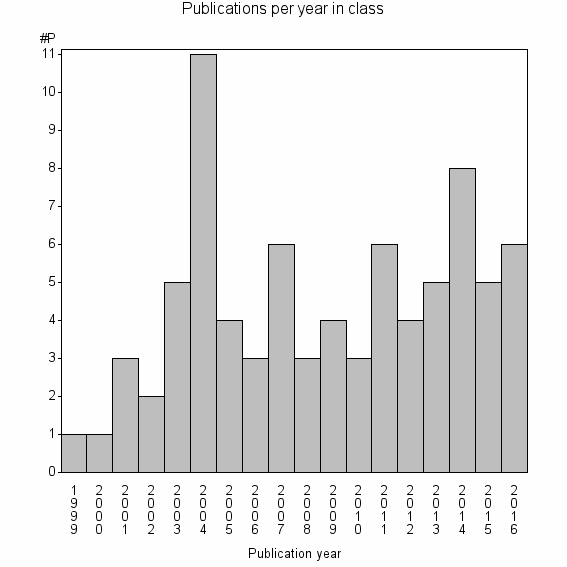
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 36483 | 80 | 18.2 | 85% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 126 | 2 | MOLECULAR BIOLOGY AND EVOLUTION//BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY | 23963 |
| 36483 | 1 | GENOMIC DATA VISUALIZATION//ALTERNATE ASSEMBLIES OF COW GENOMES//BAC FISH MAPPING | 80 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | GENOMIC DATA VISUALIZATION | authKW | 858814 | 4% | 75% | 3 |
| 2 | ALTERNATE ASSEMBLIES OF COW GENOMES | authKW | 381696 | 1% | 100% | 1 |
| 3 | BAC FISH MAPPING | authKW | 381696 | 1% | 100% | 1 |
| 4 | BHF GLASGOW CARIOVASC | address | 381696 | 1% | 100% | 1 |
| 5 | COW GENOME | authKW | 381696 | 1% | 100% | 1 |
| 6 | DATA DEFINITION FORMAT | authKW | 381696 | 1% | 100% | 1 |
| 7 | DATABASE SEARCH VIEWING | authKW | 381696 | 1% | 100% | 1 |
| 8 | EMERGENT CLUSTERS | authKW | 381696 | 1% | 100% | 1 |
| 9 | ENSEMBL PROJECT | address | 381696 | 1% | 100% | 1 |
| 10 | GFF | authKW | 381696 | 1% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 8633 | 60% | 0% | 48 |
| 2 | Biotechnology & Applied Microbiology | 2685 | 76% | 0% | 61 |
| 3 | Biochemical Research Methods | 2645 | 60% | 0% | 48 |
| 4 | Genetics & Heredity | 169 | 20% | 0% | 16 |
| 5 | Microbiology | 17 | 8% | 0% | 6 |
| 6 | Horticulture | 15 | 3% | 0% | 2 |
| 7 | Biochemistry & Molecular Biology | 10 | 14% | 0% | 11 |
| 8 | Allergy | 4 | 1% | 0% | 1 |
| 9 | Biology | 3 | 3% | 0% | 2 |
| 10 | Engineering, Biomedical | 1 | 1% | 0% | 1 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BHF GLASGOW CARIOVASC | 381696 | 1% | 100% | 1 |
| 2 | ENSEMBL PROJECT | 381696 | 1% | 100% | 1 |
| 3 | HLTH RELATED INFORMAT BIOIMAGING | 381696 | 1% | 100% | 1 |
| 4 | PLANT PATHOGEN PROGRAMME | 190847 | 1% | 50% | 1 |
| 5 | STRUCT BIOL BIOCOMP | 138795 | 3% | 18% | 2 |
| 6 | GENE EXP S EUKARYOTES | 127231 | 1% | 33% | 1 |
| 7 | GENOM BIOCOMP | 127231 | 1% | 33% | 1 |
| 8 | CORE IL PROT | 63614 | 1% | 17% | 1 |
| 9 | HEIDELBERG PERSONALIZED ONCOL DKFZ HIPO | 54526 | 1% | 14% | 1 |
| 10 | MED BIOL COMP | 52644 | 3% | 7% | 2 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOINFORMATICS | 39167 | 41% | 0% | 33 |
| 2 | BMC BIOINFORMATICS | 7328 | 15% | 0% | 12 |
| 3 | KOREAN JOURNAL OF HORTICULTURAL SCIENCE & TECHNOLOGY | 1484 | 3% | 0% | 2 |
| 4 | BMC GENOMICS | 1432 | 8% | 0% | 6 |
| 5 | GENOME RESEARCH | 1107 | 3% | 0% | 2 |
| 6 | METHODS IN MICROBIOLOGY | 871 | 1% | 0% | 1 |
| 7 | STANDARDS IN GENOMIC SCIENCES | 568 | 1% | 0% | 1 |
| 8 | BRIEFINGS IN BIOINFORMATICS | 547 | 1% | 0% | 1 |
| 9 | OMICS-A JOURNAL OF INTEGRATIVE BIOLOGY | 543 | 1% | 0% | 1 |
| 10 | ISRAEL JOURNAL OF PLANT SCIENCES | 500 | 1% | 0% | 1 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | GENOMIC DATA VISUALIZATION | 858814 | 4% | 75% | 3 | Search GENOMIC+DATA+VISUALIZATION | Search GENOMIC+DATA+VISUALIZATION |
| 2 | ALTERNATE ASSEMBLIES OF COW GENOMES | 381696 | 1% | 100% | 1 | Search ALTERNATE+ASSEMBLIES+OF+COW+GENOMES | Search ALTERNATE+ASSEMBLIES+OF+COW+GENOMES |
