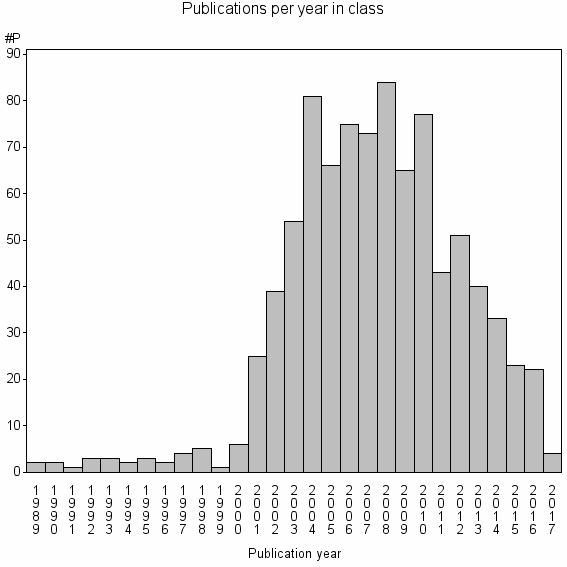
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 12669 | 889 | 34.1 | 82% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 1714 | 2 | MOLECULAR BEACON//MUTATION DETECTION//SEQUENCING BY HYBRIDIZATION | 6662 |
| 12669 | 1 | PRIMER DESIGN//SIMULTANEOUS MULTIPLE DETECTION//MICROBIAL DIAGNOSTICS | 889 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | PRIMER DESIGN | authKW | 115589 | 2% | 16% | 21 |
| 2 | SIMULTANEOUS MULTIPLE DETECTION | authKW | 103040 | 0% | 100% | 3 |
| 3 | MICROBIAL DIAGNOSTICS | authKW | 78503 | 0% | 57% | 4 |
| 4 | DIGITAL CONTENT DESIGN MANAGEMENT | address | 78055 | 1% | 45% | 5 |
| 5 | DIAGNOSTIC MICROARRAY | authKW | 71549 | 1% | 42% | 5 |
| 6 | 40 MER OLIGONUCLEOTIDE PROBES | authKW | 68693 | 0% | 100% | 2 |
| 7 | COMMUNITY GENOME ARRAY | authKW | 68693 | 0% | 100% | 2 |
| 8 | DEGENERATE PRIMER DESIGN | authKW | 68693 | 0% | 100% | 2 |
| 9 | DNA MICROARRAY CHIP | authKW | 68693 | 0% | 100% | 2 |
| 10 | MUTAGENIC PRIMER DESIGN | authKW | 68693 | 0% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biotechnology & Applied Microbiology | 8806 | 42% | 0% | 375 |
| 2 | Mathematical & Computational Biology | 6688 | 16% | 0% | 143 |
| 3 | Biochemical Research Methods | 6669 | 29% | 0% | 259 |
| 4 | Microbiology | 4124 | 29% | 0% | 258 |
| 5 | Virology | 543 | 6% | 0% | 57 |
| 6 | Biochemistry & Molecular Biology | 329 | 20% | 0% | 176 |
| 7 | Genetics & Heredity | 80 | 5% | 0% | 48 |
| 8 | Infectious Diseases | 36 | 3% | 0% | 24 |
| 9 | Chemistry, Analytical | 19 | 4% | 0% | 32 |
| 10 | Food Science & Technology | 11 | 2% | 0% | 22 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | DIGITAL CONTENT DESIGN MANAGEMENT | 78055 | 1% | 45% | 5 |
| 2 | PROGRAMA POS BIOTECNOL GENOM | 68693 | 0% | 100% | 2 |
| 3 | ISIMA LIMOS | 61821 | 0% | 60% | 3 |
| 4 | EPIDEMIOL PATHOPHYSIOL ONCOGEN VIRUS UNIT | 45794 | 0% | 67% | 2 |
| 5 | ADV GENMOIC TECHNOL | 34347 | 0% | 100% | 1 |
| 6 | AG TAUTZ | 34347 | 0% | 100% | 1 |
| 7 | AGR HIERARCHICAL UTILIZAT | 34347 | 0% | 100% | 1 |
| 8 | ALGORTIHMS STAT SYST BIOL GRP | 34347 | 0% | 100% | 1 |
| 9 | ALL RUSSIAN STATE GENET BREEDING FARM | 34347 | 0% | 100% | 1 |
| 10 | ASTOBIOL | 34347 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOINFORMATICS | 21109 | 9% | 1% | 81 |
| 2 | BMC BIOINFORMATICS | 9630 | 5% | 1% | 46 |
| 3 | JOURNAL OF MICROBIOLOGICAL METHODS | 6364 | 3% | 1% | 30 |
| 4 | MOLECULAR AND CELLULAR PROBES | 5159 | 2% | 1% | 16 |
| 5 | JOURNAL OF VIROLOGICAL METHODS | 4689 | 3% | 0% | 31 |
| 6 | NUCLEIC ACIDS RESEARCH | 4561 | 8% | 0% | 73 |
| 7 | APPLIED AND ENVIRONMENTAL MICROBIOLOGY | 4220 | 7% | 0% | 61 |
