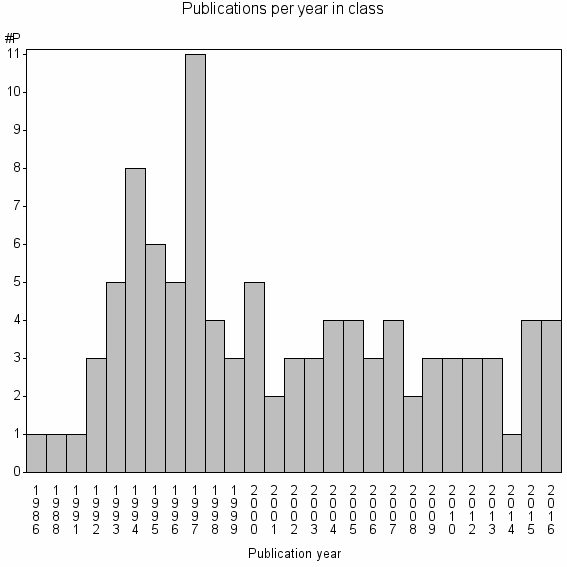
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 35297 | 99 | 30.0 | 77% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 419 | 3 | ADENOVIRUS//MERKEL CELL CARCINOMA//GENE THERAPY | 28518 |
| 564 | 2 | GENE THERAPY//ADENOVIRUS//HUMAN GENE THERAPY | 14456 |
| 35297 | 1 | S GAL//3 D REGISTRATION AND RECONSTRUCTION//3 GALACTOSIDASE | 99 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | S GAL | authKW | 411253 | 2% | 67% | 2 |
| 2 | 3 D REGISTRATION AND RECONSTRUCTION | authKW | 308441 | 1% | 100% | 1 |
| 3 | 3 GALACTOSIDASE | authKW | 308441 | 1% | 100% | 1 |
| 4 | ADSV40 BETA GAL | authKW | 308441 | 1% | 100% | 1 |
| 5 | BETA GALACTOSIDASE HISTOCHEMISTRY | authKW | 308441 | 1% | 100% | 1 |
| 6 | BLUO GAL | authKW | 308441 | 1% | 100% | 1 |
| 7 | CACO 2 HT29GLUCH COCULTURE | authKW | 308441 | 1% | 100% | 1 |
| 8 | CHIMERIC NUCLEOTIDES | authKW | 308441 | 1% | 100% | 1 |
| 9 | CRYOJANE FROZEN SECTION | authKW | 308441 | 1% | 100% | 1 |
| 10 | CRYPTIC SPLICE ACCEPTOR SITES | authKW | 308441 | 1% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Cell Biology | 221 | 25% | 0% | 25 |
| 2 | Cell & Tissue Engineering | 180 | 4% | 0% | 4 |
| 3 | Biochemical Research Methods | 161 | 14% | 0% | 14 |
| 4 | Biotechnology & Applied Microbiology | 144 | 17% | 0% | 17 |
| 5 | Anatomy & Morphology | 126 | 5% | 0% | 5 |
| 6 | Medicine, Research & Experimental | 125 | 14% | 0% | 14 |
| 7 | Genetics & Heredity | 82 | 13% | 0% | 13 |
| 8 | Microscopy | 72 | 3% | 0% | 3 |
| 9 | Food Science & Technology | 49 | 9% | 0% | 9 |
| 10 | Gastroenterology & Hepatology | 43 | 7% | 0% | 7 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MUSEUM ANAT | 308441 | 1% | 100% | 1 |
| 2 | TOKYO MED UNIV ANIM | 308441 | 1% | 100% | 1 |
| 3 | AGR BIOPROC 103 | 102812 | 1% | 33% | 1 |
| 4 | HUMAN NORMAL ANAT | 102812 | 1% | 33% | 1 |
| 5 | ANDREW L WARSHAW MD PANCREAT CANC | 77109 | 1% | 25% | 1 |
| 6 | SECT HUMAN NORMAL ANAT | 77109 | 1% | 25% | 1 |
| 7 | FUNCT FOODS HLTH PROGRAM | 44061 | 1% | 14% | 1 |
| 8 | TRANSFERT GENES | 30842 | 1% | 10% | 1 |
| 9 | NEUROL DISABIL IDINE | 20561 | 1% | 7% | 1 |
| 10 | REG BIOMED CRIB | 17134 | 1% | 6% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | HUMAN GENE THERAPY | 7491 | 9% | 0% | 9 |
