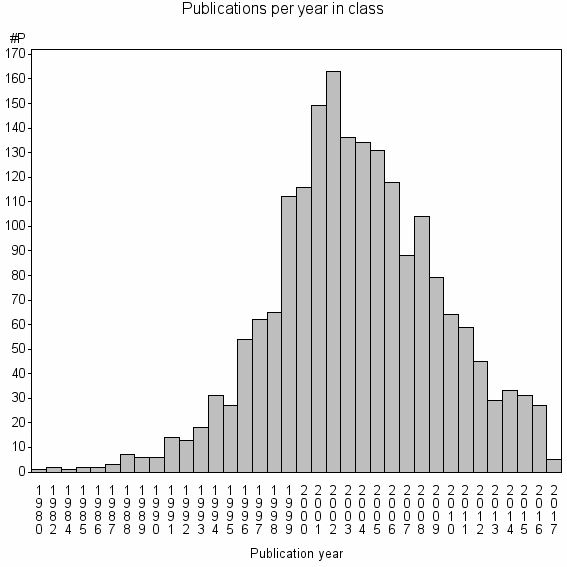
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 3649 | 1937 | 31.0 | 81% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 1714 | 2 | MOLECULAR BEACON//MUTATION DETECTION//SEQUENCING BY HYBRIDIZATION | 6662 |
| 3649 | 1 | SEQUENCING BY HYBRIDIZATION//DNA CHIP//DNA GRAPHS | 1937 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | SEQUENCING BY HYBRIDIZATION | authKW | 508128 | 2% | 92% | 35 |
| 2 | DNA CHIP | authKW | 180744 | 3% | 18% | 64 |
| 3 | DNA GRAPHS | authKW | 85815 | 0% | 78% | 7 |
| 4 | OLIGONUCLEOTIDE ARRAYS | authKW | 82346 | 1% | 29% | 18 |
| 5 | DNA SEQUENCING WITH ERRORS | authKW | 78812 | 0% | 100% | 5 |
| 6 | GAPPED PROBES | authKW | 78812 | 0% | 100% | 5 |
| 7 | CHEM SENSORS GRP | address | 77813 | 1% | 22% | 22 |
| 8 | DNA SEQUENCING BY HYBRIDIZATION | authKW | 63050 | 0% | 100% | 4 |
| 9 | NONSPECIFIC HYBRIDIZATIONS | authKW | 63050 | 0% | 100% | 4 |
| 10 | HYBRIDIZATION EFFICIENCY | authKW | 53085 | 0% | 42% | 8 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemical Research Methods | 5901 | 19% | 0% | 367 |
| 2 | Biotechnology & Applied Microbiology | 2857 | 17% | 0% | 335 |
| 3 | Chemistry, Analytical | 2408 | 16% | 0% | 314 |
| 4 | Biochemistry & Molecular Biology | 2371 | 31% | 0% | 610 |
| 5 | Genetics & Heredity | 764 | 10% | 0% | 185 |
| 6 | Mathematical & Computational Biology | 694 | 4% | 0% | 73 |
| 7 | Biophysics | 519 | 7% | 0% | 135 |
| 8 | Nanoscience & Nanotechnology | 452 | 6% | 0% | 118 |
| 9 | Chemistry, Multidisciplinary | 275 | 11% | 0% | 207 |
| 10 | Electrochemistry | 267 | 4% | 0% | 73 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CHEM SENSORS GRP | 77813 | 1% | 22% | 22 |
| 2 | DOO SAN ENGN | 50438 | 0% | 80% | 4 |
| 3 | JOINT HUMAN GENOME PROGRAM | 42031 | 0% | 67% | 4 |
| 4 | PL MICRONANOTECHNOL | 36025 | 0% | 57% | 4 |
| 5 | MOL BIOANAL BIOELECT | 35464 | 0% | 75% | 3 |
| 6 | CHEM FUNCT BIOSYST | 31525 | 0% | 100% | 2 |
| 7 | GENE EXP S DETECT | 31525 | 0% | 100% | 2 |
| 8 | PHD MICROBIOL PROGRAM | 31525 | 0% | 100% | 2 |
| 9 | TMAS2 | 31525 | 0% | 100% | 2 |
| 10 | TRANSPORT MODELING BIOANALYT SEPARAT SCI GRP | 31525 | 0% | 100% | 2 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOTECHNIQUES | 12295 | 4% | 1% | 69 |
| 2 | NUCLEIC ACIDS RESEARCH | 9553 | 8% | 0% | 156 |
| 3 | ANALYTICAL BIOCHEMISTRY | 4967 | 4% | 0% | 77 |
| 4 | INTERNATIONAL JOURNAL OF GENOME RESEARCH | 4200 | 0% | 13% | 2 |
| 5 | JOURNAL OF COMPUTATIONAL BIOLOGY | 3617 | 1% | 1% | 19 |
| 6 | MINERVA BIOTECNOLOGICA | 3564 | 1% | 2% | 11 |
| 7 | BIOSENSORS & BIOELECTRONICS | 3114 | 2% | 0% | 44 |
| 8 | GENETIC ANALYSIS-BIOMOLECULAR ENGINEERING | 2858 | 0% | 3% | 7 |
| 9 | BIOCHIP JOURNAL | 2226 | 0% | 2% | 8 |
| 10 | BIOTECHFORUM ( BTF ) | 1942 | 1% | 1% | 18 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | SEQUENCING BY HYBRIDIZATION | 508128 | 2% | 92% | 35 | Search SEQUENCING+BY+HYBRIDIZATION | Search SEQUENCING+BY+HYBRIDIZATION |
| 2 | DNA CHIP | 180744 | 3% | 18% | 64 | Search DNA+CHIP | Search DNA+CHIP |
| 3 | DNA GRAPHS | 85815 | 0% | 78% | 7 | Search DNA+GRAPHS | Search DNA+GRAPHS |
