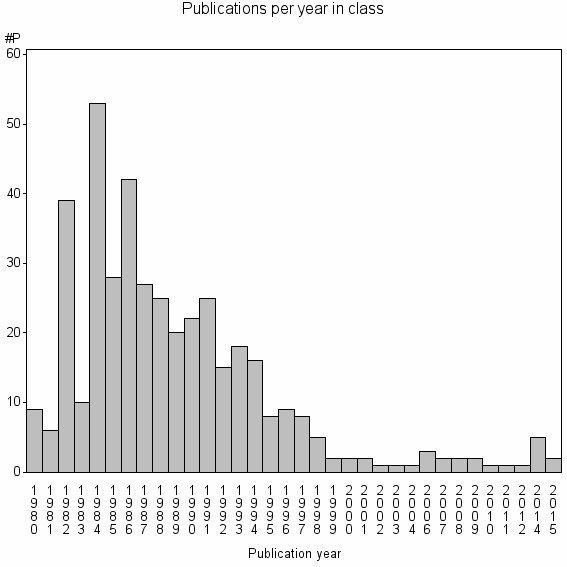
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 21589 | 413 | 17.0 | 53% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 3958 | 2 | COMPUTER APPLICATIONS IN THE BIOSCIENCES//ADAPTIVE WEIGHTED LEAST SQUARES//AMBIGRAM | 751 |
| 21589 | 1 | COMPUTER APPLICATIONS IN THE BIOSCIENCES//ADAPTIVE WEIGHTED LEAST SQUARES//AMBIGRAM | 413 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | COMPUTER APPLICATIONS IN THE BIOSCIENCES | journal | 228374 | 13% | 6% | 53 |
| 2 | ADAPTIVE WEIGHTED LEAST SQUARES | authKW | 147869 | 0% | 100% | 2 |
| 3 | AMBIGRAM | authKW | 147869 | 0% | 100% | 2 |
| 4 | ALPHABETIC SEQUENCES | authKW | 73935 | 0% | 100% | 1 |
| 5 | BINARY DNA ENCODING | authKW | 73935 | 0% | 100% | 1 |
| 6 | BINARY NUCLEOTIDE ENCODING | authKW | 73935 | 0% | 100% | 1 |
| 7 | BIOLOGICAL DATA BANKS | authKW | 73935 | 0% | 100% | 1 |
| 8 | BUCANDIN | authKW | 73935 | 0% | 100% | 1 |
| 9 | DIRECC PROYECTOS INVEST | address | 73935 | 0% | 100% | 1 |
| 10 | DNA 64 | authKW | 73935 | 0% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Computer Science, Interdisciplinary Applications | 1701 | 18% | 0% | 73 |
| 2 | Biochemistry & Molecular Biology | 1657 | 53% | 0% | 217 |
| 3 | Biochemical Research Methods | 591 | 13% | 0% | 55 |
| 4 | Medical Informatics | 246 | 3% | 0% | 13 |
| 5 | Engineering, Biomedical | 96 | 4% | 0% | 18 |
| 6 | Education, Scientific Disciplines | 85 | 2% | 0% | 10 |
| 7 | Mathematical & Computational Biology | 68 | 3% | 0% | 11 |
| 8 | Computer Science, Software Engineering | 65 | 3% | 0% | 14 |
| 9 | Computer Science, Theory & Methods | 56 | 4% | 0% | 16 |
| 10 | Computer Science, Hardware & Architecture | 46 | 2% | 0% | 9 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | DIRECC PROYECTOS INVEST | 73935 | 0% | 100% | 1 |
| 2 | HOSP INFORMAT NUMER IMAGING GRP | 73935 | 0% | 100% | 1 |
| 3 | IISE | 73935 | 0% | 100% | 1 |
| 4 | NPAL | 36966 | 0% | 50% | 1 |
| 5 | IST GENET MOL EVOLUTIONARY GENET | 24644 | 0% | 33% | 1 |
| 6 | TRANSFERT ONCOL PREDICT | 24644 | 0% | 33% | 1 |
| 7 | 110 AVE PINS OUEST | 18482 | 0% | 25% | 1 |
| 8 | VISUAL STUDIES | 14783 | 0% | 10% | 2 |
| 9 | SECT MED TECHNOL | 10560 | 0% | 14% | 1 |
| 10 | TIERARZTL | 6720 | 0% | 9% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | COMPUTER APPLICATIONS IN THE BIOSCIENCES | 228374 | 13% | 6% | 53 |
| 2 | NUCLEIC ACIDS RESEARCH | 24908 | 28% | 0% | 115 |
| 3 | COMPUTER PROGRAMS IN BIOMEDICINE | 11871 | 1% | 3% | 6 |
| 4 | BIOCHEMICAL EDUCATION | 11679 | 2% | 2% | 9 |
| 5 | BIOTECHNIQUES | 8874 | 7% | 0% | 27 |
| 6 | AUSTRALASIAN BIOTECHNOLOGY | 5404 | 1% | 2% | 3 |
| 7 | JOURNAL OF BIOINFORMATICS AND COMPUTATIONAL BIOLOGY | 1812 | 1% | 1% | 3 |
| 8 | JOURNAL OF AUDIOVISUAL MEDIA IN MEDICINE | 1678 | 0% | 2% | 1 |
| 9 | COMPUTER METHODS AND PROGRAMS IN BIOMEDICINE | 1392 | 2% | 0% | 8 |