| 2 | CELL PRESERVATION TECHNOLOGY | 5316 | 1% | 2% | 1 |
| 3 | JOURNAL OF HISTOCHEMISTRY & CYTOCHEMISTRY | 2600 | 7% | 0% | 7 |
| 4 | FOOD TECHNOLOGY | 2344 | 4% | 0% | 4 |
| 5 | CEREAL FOODS WORLD | 1983 | 3% | 0% | 3 |
| 6 | TRANSGENIC RESEARCH | 1661 | 3% | 0% | 3 |
| 7 | HISTOLOGY AND HISTOPATHOLOGY | 1334 | 4% | 0% | 4 |
| 8 | JOURNAL OF THE CANADIAN DIETETIC ASSOCIATION-REVUE DE L ASSOCIATION CANADIENNE DES DIETETISTES | 1104 | 1% | 0% | 1 |
| 9 | HISTOCHEMISTRY AND CELL BIOLOGY | 1056 | 3% | 0% | 3 |
| 10 | ANATOMICAL SCIENCE INTERNATIONAL | 717 | 1% | 0% | 1 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | S GAL | 411253 | 2% | 67% | 2 | Search S+GAL | Search S+GAL |
| 2 | 3 D REGISTRATION AND RECONSTRUCTION | 308441 | 1% | 100% | 1 | Search 3+D+REGISTRATION+AND+RECONSTRUCTION | Search 3+D+REGISTRATION+AND+RECONSTRUCTION |
| 3 | 3 GALACTOSIDASE | 308441 | 1% | 100% | 1 | Search 3+GALACTOSIDASE | Search 3+GALACTOSIDASE |
| 4 | ADSV40 BETA GAL | 308441 | 1% | 100% | 1 | Search ADSV40+BETA+GAL | Search ADSV40+BETA+GAL |
| 5 | BETA GALACTOSIDASE HISTOCHEMISTRY | 308441 | 1% | 100% | 1 | Search BETA+GALACTOSIDASE+HISTOCHEMISTRY | Search BETA+GALACTOSIDASE+HISTOCHEMISTRY |
| 6 | BLUO GAL | 308441 | 1% | 100% | 1 | Search BLUO+GAL | Search BLUO+GAL |
| 7 | CACO 2 HT29GLUCH COCULTURE | 308441 | 1% | 100% | 1 | Search CACO+2+HT29GLUCH+COCULTURE | Search CACO+2+HT29GLUCH+COCULTURE |
| 8 | CHIMERIC NUCLEOTIDES | 308441 | 1% | 100% | 1 | Search CHIMERIC+NUCLEOTIDES | Search CHIMERIC+NUCLEOTIDES |
| 9 | CRYOJANE FROZEN SECTION | 308441 | 1% | 100% | 1 | Search CRYOJANE+FROZEN+SECTION | Search CRYOJANE+FROZEN+SECTION |
| 10 | CRYPTIC SPLICE ACCEPTOR SITES | 308441 | 1% | 100% | 1 | Search CRYPTIC+SPLICE+ACCEPTOR+SITES | Search CRYPTIC+SPLICE+ACCEPTOR+SITES |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | SHIMADA, A , KOMATSU, K , NAKASHIMA, K , POSCHL, E , NIFUJI, A , (2012) IMPROVED METHODS FOR DETECTION OF BETA-GALACTOSIDASE (LACZ) ACTIVITY IN HARD TISSUE.HISTOCHEMISTRY AND CELL BIOLOGY. VOL. 137. ISSUE 6. P. 841 -847 | 10 | 48% | 2 |
| 2 | BELL, P , LIMBERIS, M , GAO, GP , WU, D , BOVE, MS , SANMIGUEL, JC , WILSON, JM , (2005) AN OPTIMIZED PROTOCOL FOR DETECTION OF E-COLI BETA-GALACTOSIDASE IN LUNG TISSUE FOLLOWING GENE TRANSFER.HISTOCHEMISTRY AND CELL BIOLOGY. VOL. 124. ISSUE 1. P. 77-85 | 11 | 44% | 30 |
