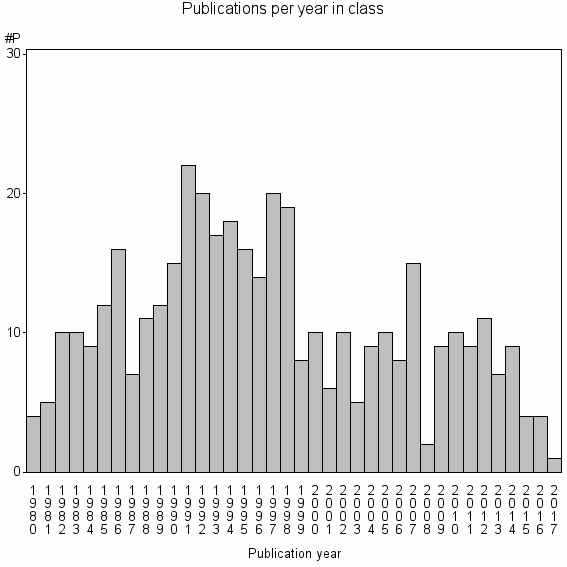
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 21813 | 404 | 34.9 | 59% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 427 | 3 | JOURNAL OF THE INSTITUTE OF BREWING//KERATINASE//CYSTATIN | 27946 |
| 1547 | 2 | KERATINASE//ALKALINE PROTEASE//ASPARTIC PROTEINASE | 7420 |
| 21813 | 1 | THERMOLYSIN//NEUTRAL PROTEASE//HALOPHILICITY | 404 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | THERMOLYSIN | authKW | 1191603 | 15% | 26% | 61 |
| 2 | NEUTRAL PROTEASE | authKW | 383089 | 5% | 24% | 21 |
| 3 | HALOPHILICITY | authKW | 302323 | 1% | 67% | 6 |
| 4 | BOVINE INTESTINE ALKALINE PHOSPHATASE | authKW | 226745 | 1% | 100% | 3 |
| 5 | FAB2 MU FRAGMENT | authKW | 226745 | 1% | 100% | 3 |
| 6 | HYDROPHOBIC INTERACTION HPLC | authKW | 170057 | 1% | 75% | 3 |
| 7 | MATRIX METALLOPROTEINASE MMP 7 | authKW | 170057 | 1% | 75% | 3 |
| 8 | SALT ACTIVATION | authKW | 168332 | 2% | 32% | 7 |
| 9 | FAB2 FRAGMENT | authKW | 160048 | 1% | 35% | 6 |
| 10 | GLYCYL D PHENYLALANINE | authKW | 151163 | 0% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 2800 | 67% | 0% | 272 |
| 2 | Biotechnology & Applied Microbiology | 637 | 18% | 0% | 72 |
| 3 | Biophysics | 545 | 14% | 0% | 57 |
| 4 | Food Science & Technology | 271 | 10% | 0% | 42 |
| 5 | Biochemical Research Methods | 268 | 9% | 0% | 38 |
| 6 | Chemistry, Applied | 112 | 6% | 0% | 23 |
| 7 | Microbiology | 107 | 8% | 0% | 33 |
| 8 | Agronomy | 10 | 2% | 0% | 8 |
| 9 | Chemistry, Medicinal | 9 | 2% | 0% | 9 |
| 10 | Cell Biology | 4 | 4% | 0% | 16 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BRANCH BISCUITS | 75582 | 0% | 100% | 1 |
| 2 | BRANCHE BOISSONS | 75582 | 0% | 100% | 1 |
| 3 | DANISCO US INC | 75582 | 0% | 100% | 1 |
| 4 | EXPTL PATHOL CHIGUSA KU | 75582 | 0% | 100% | 1 |
| 5 | GENENCOR INT BV | 75582 | 0% | 100% | 1 |
| 6 | GRONINGEN BIOMOL SCI BIOTECCHNOL | 75582 | 0% | 100% | 1 |
| 7 | GRP RECH BIOMOL MICROENVRIONM METAB | 75582 | 0% | 100% | 1 |
| 8 | INSERM U 314 | 75582 | 0% | 100% | 1 |
| 9 | ISSBM | 75582 | 0% | 100% | 1 |
| 10 | MED BIOL MOL STRUCT BIOL | 75582 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF BIOCHEMISTRY | 12428 | 10% | 0% | 42 |
| 2 | PROTEIN ENGINEERING | 8541 | 4% | 1% | 16 |
