 2 | LIFE SCI 4 401 | address | 652315 | 3% | 69% | 11 |
| 3 | N TERMINOMICS | authKW | 368024 | 2% | 53% | 8 |
| 4 | TERMINOMICS | authKW | 345030 | 1% | 100% | 4 |
| 5 | DIAGONAL CHROMATOGRAPHY | authKW | 345026 | 2% | 67% | 6 |
| 6 | ORAL BIOL MED SCI | address | 260608 | 10% | 8% | 37 |
| 7 | COFRADIC | authKW | 240913 | 3% | 31% | 9 |
| 8 | N TERMINAL SEQUENCE ANALYSIS | authKW | 230017 | 1% | 67% | 4 |
| 9 | POSITIONAL PROTEOMICS | authKW | 197156 | 1% | 57% | 4 |
| 10 | BR SIGNATURE | authKW | 172515 | 1% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemical Research Methods | 6231 | 44% | 0% | 156 |
| 2 | Biochemistry & Molecular Biology | 1160 | 48% | 0% | 170 |
| 3 | Chemistry, Analytical | 337 | 14% | 0% | 51 |
| 4 | Mathematical & Computational Biology | 272 | 5% | 0% | 19 |
| 5 | Biotechnology & Applied Microbiology | 99 | 8% | 0% | 30 |
| 6 | Chemistry, Medicinal | 34 | 4% | 0% | 13 |
| 7 | Biophysics | 31 | 5% | 0% | 16 |
| 8 | Spectroscopy | 28 | 3% | 0% | 11 |
| 9 | Cell Biology | 16 | 6% | 0% | 20 |
| 10 | Genetics & Heredity | 5 | 3% | 0% | 11 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | LIFE SCI 4 401 | 652315 | 3% | 69% | 11 |
| 2 | ORAL BIOL MED SCI | 260608 | 10% | 8% | 37 |
| 3 | AG SYSTEMAT PROTEOMFOR BIOANALYT | 115009 | 1% | 67% | 2 |
| 4 | MED PROT | 96050 | 7% | 4% | 25 |
| 5 | AG SYSTEMAT PROTEOME BIOANALYT | 86257 | 0% | 100% | 1 |
| 6 | BIOCATALYST CHARACTERIZAT DESIGN | 86257 | 0% | 100% | 1 |
| 7 | BIOCHEM ORAL BIOL MED SCI | 86257 | 0% | 100% | 1 |
| 8 | COMPETENCE SYST PHYSIOL METABOL DIS | 86257 | 0% | 100% | 1 |
| 9 | COMPUTAT BIOL MOL BIOPHYS PROGRAM | 86257 | 0% | 100% | 1 |
| 10 | CORP BIOL | 86257 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOLECULAR & CELLULAR PROTEOMICS | 17482 | 7% | 1% | 25 |
| 2 | PROTEOMICS | 15432 | 9% | 1% | 32 |
| 3 | BIOLOGICAL CHEMISTRY | 6054 | 4% | 0% | 15 |
| 4 | JOURNAL OF PROTEOME RESEARCH | 4070 | 5% | 0% | 17 |
| 5 | NATURE METHODS | 2487 | 2% | 0% | 7 |
| 6 | CURRENT OPINION IN CHEMICAL BIOLOGY | 2238 | 2% | 0% | 7 |
| 7 | DATABASE-THE JOURNAL OF BIOLOGICAL DATABASES AND CURATION | 1187 | 1% | 0% | 3 |
| 8 | EXPERT REVIEW OF PROTEOMICS | 1055 | 1% | 0% | 3 |
| 9 | NATURE PROTOCOLS | 945 | 1% | 0% | 5 |
| 10 | NATURE BIOTECHNOLOGY | 815 | 1% | 0% | 5 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DEGRADOMICS | 745848 | 6% | 41% | 21 | Search DEGRADOMICS | Search DEGRADOMICS |
| 2 | N TERMINOMICS | 368024 | 2% | 53% | 8 | Search N+TERMINOMICS | Search N+TERMINOMICS |
| 3 | TERMINOMICS | 345030 | 1% | 100% | 4 | Search TERMINOMICS | Search TERMINOMICS |
| 4 | DIAGONAL CHROMATOGRAPHY | 345026 | 2% | 67% | 6 | Search DIAGONAL+CHROMATOGRAPHY | Search DIAGONAL+CHROMATOGRAPHY |
| 5 | COFRADIC | 240913 | 3% | 31% | 9 | Search COFRADIC | Search COFRADIC |
