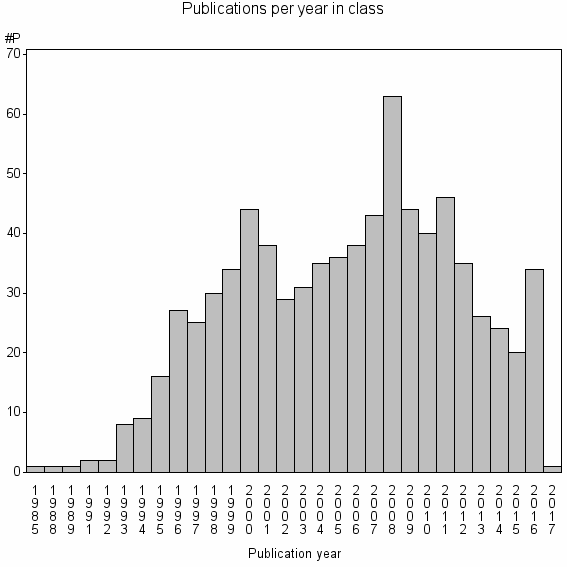
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 14228 | 783 | 38.7 | 90% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 478 | 3 | PHAGE DISPLAY//CELL FREE PROTEIN SYNTHESIS//ANTIBODY ENGINEERING | 23687 |
| 1537 | 2 | CELL FREE PROTEIN SYNTHESIS//INTEIN//PROTEIN SPLICING | 7453 |
| 14228 | 1 | BIMOLECULAR FLUORESCENCE COMPLEMENTATION//SPLIT UBIQUITIN//PROTEIN FRAGMENT COMPLEMENTATION ASSAY | 783 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | BIMOLECULAR FLUORESCENCE COMPLEMENTATION | authKW | 278671 | 3% | 26% | 27 |
| 2 | SPLIT UBIQUITIN | authKW | 246543 | 2% | 45% | 14 |
| 3 | PROTEIN FRAGMENT COMPLEMENTATION ASSAY | authKW | 244076 | 2% | 48% | 13 |
| 4 | BIFC | authKW | 192760 | 3% | 21% | 23 |
| 5 | PROTEIN FRAGMENT COMPLEMENTATION | authKW | 157272 | 1% | 37% | 11 |
| 6 | BIMOLECULAR FLUORESCENCE COMPLEMENTATION BIFC | authKW | 149974 | 1% | 38% | 10 |
| 7 | MASPIT | authKW | 116990 | 0% | 100% | 3 |
| 8 | SPLIT REPORTERS | authKW | 116990 | 0% | 100% | 3 |
| 9 | INTERACTION TRAP | authKW | 89131 | 1% | 57% | 4 |
| 10 | PROTEIN COMPLEMENTATION ASSAY | authKW | 81237 | 1% | 42% | 5 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemical Research Methods | 6303 | 30% | 0% | 236 |
| 2 | Biochemistry & Molecular Biology | 3307 | 54% | 0% | 421 |
| 3 | Biotechnology & Applied Microbiology | 652 | 13% | 0% | 105 |
| 4 | Cell Biology | 418 | 14% | 0% | 106 |
| 5 | Chemistry, Analytical | 398 | 11% | 0% | 86 |
| 6 | Biophysics | 153 | 6% | 0% | 48 |
| 7 | Genetics & Heredity | 95 | 6% | 0% | 47 |
| 8 | Microbiology | 39 | 4% | 0% | 35 |
| 9 | Plant Sciences | 38 | 5% | 0% | 37 |
| 10 | Mycology | 26 | 1% | 0% | 7 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ADV DISCOVERY SCI | 38997 | 0% | 100% | 1 |
| 2 | BIO3 BIOINFORMAT MOL GENET BIOL | 38997 | 0% | 100% | 1 |
| 3 | BIOENGNBIO X PROGRAMMIPS | 38997 | 0% | 100% | 1 |
| 4 | BIOL MICROBIOL GENET | 38997 | 0% | 100% | 1 |
| 5 | BIOMOL ENGN SURUGA KU | 38997 | 0% | 100% | 1 |
| 6 | BIOPROC DEV MFG | 38997 | 0% | 100% | 1 |
| 7 | BOT BOT GARTEN MOL ENTWICKLUNGSBIOL PFLANZ | 38997 | 0% | 100% | 1 |
| 8 | CANC E17124 | 38997 | 0% | 100% | 1 |
| 9 | CELL DEV BIOL WORKING GRP | 38997 | 0% | 100% | 1 |
| 10 | CELLULAR BIOCHEM ZMBE | 38997 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOTECHNIQUES | 16050 | 6% | 1% | 50 |
| 2 | METHODS | 5112 | 3% | 1% | 20 |
| 3 | NATURE METHODS | 3874 | 2% | 1% | 13 |
| 4 | CURRENT OPINION IN BIOTECHNOLOGY | 2405 | 2% | 1% | 12 |