| 3 | BAC FISH MAPPING | 381696 | 1% | 100% | 1 | Search BAC+FISH+MAPPING | Search BAC+FISH+MAPPING |
| 4 | COW GENOME | 381696 | 1% | 100% | 1 | Search COW+GENOME | Search COW+GENOME |
| 5 | DATA DEFINITION FORMAT | 381696 | 1% | 100% | 1 | Search DATA+DEFINITION+FORMAT | Search DATA+DEFINITION+FORMAT |
| 6 | DATABASE SEARCH VIEWING | 381696 | 1% | 100% | 1 | Search DATABASE+SEARCH+VIEWING | Search DATABASE+SEARCH+VIEWING |
| 7 | EMERGENT CLUSTERS | 381696 | 1% | 100% | 1 | Search EMERGENT+CLUSTERS | Search EMERGENT+CLUSTERS |
| 8 | GFF | 381696 | 1% | 100% | 1 | Search GFF | Search GFF |
| 9 | INFERENCE OF BIOLOGICAL PATHWAYS | 381696 | 1% | 100% | 1 | Search INFERENCE+OF+BIOLOGICAL+PATHWAYS | Search INFERENCE+OF+BIOLOGICAL+PATHWAYS |
| 10 | NONGLOBULAR PROTEINS | 381696 | 1% | 100% | 1 | Search NONGLOBULAR+PROTEINS | Search NONGLOBULAR+PROTEINS |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | KRZYWINSKI, M , SCHEIN, J , BIROL, I , CONNORS, J , GASCOYNE, R , HORSMAN, D , JONES, SJ , MARRA, MA , (2009) CIRCOS: AN INFORMATION AESTHETIC FOR COMPARATIVE GENOMICS.GENOME RESEARCH. VOL. 19. ISSUE 9. P. 1639 -1645 | 11 | 32% | 1777 |
| 2 | CRABTREE, J , AGRAWAL, S , MAHURKAR, A , MYERS, GS , RASKO, DA , WHITE, O , (2014) CIRCLEATOR: FLEXIBLE CIRCULAR VISUALIZATION OF GENOME-ASSOCIATED DATA WITH BIOPERL AND SVG.BIOINFORMATICS. VOL. 30. ISSUE 21. P. 3125 -3127 | 9 | 53% | 4 |
| 3 | ZHANG, HE , MELTZER, PS , DAVIS, SR , (2016) CAOMICSV: AN R PACKAGE FOR VISUALIZING MULTIDIMENSIONAL CANCER GENOMIC DATA.BMC BIOINFORMATICS. VOL. 17. ISSUE . P. - | 6 | 60% | 0 |
| 4 | CUI, Y , CHEN, XW , LUO, HX , FAN, Z , LUO, JJ , HE, SM , YUE, HY , ZHANG, P , CHEN, RS , (2016) BIOCIRCOS.JS: AN INTERACTIVE CIRCOS JAVASCRIPT LIBRARY FOR BIOLOGICAL DATA VISUALIZATION ON WEB APPLICATIONS.BIOINFORMATICS. VOL. 32. ISSUE 11. P. 1740 -1742 | 5 | 56% | 0 |
| 5 | NAQUIN, D , D'AUBENTON-CARAFA, Y , THERMES, C , SILVAIN, M , (2014) CIRCUS: A PACKAGE FOR CIRCOS DISPLAY OF STRUCTURAL GENOME VARIATIONS FROM PAIRED-END AND MATE-PAIR SEQUENCING DATA.BMC BIOINFORMATICS. VOL. 15. ISSUE . P. - | 4 | 67% | 7 |
| 6 | CHEONG, WH , TAN, YC , YAP, SJ , NG, KP , (2015) CLICO FS: AN INTERACTIVE WEB-BASED SERVICE OF CIRCOS.BIOINFORMATICS. VOL. 31. ISSUE 22. P. 3685 -3687 | 4 | 67% | 1 |
| 7 | GU, ZG , GU, L , EILS, R , SCHLESNER, M , BRORS, B , (2014) CIRCLIZE IMPLEMENTS AND ENHANCES CIRCULAR VISUALIZATION IN R.BIOINFORMATICS. VOL. 30. ISSUE 19. P. 2811 -2812 | 3 | 75% | 21 |
| 8 | AN, JY , LAI, J , SAJJANHAR, A , BATRA, J , WANG, CW , NELSON, CC , (2015) J-CIRCOS: AN INTERACTIVE CIRCOS PLOTTER.BIOINFORMATICS. VOL. 31. ISSUE 9. P. 1463 -1465 | 4 | 57% | 4 |
| 9 | ALIKHAN, NF , PETTY, NK , BEN ZAKOUR, NL , BEATSON, SA , (2011) BLAST RING IMAGE GENERATOR (BRIG): SIMPLE PROKARYOTE GENOME COMPARISONS.BMC GENOMICS. VOL. 12. ISSUE . P. - | 8 | 18% | 320 |
| 10 | TANYALCIN, I , AL ASSAF, C , GHELDOF, A , STOUFFS, K , LISSENS, W , JANSEN, AC , (2016) I-PV: A CIRCOS MODULE FOR INTERACTIVE PROTEIN SEQUENCE VISUALIZATION.BIOINFORMATICS. VOL. 32. ISSUE 3. P. 447 -449 | 4 | 57% | 0 |
Classes with closest relation at Level 1 |