| 4 | OLIGONUCLEOTIDE ARRAYS | 82346 | 1% | 29% | 18 | Search OLIGONUCLEOTIDE+ARRAYS | Search OLIGONUCLEOTIDE+ARRAYS |
| 5 | DNA SEQUENCING WITH ERRORS | 78812 | 0% | 100% | 5 | Search DNA+SEQUENCING+WITH+ERRORS | Search DNA+SEQUENCING+WITH+ERRORS |
| 6 | GAPPED PROBES | 78812 | 0% | 100% | 5 | Search GAPPED+PROBES | Search GAPPED+PROBES |
| 7 | DNA SEQUENCING BY HYBRIDIZATION | 63050 | 0% | 100% | 4 | Search DNA+SEQUENCING+BY+HYBRIDIZATION | Search DNA+SEQUENCING+BY+HYBRIDIZATION |
| 8 | NONSPECIFIC HYBRIDIZATIONS | 63050 | 0% | 100% | 4 | Search NONSPECIFIC+HYBRIDIZATIONS | Search NONSPECIFIC+HYBRIDIZATIONS |
| 9 | HYBRIDIZATION EFFICIENCY | 53085 | 0% | 42% | 8 | Search HYBRIDIZATION+EFFICIENCY | Search HYBRIDIZATION+EFFICIENCY |
| 10 | HETEROBIFUNCTIONAL REAGENT | 50438 | 0% | 80% | 4 | Search HETEROBIFUNCTIONAL+REAGENT | Search HETEROBIFUNCTIONAL+REAGENT |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | HELLER, MJ , (2002) DNA MICROARRAY TECHNOLOGY: DEVICES, SYSTEMS, AND APPLICATIONS.ANNUAL REVIEW OF BIOMEDICAL ENGINEERING. VOL. 4. ISSUE . P. 129 -153 | 78 | 44% | 578 |
| 2 | HARRISON, A , BINDER, H , BUHOT, A , BURDEN, CJ , CARLON, E , GIBAS, C , GAMBLE, LJ , HALPERIN, A , HOOYBERGHS, J , KREIL, DP , ET AL (2013) PHYSICO-CHEMICAL FOUNDATIONS UNDERPINNING MICROARRAY AND NEXT-GENERATION SEQUENCING EXPERIMENTS.NUCLEIC ACIDS RESEARCH. VOL. 41. ISSUE 5. P. 2779 -2796 | 52 | 66% | 12 |
| 3 | BEAUCAGE, SL , (2001) STRATEGIES IN THE PREPARATION OF DNA OLIGONUCLEOTIDE ARRAYS FOR DIAGNOSTIC APPLICATIONS.CURRENT MEDICINAL CHEMISTRY. VOL. 8. ISSUE 10. P. 1213-1244 | 55 | 64% | 95 |
| 4 | GAO, XL , GULARI, E , ZHOU, XC , (2004) IN SITU SYNTHESIS OF OLIGONUCLEOTIDE MICROARRAYS.BIOPOLYMERS. VOL. 73. ISSUE 5. P. 579 -596 | 45 | 66% | 139 |
| 5 | KINOSHITA, K , FUJIMOTO, K , YAKABE, T , SAITO, S , HAMAGUCHI, Y , KIKUCHI, T , NONAKA, K , MURATA, S , MASUDA, D , TAKADA, W , ET AL (2007) MULTIPLE PRIMER EXTENSION BY DNA POLYMERASE ON A NOVEL PLASTIC DNA ARRAY COATED WITH A BIOCOMPATIBLE POLYMER.NUCLEIC ACIDS RESEARCH. VOL. 35. ISSUE 1. P. - | 36 | 84% | 22 |
| 6 | HACIA, JG , (1999) RESEQUENCING AND MUTATIONAL ANALYSIS USING OLIGONUCLEOTIDE MICROARRAYS.NATURE GENETICS. VOL. 21. ISSUE . P. 42 -47 | 37 | 73% | 357 |
| 7 | HACIA, JG , COLLINS, FS , (1999) MUTATIONAL ANALYSIS USING OLIGONUCLEOTIDE MICROARRAYS.JOURNAL OF MEDICAL GENETICS. VOL. 36. ISSUE 10. P. 730-736 | 42 | 81% | 105 |
| 8 | PIRRUNG, MC , (2002) HOW TO MAKE A DNA CHIP.ANGEWANDTE CHEMIE-INTERNATIONAL EDITION. VOL. 41. ISSUE 8. P. 1277-+ | 47 | 67% | 74 |
| 9 | POZHITKOV, AE , BOUBE, I , BROUWER, MH , NOBLE, PA , (2010) BEYOND AFFYMETRIX ARRAYS: EXPANDING THE SET OF KNOWN HYBRIDIZATION ISOTHERMS AND OBSERVING PRE-WASH SIGNAL INTENSITIES.NUCLEIC ACIDS RESEARCH. VOL. 38. ISSUE 5. P. - | 32 | 84% | 13 |
| 10 | KETOMAKI, K , HAKALA, H , KURONEN, O , LONNBERG, H , (2003) HYBRIDIZATION PROPERTIES OF SUPPORT-BOUND OLIGONUCLEOTIDES: THE EFFECT OF THE SITE OF IMMOBILIZATION ON THE STABILITY AND SELECTIVITY OF DUPLEX FORMATION.BIOCONJUGATE CHEMISTRY. VOL. 14. ISSUE 4. P. 811-816 | 43 | 78% | 6 |
Classes with closest relation at Level 1 |