| 5 | METHODS IN MICROBIOLOGY | 2221 | 1% | 1% | 5 |
| 6 | NATURE BIOTECHNOLOGY | 2124 | 2% | 0% | 12 |
| 7 | METHODS IN ENZYMOLOGY | 2112 | 4% | 0% | 30 |
| 8 | ACS CHEMICAL BIOLOGY | 1797 | 1% | 0% | 10 |
| 9 | ANALYTICAL BIOCHEMISTRY | 1739 | 4% | 0% | 29 |
| 10 | NATURE PROTOCOLS | 1380 | 1% | 0% | 9 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | STYNEN, B , TOURNU, H , TAVERNIER, J , VAN DIJCK, P , (2012) DIVERSITY IN GENETIC IN VIVO METHODS FOR PROTEIN-PROTEIN INTERACTION STUDIES: FROM THE YEAST TWO-HYBRID SYSTEM TO THE MAMMALIAN SPLIT-LUCIFERASE SYSTEM.MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS. VOL. 76. ISSUE 2. P. 331 -382 | 188 | 28% | 50 |
| 2 | WEHR, MC , ROSSNER, MJ , (2016) SPLIT PROTEIN BIOSENSOR ASSAYS IN MOLECULAR PHARMACOLOGICAL STUDIES.DRUG DISCOVERY TODAY. VOL. 21. ISSUE 3. P. 415 -429 | 48 | 53% | 3 |
| 3 | JOHNSSON, N , MULLER, J , (2008) SPLIT-UBIQUITIN AND THE SPLIT-PROTEIN SENSORS: CHESSMAN FOR THE ENDGAME.CHEMBIOCHEM. VOL. 9. ISSUE 13. P. 2029 -2038 | 46 | 65% | 21 |
| 4 | MICHNICK, SW , EAR, PH , MANDERSON, EN , REMY, I , STEFAN, E , (2007) UNIVERSAL STRATEGIES IN RESEARCH AND DRUG DISCOVERY BASED ON PROTEIN-FRAGMENT COMPLEMENTATION ASSAYS.NATURE REVIEWS DRUG DISCOVERY. VOL. 6. ISSUE 7. P. 569 -582 | 46 | 48% | 161 |
| 5 | VENTURA, S , (2011) BIMOLECULAR FLUORESCENCE COMPLEMENTATION: ILLUMINATING CELLULAR PROTEIN INTERACTIONS.CURRENT MOLECULAR MEDICINE. VOL. 11. ISSUE 7. P. 582-598 | 48 | 51% | 7 |
| 6 | SHEKHAWAT, SS , GHOSH, I , (2011) SPLIT-PROTEIN SYSTEMS: BEYOND BINARY PROTEIN-PROTEIN INTERACTIONS.CURRENT OPINION IN CHEMICAL BIOLOGY. VOL. 15. ISSUE 6. P. 789 -797 | 33 | 60% | 49 |
| 7 | MORELL, M , VENTURA, S , AVILES, FX , (2009) PROTEIN COMPLEMENTATION ASSAYS: APPROACHES FOR THE IN VIVO ANALYSIS OF PROTEIN INTERACTIONS.FEBS LETTERS. VOL. 583. ISSUE 11. P. 1684-1691 | 38 | 60% | 23 |
| 8 | ZHU, ZX , YU, ZM , TAYLOR, JL , WU, YH , NI, J , (2016) THE APPLICATION OF YEAST HYBRID SYSTEMS IN PROTEIN INTERACTION ANALYSIS.MOLECULAR BIOLOGY. VOL. 50. ISSUE 5. P. 663 -670 | 29 | 71% | 0 |
| 9 | SUTER, B , KITTANAKOM, S , STAGLJAR, I , (2008) TWO-HYBRID TECHNOLOGIES IN PROTEOMICS RESEARCH.CURRENT OPINION IN BIOTECHNOLOGY. VOL. 19. ISSUE 4. P. 316-323 | 37 | 59% | 57 |
| 10 | MILLER, KE , KIM, Y , HUH, WK , PARK, HO , (2015) BIMOLECULAR FLUORESCENCE COMPLEMENTATION (BIFC) ANALYSIS: ADVANCES AND RECENT APPLICATIONS FOR GENOME-WIDE INTERACTION STUDIES.JOURNAL OF MOLECULAR BIOLOGY. VOL. 427. ISSUE 11. P. 2039 -2055 | 40 | 36% | 16 |
Classes with closest relation at Level 1 |