| 3 | JOURNAL OF FERMENTATION AND BIOENGINEERING | 2760 | 2% | 0% | 9 |
| 4 | BIOSCIENCE BIOTECHNOLOGY AND BIOCHEMISTRY | 2491 | 5% | 0% | 20 |
| 5 | INTERNATIONAL JOURNAL OF PEPTIDE AND PROTEIN RESEARCH | 2431 | 2% | 0% | 9 |
| 6 | MOLECULAR BIOLOGY | 1291 | 2% | 0% | 9 |
| 7 | JOURNAL OF BIOCHEMICAL AND BIOPHYSICAL METHODS | 973 | 1% | 0% | 5 |
| 8 | BIOCHIMICA ET BIOPHYSICA ACTA-PROTEIN STRUCTURE AND MOLECULAR ENZYMOLOGY | 867 | 1% | 0% | 5 |
| 9 | SEIBUTSU-KOGAKU KAISHI-JOURNAL OF THE SOCIETY FOR FERMENTATION AND BIOENGINEERING | 746 | 0% | 1% | 1 |
| 10 | ADVANCES IN BIOPHYSICS | 725 | 0% | 1% | 1 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | THERMOLYSIN | 1191603 | 15% | 26% | 61 | Search THERMOLYSIN | Search THERMOLYSIN |
| 2 | NEUTRAL PROTEASE | 383089 | 5% | 24% | 21 | Search NEUTRAL+PROTEASE | Search NEUTRAL+PROTEASE |
| 3 | HALOPHILICITY | 302323 | 1% | 67% | 6 | Search HALOPHILICITY | Search HALOPHILICITY |
| 4 | BOVINE INTESTINE ALKALINE PHOSPHATASE | 226745 | 1% | 100% | 3 | Search BOVINE+INTESTINE+ALKALINE+PHOSPHATASE | Search BOVINE+INTESTINE+ALKALINE+PHOSPHATASE |
| 5 | FAB2 MU FRAGMENT | 226745 | 1% | 100% | 3 | Search FAB2+MU+FRAGMENT | Search FAB2+MU+FRAGMENT |
| 6 | HYDROPHOBIC INTERACTION HPLC | 170057 | 1% | 75% | 3 | Search HYDROPHOBIC+INTERACTION+HPLC | Search HYDROPHOBIC+INTERACTION+HPLC |
| 7 | MATRIX METALLOPROTEINASE MMP 7 | 170057 | 1% | 75% | 3 | Search MATRIX+METALLOPROTEINASE+MMP+7 | Search MATRIX+METALLOPROTEINASE+MMP+7 |
| 8 | SALT ACTIVATION | 168332 | 2% | 32% | 7 | Search SALT+ACTIVATION | Search SALT+ACTIVATION |
| 9 | FAB2 FRAGMENT | 160048 | 1% | 35% | 6 | Search FAB2+FRAGMENT | Search FAB2+FRAGMENT |
| 10 | GLYCYL D PHENYLALANINE | 151163 | 0% | 100% | 2 | Search GLYCYL+D+PHENYLALANINE | Search GLYCYL+D+PHENYLALANINE |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | KUSANO, M , YASUKAWA, K , INOUYE, K , (2009) INSIGHTS INTO THE CATALYTIC ROLES OF THE POLYPEPTIDE REGIONS IN THE ACTIVE SITE OF THERMOLYSIN AND GENERATION OF THE THERMOLYSIN VARIANTS WITH HIGH ACTIVITY AND STABILITY.JOURNAL OF BIOCHEMISTRY. VOL. 145. ISSUE 1. P. 103 -113 | 30 | 83% | 6 |
| 2 | KOJIMA, K , NAKATA, H , INOUYE, K , (2014) INVOLVEMENT OF VAL 315 LOCATED IN THE C-TERMINAL REGION OF THERMOLYSIN IN ITS EXPRESSION IN ESCHERICHIA COLI AND ITS THERMAL STABILITY.BIOCHIMICA ET BIOPHYSICA ACTA-PROTEINS AND PROTEOMICS. VOL. 1844. ISSUE 2. P. 330-338 | 25 | 83% | 2 |