| 3 | TRIFONOV, S , YAMASHITA, Y , KASE, M , MARUYAMA, M , SUGIMOTO, T , (2016) OVERVIEW AND ASSESSMENT OF THE HISTOCHEMICAL METHODS AND REAGENTS FOR THE DETECTION OF BETA-GALACTOSIDASE ACTIVITY IN TRANSGENIC ANIMALS.ANATOMICAL SCIENCE INTERNATIONAL. VOL. 91. ISSUE 1. P. 56 -67 | 10 | 37% | 0 |
| 4 | KOPP, HG , HOOPER, AT , SHMELKOV, SV , RAFII, S , (2007) BETA-GALACTOSIDASE STAINING ON BONE MARROW. THE OSTEOCLAST PITFALL.HISTOLOGY AND HISTOPATHOLOGY. VOL. 22. ISSUE 9. P. 971 -976 | 6 | 50% | 21 |
| 5 | GIOGLIO, L , DE ANGELIS, MGC , BORATTO, R , POGGI, P , (2002) AN IMPROVED METHOD FOR BETA-GALACTOSIDASE ACTIVITY DETECTION ON MUSCLE TISSUE. A LIGHT AND ELECTRON MICROSCOPIC STUDY.ANNALS OF ANATOMY-ANATOMISCHER ANZEIGER. VOL. 184. ISSUE 2. P. 153-157 | 5 | 63% | 4 |
| 6 | RAVAI, M , (1996) QUALITY CHARACTERISTICS OF RASPBERRIES AND BLACKBERRIES.CEREAL FOODS WORLD. VOL. 41. ISSUE 10. P. 772-775 | 3 | 100% | 2 |
| 7 | BOLON, B , (2008) WHOLE MOUNT ENZYME HISTOCHEMISTRY AS A RAPID SCREEN AT NECROPSY FOR EXPRESSION OF BETA-GALACTOSIDASE (LACZ)-BEARING TRANSGENES: CONSIDERATIONS FOR SEPARATING SPECIFIC LACZ ACTIVITY FROM NONSPECIFIC (ENDOGENOUS) GALACTOSIDASE ACTIVITY.TOXICOLOGIC PATHOLOGY. VOL. 36. ISSUE 2. P. 265 -276 | 8 | 25% | 8 |
| 8 | MERKWITZ, C , BLASCHUK, O , SCHULZ, A , RICKEN, AM , (2016) COMMENTS ON METHODS TO SUPPRESS ENDOGENOUS -GALACTOSIDASE ACTIVITY IN MOUSE TISSUES EXPRESSING THE LACZ REPORTER GENE.JOURNAL OF HISTOCHEMISTRY & CYTOCHEMISTRY. VOL. 64. ISSUE 10. P. 579 -586 | 3 | 50% | 0 |
| 9 | AOYAMA, N , MOLIN, DGM , MENTINK, MMT , KOERTEN, HK , DE RUITER, MC , GITTENBERGER-DE GROOT, AC , POELMANN, RE , (2004) CHANGING INTRACELLULAR COMPARTMENTALIZATION OF BETA-GALACTOSIDASE IN THE ROSA26 REPORTER MOUSE DURING EMBRYONIC DEVELOPMENT: A LIGHT- AND ELECTRON-MICROSCOPIC STUDY.ANATOMICAL RECORD PART A-DISCOVERIES IN MOLECULAR CELLULAR AND EVOLUTIONARY BIOLOGY. VOL. 279A. ISSUE 2. P. 740-748 | 7 | 28% | 4 |
| 10 | BUCKINX, R , TIMMERMANS, JP , (2016) TARGETING THE GASTROINTESTINAL TRACT WITH VIRAL VECTORS: STATE OF THE ART AND POSSIBLE APPLICATIONS IN RESEARCH AND THERAPY.HISTOCHEMISTRY AND CELL BIOLOGY. VOL. 146. ISSUE 6. P. 709 -720 | 10 | 13% | 1 |
Classes with closest relation at Level 1 |