| 8 | RUSSIAN JOURNAL OF BIOORGANIC CHEMISTRY | 1525 | 1% | 1% | 8 |
| 9 | COMPARATIVE AND FUNCTIONAL GENOMICS | 1423 | 0% | 1% | 4 |
| 10 | JOURNAL OF CLINICAL MICROBIOLOGY | 1367 | 4% | 0% | 33 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | LESKI, TA , MALANOSKI, AP , STENGER, DA , LIN, BC , (2010) TARGET AMPLIFICATION FOR BROAD SPECTRUM MICROBIAL DIAGNOSTICS AND DETECTION.FUTURE MICROBIOLOGY. VOL. 5. ISSUE 2. P. 191-203 | 57 | 49% | 4 |
| 2 | DUGAT-BONY, E , PEYRETAILLADE, E , PARISOT, N , BIDERRE-PETIT, C , JAZIRI, F , HILL, D , RIMOUR, S , PEYRET, P , (2012) DETECTING UNKNOWN SEQUENCES WITH DNA MICROARRAYS: EXPLORATIVE PROBE DESIGN STRATEGIES.ENVIRONMENTAL MICROBIOLOGY. VOL. 14. ISSUE 2. P. 356 -371 | 58 | 44% | 12 |
| 3 | MILLER, SM , TOURLOUSSE, DM , STEDTFELD, RD , BAUSHKE, SW , HERZOG, AB , WICK, LM , ROUILLARD, JM , GULARI, E , TIEDJE, JM , HASHSHAM, SA , (2008) IN SITU-SYNTHESIZED VIRULENCE AND MARKER GENE BIOCHIP FOR DETECTION OF BACTERIAL PATHOGENS IN WATER.APPLIED AND ENVIRONMENTAL MICROBIOLOGY. VOL. 74. ISSUE 7. P. 2200-2209 | 42 | 63% | 19 |
| 4 | LOY, A , BODROSSY, L , (2006) HIGHLY PARALLEL MICROBIAL DIAGNOSTICS USING OLIGONUCLEOTIDE MICROARRAYS.CLINICA CHIMICA ACTA. VOL. 363. ISSUE 1-2. P. 106-119 | 48 | 41% | 78 |
| 5 | LESKI, TA , LIN, BC , MALANOSKI, AP , STENGER, DA , (2012) APPLICATION OF RESEQUENCING MICROARRAYS IN MICROBIAL DETECTION AND CHARACTERIZATION.FUTURE MICROBIOLOGY. VOL. 7. ISSUE 5. P. 625-637 | 38 | 49% | 4 |
| 6 | GENTRY, TJ , WICKHAM, GS , SCHADT, CW , HE, Z , ZHOU, J , (2006) MICROARRAY APPLICATIONS IN MICROBIAL ECOLOGY RESEARCH.MICROBIAL ECOLOGY. VOL. 52. ISSUE 2. P. 159-175 | 45 | 40% | 89 |
| 7 | SESSITSCH, A , HACKL, E , WENZL, P , KILIAN, A , KOSTIC, T , STRALIS-PAVESE, N , SANDJONG, BT , BODROSSY, L , (2006) DIAGNOSTIC MICROBIAL MICROARRAYS IN SOIL ECOLOGY.NEW PHYTOLOGIST. VOL. 171. ISSUE 4. P. 719-736 | 46 | 38% | 45 |
| 8 | HE, ZL , DENG, Y , ZHOU, JZ , (2012) DEVELOPMENT OF FUNCTIONAL GENE MICROARRAYS FOR MICROBIAL COMMUNITY ANALYSIS.CURRENT OPINION IN BIOTECHNOLOGY. VOL. 23. ISSUE 1. P. 49-55 | 27 | 56% | 15 |
| 9 | BODROSSY, L , SESSITSCH, A , (2004) OLIGONUCLEOTIDE MICROARRAYS IN MICROBIAL DIAGNOSTICS.CURRENT OPINION IN MICROBIOLOGY. VOL. 7. ISSUE 3. P. 245-254 | 30 | 51% | 151 |
| 10 | KIM, DH , LEE, BK , KIM, YD , RHEE, SK , KIM, YC , (2010) DETECTION OF REPRESENTATIVE ENTEROPATHOGENIC BACTERIA, VIBRIO SPP., PATHOGENIC ESCHERICHIA COLI, SALMONELLA SPP., SHIGELLA SPP., AND YERSINIA ENTEROCOLITICA, USING A VIRULENCE FACTOR GENE-BASED OLIGONUCLEOTIDE MICROARRAY.JOURNAL OF MICROBIOLOGY. VOL. 48. ISSUE 5. P. 682-688 | 23 | 66% | 10 |
Classes with closest relation at Level 1 |