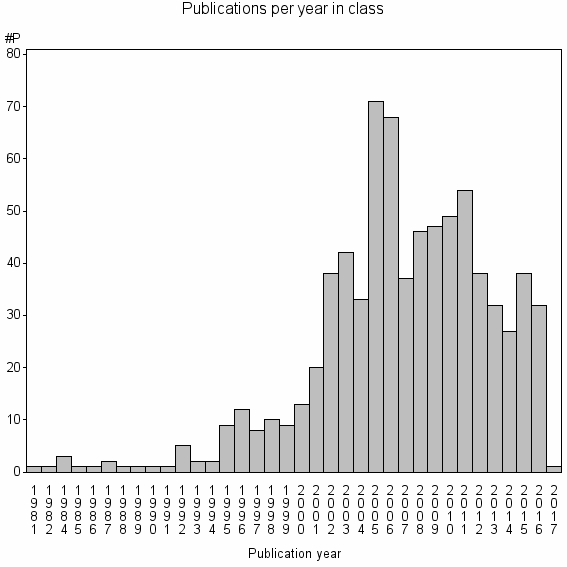
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 14677 | 756 | 23.8 | 59% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 126 | 2 | MOLECULAR BIOLOGY AND EVOLUTION//BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY | 23963 |
| 14677 | 1 | GENOME REARRANGEMENT//SORTING BY REVERSALS//GENOME REARRANGEMENTS | 756 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | GENOME REARRANGEMENT | authKW | 1526184 | 10% | 48% | 78 |
| 2 | SORTING BY REVERSALS | authKW | 981455 | 4% | 90% | 27 |
| 3 | GENOME REARRANGEMENTS | authKW | 749627 | 6% | 40% | 46 |
| 4 | BREAKPOINT GRAPH | authKW | 565450 | 2% | 100% | 14 |
| 5 | COMMON INTERVALS | authKW | 527752 | 2% | 93% | 14 |
| 6 | DCJ | authKW | 444282 | 1% | 100% | 11 |
| 7 | SORTING BY TRANSPOSITIONS | authKW | 363504 | 1% | 100% | 9 |
| 8 | GENE TEAMS | authKW | 323114 | 1% | 100% | 8 |
| 9 | CONSECUTIVE ONES PROPERTY | authKW | 263862 | 2% | 47% | 14 |
| 10 | BREAKPOINT DISTANCE | authKW | 258488 | 1% | 80% | 8 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 33833 | 39% | 0% | 293 |
| 2 | Biochemical Research Methods | 9939 | 38% | 0% | 289 |
| 3 | Biotechnology & Applied Microbiology | 4925 | 35% | 0% | 261 |
| 4 | Computer Science, Theory & Methods | 4267 | 21% | 0% | 159 |
| 5 | Statistics & Probability | 4164 | 18% | 0% | 139 |
| 6 | Computer Science, Interdisciplinary Applications | 4070 | 20% | 0% | 152 |
| 7 | Computer Science, Information Systems | 1115 | 10% | 0% | 79 |
| 8 | Mathematics, Applied | 894 | 15% | 0% | 111 |
| 9 | Genetics & Heredity | 667 | 13% | 0% | 102 |
| 10 | Mathematics, Interdisciplinary Applications | 309 | 5% | 0% | 38 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | DATA MIN SEARCH GRP | 168287 | 1% | 83% | 5 |
| 2 | EPFL IC LCBB | 121168 | 0% | 100% | 3 |
| 3 | AG GENOMINFORMAT | 100958 | 1% | 25% | 10 |
| 4 | LACIM | 94646 | 3% | 12% | 19 |
| 5 | EQUIPE EBM | 80779 | 0% | 100% | 2 |
| 6 | IIS LCBB | 80779 | 0% | 100% | 2 |
| 7 | PUBL HLTH INFORMAT FELLOW PROGRAM | 80779 | 0% | 100% | 2 |
| 8 | TEAM BAMBOO | 80779 | 0% | 100% | 2 |
| 9 | IGM INFO | 59389 | 1% | 29% | 5 |
| 10 | TECH FAK | 59177 | 3% | 6% | 26 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF COMPUTATIONAL BIOLOGY | 248949 | 13% | 6% | 98 |
| 2 | IEEE-ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS | 43717 | 5% | 3% | 35 |
| 3 | BMC BIOINFORMATICS | 20645 | 8% | 1% | 62 |
| 4 | ALGORITHMS FOR MOLECULAR BIOLOGY | 14253 | 1% | 4% | 10 |
| 5 | BIOINFORMATICS | 10205 | 7% | 0% | 52 |
| 6 | THEORETICAL COMPUTER SCIENCE | 7921 | 6% | 0% | 46 |
| 7 | LECTURE NOTES IN COMPUTER SCIENCE | 6347 | 16% | 0% | 119 |
| 8 | SIAM JOURNAL ON DISCRETE MATHEMATICS | 3847 | 2% | 1% | 14 |
| 9 | GENE-COMBIS | 3364 | 0% | 8% | 1 |
| 10 | DISCRETE APPLIED MATHEMATICS | 3238 | 3% | 0% | 23 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | GENOME REARRANGEMENT | 1526184 | 10% | 48% | 78 | Search GENOME+REARRANGEMENT | Search GENOME+REARRANGEMENT |
| 2 | SORTING BY REVERSALS | 981455 | 4% | 90% | 27 | Search SORTING+BY+REVERSALS | Search SORTING+BY+REVERSALS |