| 10 | METHODS OF BIOCHEMICAL ANALYSIS | 1367 | 0% | 2% | 1 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ADAPTIVE WEIGHTED LEAST SQUARES | 147869 | 0% | 100% | 2 | Search ADAPTIVE+WEIGHTED+LEAST+SQUARES | Search ADAPTIVE+WEIGHTED+LEAST+SQUARES |
| 2 | AMBIGRAM | 147869 | 0% | 100% | 2 | Search AMBIGRAM | Search AMBIGRAM |
| 3 | ALPHABETIC SEQUENCES | 73935 | 0% | 100% | 1 | Search ALPHABETIC+SEQUENCES | Search ALPHABETIC+SEQUENCES |
| 4 | BINARY DNA ENCODING | 73935 | 0% | 100% | 1 | Search BINARY+DNA+ENCODING | Search BINARY+DNA+ENCODING |
| 5 | BINARY NUCLEOTIDE ENCODING | 73935 | 0% | 100% | 1 | Search BINARY+NUCLEOTIDE+ENCODING | Search BINARY+NUCLEOTIDE+ENCODING |
| 6 | BIOLOGICAL DATA BANKS | 73935 | 0% | 100% | 1 | Search BIOLOGICAL+DATA+BANKS | Search BIOLOGICAL+DATA+BANKS |
| 7 | BUCANDIN | 73935 | 0% | 100% | 1 | Search BUCANDIN | Search BUCANDIN |
| 8 | DNA 64 | 73935 | 0% | 100% | 1 | Search DNA+64 | Search DNA+64 |
| 9 | FIS PREDICTION | 73935 | 0% | 100% | 1 | Search FIS+PREDICTION | Search FIS+PREDICTION |
| 10 | GENERIC SEQUENCE | 73935 | 0% | 100% | 1 | Search GENERIC+SEQUENCE | Search GENERIC+SEQUENCE |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | BISHOP, MJ , (1984) SOFTWARE FOR MOLECULAR-BIOLOGY .3. GENERAL-ANALYSIS PROGRAMS CLASSIFIED ACCORDING TO THE COMPUTER ON WHICH THEY RUN.BIOESSAYS. VOL. 1. ISSUE 3. P. 126-129 | 42 | 76% | 0 |
| 2 | KORN, LJ , QUEEN, C , (1984) ANALYSIS OF BIOLOGICAL SEQUENCES ON SMALL COMPUTERS.DNA-A JOURNAL OF MOLECULAR & CELLULAR BIOLOGY. VOL. 3. ISSUE 6. P. 421-436 | 49 | 56% | 30 |
| 3 | CAMPIONEPICCARDO, J , RUBEN, M , (1986) AN INTEGRATED SOFTWARE SYSTEM FOR MICROCOMPUTER MANAGEMENT OF RECOMBINANT-DNA DATA.NUCLEIC ACIDS RESEARCH. VOL. 14. ISSUE 1. P. 571-574 | 16 | 94% | 2 |
| 4 | AKBARI, A , ALBREGTSEN, F , LINGJAERDE, OC , (2002) ADAPTIVE WEIGHTED LEAST SQUARES METHOD FOR THE ESTIMATION OF DNA FRAGMENT LENGTHS FROM AGAROSE GELS.ELECTROPHORESIS. VOL. 23. ISSUE 2. P. 176-181 | 6 | 100% | 4 |
| 5 | CRABBE, MJC , (1985) COMPUTERS IN BIOCHEMICAL-ANALYSIS.METHODS OF BIOCHEMICAL ANALYSIS. VOL. 31. ISSUE . P. 417-474 | 37 | 28% | 3 |
| 6 | CAMPIONEPICCARDO, J , (1988) ALGORITHMS FOR DETERMINING THE FATE OF SITES AND DOMAIN BOUNDARIES IN COMPUTER-SIMULATIONS OF RECOMBINANT DNA PROCEDURES.COMPUTER APPLICATIONS IN THE BIOSCIENCES. VOL. 4. ISSUE 2. P. 259-264 | 8 | 100% | 0 |
| 7 | JENSON, HB , (1987) 2 COMPUTER-PROGRAMS FOR RAPID ENTRY OF DNA-SEQUENCE DATA.COMPUTER APPLICATIONS IN THE BIOSCIENCES. VOL. 3. ISSUE 4. P. 283-286 | 9 | 90% | 2 |
| 8 | JUNGCK, JR , FRIEDMAN, RM , (1984) MATHEMATICAL TOOLS FOR MOLECULAR-GENETICS DATA - AN ANNOTATED-BIBLIOGRAPHY.BULLETIN OF MATHEMATICAL BIOLOGY. VOL. 46. ISSUE 4. P. 699-744 | 33 | 29% | 10 |
| 9 | CARRINGTON, JM , THOMPSON, MP , (1992) CAL IN THE BIOCHEMICAL LABORATORY.BIOCHEMICAL EDUCATION. VOL. 20. ISSUE 1. P. 32-35 | 6 | 100% | 3 |
| 10 | BISHOP, MJ , (1984) SOFTWARE FOR MOLECULAR-BIOLOGY .2. RESTRICTION MAPPING AND DNA SEQUENCING PROGRAMS.BIOESSAYS. VOL. 1. ISSUE 2. P. 75-77 | 14 | 67% | 0 |
Classes with closest relation at Level 1 |