| 6 | N TERMINAL SEQUENCE ANALYSIS | 230017 | 1% | 67% | 4 | Search N+TERMINAL+SEQUENCE+ANALYSIS | Search N+TERMINAL+SEQUENCE+ANALYSIS |
| 7 | POSITIONAL PROTEOMICS | 197156 | 1% | 57% | 4 | Search POSITIONAL+PROTEOMICS | Search POSITIONAL+PROTEOMICS |
| 8 | BR SIGNATURE | 172515 | 1% | 100% | 2 | Search BR+SIGNATURE | Search BR+SIGNATURE |
| 9 | C TERMINOMICS | 172515 | 1% | 100% | 2 | Search C+TERMINOMICS | Search C+TERMINOMICS |
| 10 | N TERMINAL PEPTIDES | 172511 | 1% | 50% | 4 | Search N+TERMINAL+PEPTIDES | Search N+TERMINAL+PEPTIDES |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | VIZOVISEK, M , VIDMAR, R , FONOVIC, M , TURK, B , (2016) CURRENT TRENDS AND CHALLENGES IN PROTEOMIC IDENTIFICATION OF PROTEASE SUBSTRATES.BIOCHIMIE. VOL. 122. ISSUE . P. 77 -87 | 57 | 57% | 3 |
| 2 | ROGERS, LD , OVERALL, CM , (2013) PROTEOLYTIC POST-TRANSLATIONAL MODIFICATION OF PROTEINS: PROTEOMIC TOOLS AND METHODOLOGY.MOLECULAR & CELLULAR PROTEOMICS. VOL. 12. ISSUE 12. P. 3532 -3542 | 53 | 62% | 24 |
| 3 | SHAHINIAN, H , THOLEN, S , SCHILLING, O , (2013) PROTEOMIC IDENTIFICATION OF PROTEASE CLEAVAGE SITES: CELL-BIOLOGICAL AND BIOMEDICAL APPLICATIONS.EXPERT REVIEW OF PROTEOMICS. VOL. 10. ISSUE 5. P. 421 -433 | 58 | 48% | 5 |
| 4 | CHEN, LF , SHAN, YC , WENG, YJ , SUI, ZG , ZHANG, XD , LIANG, Z , ZHANG, LH , ZHANG, YK , (2016) HYDROPHOBIC TAGGING-ASSISTED N-TERMINI ENRICHMENT FOR IN-DEPTH N-TERMINOME ANALYSIS.ANALYTICAL CHEMISTRY. VOL. 88. ISSUE 17. P. 8390 -8395 | 31 | 78% | 0 |
| 5 | HUESGEN, PF , OVERALL, CM , (2012) N- AND C-TERMINAL DEGRADOMICS: NEW APPROACHES TO REVEAL BIOLOGICAL ROLES FOR PLANT PROTEASES FROM SUBSTRATE IDENTIFICATION.PHYSIOLOGIA PLANTARUM. VOL. 145. ISSUE 1. P. 5-17 | 28 | 76% | 14 |
| 6 | MARINO, G , ECICHARD, U , OVERALL, CM , (2015) PROTEIN TERMINI AND THEIR MODIFICATIONS REVEALED BY POSITIONAL PROTEOMICS.ACS CHEMICAL BIOLOGY. VOL. 10. ISSUE 8. P. 1754 -1764 | 39 | 43% | 7 |
| 7 | LANGE, PF , HUESGEN, PF , OVERALL, CM , (2012) TOPFIND 2.0-LINKING PROTEIN TERMINI WITH PROTEOLYTIC PROCESSING AND MODIFICATIONS ALTERING PROTEIN FUNCTION.NUCLEIC ACIDS RESEARCH. VOL. 40. ISSUE D1. P. D351 -D361 | 25 | 76% | 13 |
| 8 | CHEN, LF , SHAN, YC , WENG, YJ , YUAN, HM , ZHANG, S , FAN, RL , SUI, ZG , ZHANG, XD , ZHANG, LH , ZHANG, YK , (2016) DEPLETION OF INTERNAL PEPTIDES BY SITE-SELECTIVE BLOCKING, PHOSPHATE LABELING, AND TIO2 ADSORPTION FOR IN-DEPTH ANALYSIS OF C-TERMINOME.ANALYTICAL AND BIOANALYTICAL CHEMISTRY. VOL. 408. ISSUE 14. P. 3867 -3874 | 24 | 73% | 0 |
| 9 | KLINGLER, D , HARDT, M , (2012) PROFILING PROTEASE ACTIVITIES BY DYNAMIC PROTEOMICS WORKFLOWS.PROTEOMICS. VOL. 12. ISSUE 4-5. P. 587 -596 | 37 | 46% | 11 |
| 10 | VAN DEN BERG, BHJ , THOLEY, A , (2012) MASS SPECTROMETRY-BASED PROTEOMICS STRATEGIES FOR PROTEASE CLEAVAGE SITE IDENTIFICATION.PROTEOMICS. VOL. 12. ISSUE 4-5. P. 516 -529 | 39 | 44% | 11 |
Classes with closest relation at Level 1 |