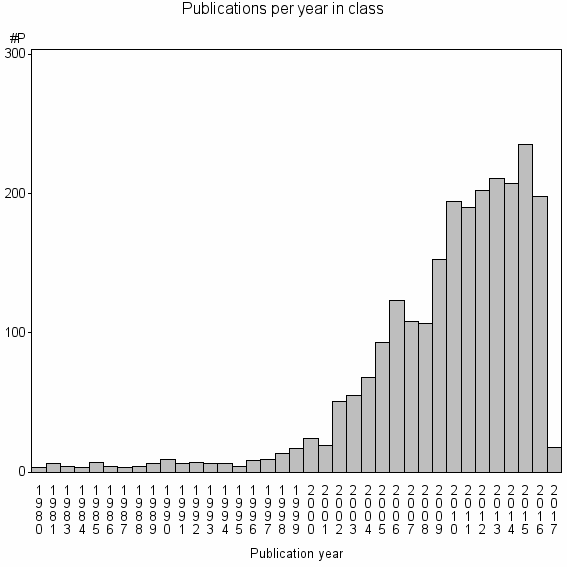
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 2029 | 2381 | 46.8 | 80% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 548 | 2 | BMC SYSTEMS BIOLOGY//MATHEMATICAL & COMPUTATIONAL BIOLOGY//BOOLEAN NETWORKS | 14655 |
| 2029 | 1 | FLUX BALANCE ANALYSIS//METABOLIC NETWORK//CONSTRAINT BASED MODELING | 2381 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | FLUX BALANCE ANALYSIS | authKW | 1439746 | 7% | 63% | 177 |
| 2 | METABOLIC NETWORK | authKW | 1006912 | 8% | 42% | 188 |
| 3 | CONSTRAINT BASED MODELING | authKW | 634622 | 3% | 71% | 70 |
| 4 | ELEMENTARY FLUX MODES | authKW | 381825 | 2% | 66% | 45 |
| 5 | GENOME SCALE METABOLIC MODEL | authKW | 364241 | 2% | 57% | 50 |
| 6 | ELEMENTARY MODES | authKW | 288498 | 1% | 75% | 30 |
| 7 | METABOLIC MODELING | authKW | 230813 | 2% | 33% | 55 |
| 8 | BMC SYSTEMS BIOLOGY | journal | 225611 | 7% | 10% | 168 |
| 9 | GENOME SCALE MODEL | authKW | 188407 | 1% | 64% | 23 |
| 10 | METABOLIC NETWORK RECONSTRUCTION | authKW | 131888 | 1% | 86% | 12 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Mathematical & Computational Biology | 88354 | 35% | 1% | 841 |
| 2 | Biotechnology & Applied Microbiology | 18671 | 38% | 0% | 898 |
| 3 | Biochemical Research Methods | 13254 | 25% | 0% | 601 |
| 4 | Biochemistry & Molecular Biology | 1291 | 23% | 0% | 543 |
| 5 | Biology | 1148 | 7% | 0% | 163 |
| 6 | Genetics & Heredity | 402 | 7% | 0% | 162 |
| 7 | Microbiology | 311 | 6% | 0% | 148 |
| 8 | Computer Science, Interdisciplinary Applications | 297 | 4% | 0% | 87 |
| 9 | Multidisciplinary Sciences | 97 | 1% | 0% | 32 |
| 10 | Biophysics | 78 | 3% | 0% | 77 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | COMPUTAT SYST BIOTECHNOL | 112268 | 1% | 49% | 18 |
| 2 | ARTIFICIAL INTELLIGENCE BIOINFORMAT GRP | 60179 | 1% | 36% | 13 |
| 3 | SYST BIOL MATH MODELING GRP | 58552 | 1% | 35% | 13 |
| 4 | CELL SYST MODELLING GRP | 51291 | 0% | 100% | 4 |
| 5 | IND ENGN MECH ENGN COMP SCI | 48849 | 1% | 21% | 18 |
| 6 | METAB BIOMOL ENGN | 42164 | 1% | 14% | 24 |
| 7 | BIOCOMP STRUCT PROGRAM | 38468 | 0% | 100% | 3 |
| 8 | INT MAX PLANCK COMPUTAT BIOL SCI COMP I | 38468 | 0% | 100% | 3 |
| 9 | MOL BIOMOL INFORMAT NCMLS | 38468 | 0% | 100% | 3 |
| 10 | GORTNER 240 | 37082 | 0% | 32% | 9 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BMC SYSTEMS BIOLOGY | 225611 | 7% | 10% | 168 |
| 2 | METABOLIC ENGINEERING | 46253 | 3% | 6% | 62 |
| 3 | MOLECULAR SYSTEMS BIOLOGY | 45767 | 2% | 7% | 52 |
| 4 | BIOINFORMATICS | 35883 | 7% | 2% | 173 |
| 5 | PLOS COMPUTATIONAL BIOLOGY | 31040 | 4% | 2% | 106 |
| 6 | IEE PROCEEDINGS SYSTEMS BIOLOGY | 18674 | 0% | 13% | 11 |
| 7 | BMC BIOINFORMATICS | 13411 | 4% | 1% | 89 |
| 8 | CURRENT OPINION IN BIOTECHNOLOGY | 11143 | 2% | 2% | 45 |
| 9 | BIOTECHNOLOGY JOURNAL | 10162 | 1% | 3% | 28 |
| 10 | MOLECULAR BIOSYSTEMS | 7152 | 2% | 1% | 39 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | FLUX BALANCE ANALYSIS | 1439746 | 7% | 63% | 177 | Search FLUX+BALANCE+ANALYSIS | Search FLUX+BALANCE+ANALYSIS |
| 2 | METABOLIC NETWORK | 1006912 | 8% | 42% | 188 | Search METABOLIC+NETWORK | Search METABOLIC+NETWORK |