| 3 | GENOME REARRANGEMENTS | 749627 | 6% | 40% | 46 | Search GENOME+REARRANGEMENTS | Search GENOME+REARRANGEMENTS |
| 4 | BREAKPOINT GRAPH | 565450 | 2% | 100% | 14 | Search BREAKPOINT+GRAPH | Search BREAKPOINT+GRAPH |
| 5 | COMMON INTERVALS | 527752 | 2% | 93% | 14 | Search COMMON+INTERVALS | Search COMMON+INTERVALS |
| 6 | DCJ | 444282 | 1% | 100% | 11 | Search DCJ | Search DCJ |
| 7 | SORTING BY TRANSPOSITIONS | 363504 | 1% | 100% | 9 | Search SORTING+BY+TRANSPOSITIONS | Search SORTING+BY+TRANSPOSITIONS |
| 8 | GENE TEAMS | 323114 | 1% | 100% | 8 | Search GENE+TEAMS | Search GENE+TEAMS |
| 9 | CONSECUTIVE ONES PROPERTY | 263862 | 2% | 47% | 14 | Search CONSECUTIVE+ONES+PROPERTY | Search CONSECUTIVE+ONES+PROPERTY |
| 10 | BREAKPOINT DISTANCE | 258488 | 1% | 80% | 8 | Search BREAKPOINT+DISTANCE | Search BREAKPOINT+DISTANCE |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | LIN, YC , TANG, CY , (2006) EXPOSING PHYLOGENETIC RELATIONSHIPS BY GENOME REARRANGEMENT.ADVANCES IN COMPUTERS , VOL 68. VOL. 68. ISSUE . P. 1 -57 | 68 | 67% | 1 |
| 2 | RAJARAMAN, A , MA, J , (2016) RECONSTRUCTING ANCESTRAL GENE ORDERS WITH DUPLICATIONS GUIDED BY SYNTENY LEVEL GENOME RECONSTRUCTION.BMC BIOINFORMATICS. VOL. 17. ISSUE . P. - | 27 | 87% | 0 |
| 3 | HUANG, YL , HUANG, CC , TANG, CY , LU, CL , (2010) SORT(2): A TOOL FOR SORTING GENOMES AND RECONSTRUCTING PHYLOGENETIC TREES BY REVERSALS, GENERALIZED TRANSPOSITIONS AND TRANSLOCATIONS.NUCLEIC ACIDS RESEARCH. VOL. 38. ISSUE . P. W221 -W227 | 25 | 96% | 5 |
| 4 | SANKOFF, D , ZHENG, CF , MUNOZ, A , YANG, ZY , ADAM, Z , WARREN, R , CHOI, V , ZHU, Q , (2010) ISSUES IN THE RECONSTRUCTION OF GENE ORDER EVOLUTION.JOURNAL OF COMPUTER SCIENCE AND TECHNOLOGY. VOL. 25. ISSUE 1. P. 10 -25 | 27 | 87% | 1 |
| 5 | FEIJAO, P , MEIDANIS, J , (2011) SCJ: A BREAKPOINT-LIKE DISTANCE THAT SIMPLIFIES SEVERAL REARRANGEMENT PROBLEMS.IEEE-ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS. VOL. 8. ISSUE 5. P. 1318 -1329 | 24 | 92% | 9 |
| 6 | LI, ZM , WANG, LH , ZHANG, KH , (2006) ALGORITHMIC APPROACHES FOR GENOME REARRANGEMENT: A REVIEW.IEEE TRANSACTIONS ON SYSTEMS MAN AND CYBERNETICS PART C-APPLICATIONS AND REVIEWS. VOL. 36. ISSUE 5. P. 636 -648 | 31 | 82% | 9 |
| 7 | FEIJAO, P , ARAUJO, E , (2016) FAST ANCESTRAL GENE ORDER RECONSTRUCTION OF GENOMES WITH UNEQUAL GENE CONTENT.BMC BIOINFORMATICS. VOL. 17. ISSUE . P. - | 20 | 100% | 0 |
| 8 | CHEN, KT , HUANG, KH , LU, CL , (2011) SORTING PERMUTATIONS BY CUT-CIRCULARIZE-LINEARIZE-AND-PASTE OPERATIONS.BMC GENOMICS. VOL. 12. ISSUE . P. - | 21 | 100% | 1 |
| 9 | HUANG, YL , LU, CL , (2010) SORTING BY REVERSALS, GENERALIZED TRANSPOSITIONS, AND TRANSLOCATIONS USING PERMUTATION GROUPS.JOURNAL OF COMPUTATIONAL BIOLOGY. VOL. 17. ISSUE 5. P. 685 -705 | 21 | 100% | 5 |
| 10 | AVDEYEV, P , JIANG, S , AGANEZOV, S , HU, F , ALEKSEYEV, MA , (2016) RECONSTRUCTION OF ANCESTRAL GENOMES IN PRESENCE OF GENE GAIN AND LOSS.JOURNAL OF COMPUTATIONAL BIOLOGY. VOL. 23. ISSUE 3. P. 150 -164 | 15 | 100% | 3 |
Classes with closest relation at Level 1 |