| 3 | TATSUMI, C , HASHIDA, Y , YASUKAWA, K , INOUYE, K , (2007) EFFECTS OF SITE-DIRECTED MUTAGENESIS OF THE SURFACE RESIDUES GLN128 AND GLN225 OF THERMOLYSIN ON ITS CATALYTIC ACTIVITY.JOURNAL OF BIOCHEMISTRY. VOL. 141. ISSUE 6. P. 835-842 | 29 | 81% | 5 |
| 4 | YASUKAWA, K , INOUYE, K , (2007) IMPROVING THE ACTIVITY AND STABILITY OF THERMOLYSIN BY SITE-DIRECTED MUTAGENESIS.BIOCHIMICA ET BIOPHYSICA ACTA-PROTEINS AND PROTEOMICS. VOL. 1774. ISSUE 10. P. 1281-1288 | 27 | 82% | 18 |
| 5 | MENACH, E , YASUKAWA, K , INOUYE, K , (2010) EFFECTS OF SITE-DIRECTED MUTAGENESIS OF THE LOOP RESIDUE OF THE N-TERMINAL DOMAIN GLY117 OF THERMOLYSIN ON ITS CATALYTIC ACTIVITY.BIOSCIENCE BIOTECHNOLOGY AND BIOCHEMISTRY. VOL. 74. ISSUE 12. P. 2457 -2462 | 25 | 83% | 4 |
| 6 | HASHIDA, Y , INOUYE, K , (2007) MOLECULAR MECHANISM OF THE INHIBITORY EFFECT OF COBALT ION ON THERMOLYSIN ACTIVITY AND THE SUPPRESSIVE EFFECT OF CALCIUM ION ON THE COBALT ION-DEPENDENT INACTIVATION OF THERMOLYSIN.JOURNAL OF BIOCHEMISTRY. VOL. 141. ISSUE 6. P. 879-888 | 26 | 81% | 5 |
| 7 | TAKITA, T , AONO, T , SAKURAMA, H , ITOH, T , WADA, T , MINODA, M , YASUKAWA, K , INOUYE, K , (2008) EFFECTS OF INTRODUCING NEGATIVE CHARGES INTO THE MOLECULAR SURFACE OF THERMOLYSIN BY SITE-DIRECTED MUTAGENESIS ON ITS ACTIVITY AND STABILITY.BIOCHIMICA ET BIOPHYSICA ACTA-PROTEINS AND PROTEOMICS. VOL. 1784. ISSUE 3. P. 481 -488 | 24 | 77% | 8 |
| 8 | KAWASAKI, Y , YASUKAWA, K , INOUYE, K , (2013) EFFECTS OF SITE-DIRECTED MUTAGENESIS IN THE N-TERMINAL DOMAIN OF THERMOLYSIN ON ITS STABILIZATION.JOURNAL OF BIOCHEMISTRY. VOL. 153. ISSUE 1. P. 85-92 | 21 | 75% | 4 |
| 9 | MENACH, E , YASUKAWA, K , INOUYE, K , (2013) EFFECTS OF MUTATIONS OF THERMOLYSIN, ASN116 TO ASP AND ASP150 TO GLU, ON SALT-INDUCED ACTIVATION AND STABILIZATION.BIOSCIENCE BIOTECHNOLOGY AND BIOCHEMISTRY. VOL. 77. ISSUE 4. P. 741 -746 | 18 | 75% | 1 |
| 10 | HASHIDA, Y , INOUYE, K , (2007) KINETIC ANALYSIS OF THE ACTIVATION-AND-INHIBITION DUAL EFFECTS OF COBALT ION ON THERMOLYSIN ACTIVITY.JOURNAL OF BIOCHEMISTRY. VOL. 141. ISSUE 6. P. 843-853 | 23 | 64% | 9 |
Classes with closest relation at Level 1 |