| 3 | CONSTRAINT BASED MODELING | 634622 | 3% | 71% | 70 | Search CONSTRAINT+BASED+MODELING | Search CONSTRAINT+BASED+MODELING |
| 4 | ELEMENTARY FLUX MODES | 381825 | 2% | 66% | 45 | Search ELEMENTARY+FLUX+MODES | Search ELEMENTARY+FLUX+MODES |
| 5 | GENOME SCALE METABOLIC MODEL | 364241 | 2% | 57% | 50 | Search GENOME+SCALE+METABOLIC+MODEL | Search GENOME+SCALE+METABOLIC+MODEL |
| 6 | ELEMENTARY MODES | 288498 | 1% | 75% | 30 | Search ELEMENTARY+MODES | Search ELEMENTARY+MODES |
| 7 | METABOLIC MODELING | 230813 | 2% | 33% | 55 | Search METABOLIC+MODELING | Search METABOLIC+MODELING |
| 8 | GENOME SCALE MODEL | 188407 | 1% | 64% | 23 | Search GENOME+SCALE+MODEL | Search GENOME+SCALE+MODEL |
| 9 | METABOLIC NETWORK RECONSTRUCTION | 131888 | 1% | 86% | 12 | Search METABOLIC+NETWORK+RECONSTRUCTION | Search METABOLIC+NETWORK+RECONSTRUCTION |
| 10 | GENOME SCALE METABOLIC NETWORK | 120385 | 1% | 72% | 13 | Search GENOME+SCALE+METABOLIC+NETWORK | Search GENOME+SCALE+METABOLIC+NETWORK |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | LEWIS, NE , NAGARAJAN, H , PALSSON, BO , (2012) CONSTRAINING THE METABOLIC GENOTYPE-PHENOTYPE RELATIONSHIP USING A PHYLOGENY OF IN SILICO METHODS.NATURE REVIEWS MICROBIOLOGY. VOL. 10. ISSUE 4. P. 291 -305 | 124 | 89% | 205 |
| 2 | O'BRIEN, EJ , MONK, JM , PALSSON, BO , (2015) USING GENOME-SCALE MODELS TO PREDICT BIOLOGICAL CAPABILITIES.CELL. VOL. 161. ISSUE 5. P. 971 -987 | 79 | 90% | 33 |
| 3 | BOURGUIGNON, PY , SCHACHTER, V , DUROT, M , (2009) GENOME-SCALE MODELS OF BACTERIAL METABOLISM: RECONSTRUCTION AND APPLICATIONS.FEMS MICROBIOLOGY REVIEWS. VOL. 33. ISSUE 1. P. 164 -190 | 130 | 65% | 119 |
| 4 | BORDBAR, A , MONK, JM , KING, ZA , PALSSON, BO , (2014) CONSTRAINT-BASED MODELS PREDICT METABOLIC AND ASSOCIATED CELLULAR FUNCTIONS.NATURE REVIEWS GENETICS. VOL. 15. ISSUE 2. P. 107-120 | 74 | 73% | 111 |
| 5 | MAIA, P , ROCHA, M , ROCHA, I , (2016) IN SILICO CONSTRAINT-BASED STRAIN OPTIMIZATION METHODS: THE QUEST FOR OPTIMAL CELL FACTORIES.MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS. VOL. 80. ISSUE 1. P. 45 -67 | 110 | 71% | 2 |
| 6 | MCCLOSKEY, D , PALSSON, BO , FEIST, AM , (2013) BASIC AND APPLIED USES OF GENOME-SCALE METABOLIC NETWORK RECONSTRUCTIONS OF ESCHERICHIA COLI.MOLECULAR SYSTEMS BIOLOGY. VOL. 9. ISSUE . P. - | 92 | 70% | 90 |
| 7 | OBERHARDT, MA , PALSSON, BO , PAPIN, JA , (2009) APPLICATIONS OF GENOME-SCALE METABOLIC RECONSTRUCTIONS.MOLECULAR SYSTEMS BIOLOGY. VOL. 5. ISSUE . P. - | 94 | 62% | 299 |
| 8 | SCHELLENBERGER, J , QUE, R , FLEMING, RMT , THIELE, I , ORTH, JD , FEIST, AM , ZIELINSKI, DC , BORDBAR, A , LEWIS, NE , RAHMANIAN, S , ET AL (2011) QUANTITATIVE PREDICTION OF CELLULAR METABOLISM WITH CONSTRAINT-BASED MODELS: THE COBRA TOOLBOX V2.0.NATURE PROTOCOLS. VOL. 6. ISSUE 9. P. 1290-1307 | 55 | 83% | 343 |
| 9 | PRICE, ND , REED, JL , PALSSON, BO , (2004) GENOME-SCALE MODELS OF MICROBIAL CELLS: EVALUATING THE CONSEQUENCES OF CONSTRAINTS.NATURE REVIEWS MICROBIOLOGY. VOL. 2. ISSUE 11. P. 886-897 | 74 | 77% | 567 |
| 10 | FEIST, AM , PALSSON, BO , (2008) THE GROWING SCOPE OF APPLICATIONS OF GENOME-SCALE METABOLIC RECONSTRUCTIONS USING ESCHERICHIA COLI.NATURE BIOTECHNOLOGY. VOL. 26. ISSUE 6. P. 659 -667 | 73 | 74% | 259 |
Classes with closest relation at Level 1 |