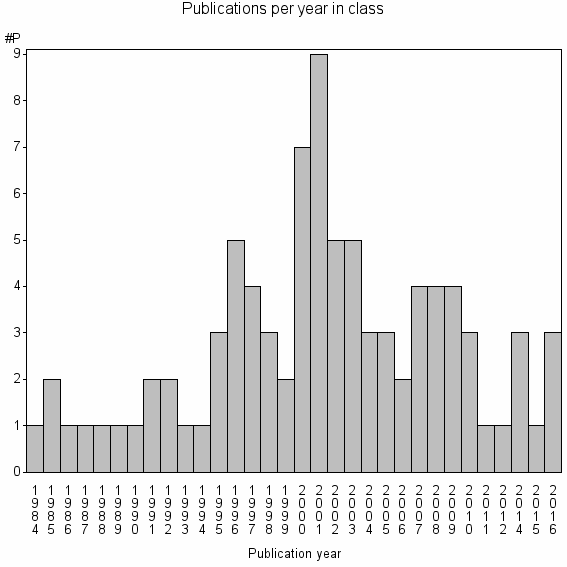
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 35939 | 89 | 25.9 | 72% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 4 | 4 | ENVIRONMENTAL SCIENCES//GEOSCIENCES, MULTIDISCIPLINARY//METEOROLOGY & ATMOSPHERIC SCIENCES | 1766162 |
| 50 | 3 | ANAEROBIC DIGESTION//MICROBIAL FUEL CELL//WATER SCIENCE AND TECHNOLOGY | 91931 |
| 1312 | 2 | MICROBIOLOGY//PLANCTOMYCETES//PROPIDIUM MONOAZIDE | 8512 |
| 35939 | 1 | ACTIVATED BIOMASS//EFFLUENT TREATMENT PLANT ETP//STAIRCASE ELECTROPHORESIS | 89 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | ACTIVATED BIOMASS | authKW | 857738 | 6% | 50% | 5 |
| 2 | EFFLUENT TREATMENT PLANT ETP | authKW | 686194 | 2% | 100% | 2 |
| 3 | STAIRCASE ELECTROPHORESIS | authKW | 686194 | 2% | 100% | 2 |
| 4 | UNIDENTIFIED BACTERIA | authKW | 686194 | 2% | 100% | 2 |
| 5 | CATABOLIC POTENTIAL | authKW | 617573 | 3% | 60% | 3 |
| 6 | ENVIRONM MODELING GENOM | address | 514643 | 3% | 50% | 3 |
| 7 | LMW RNA | authKW | 441122 | 3% | 43% | 3 |
| 8 | ENVIRONM GENOM UNIT | address | 408431 | 11% | 12% | 10 |
| 9 | 5S RRNA SCAFFOLD | authKW | 343097 | 1% | 100% | 1 |
| 10 | ANTI DNA RNA ANTIBODY | authKW | 343097 | 1% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biotechnology & Applied Microbiology | 1169 | 48% | 0% | 43 |
| 2 | Microbiology | 878 | 42% | 0% | 37 |
| 3 | Agricultural Engineering | 426 | 9% | 0% | 8 |
| 4 | Biochemical Research Methods | 388 | 22% | 0% | 20 |
| 5 | Mathematical & Computational Biology | 153 | 8% | 0% | 7 |
| 6 | Energy & Fuels | 59 | 9% | 0% | 8 |
| 7 | Engineering, Environmental | 22 | 4% | 0% | 4 |
| 8 | Chemistry, Analytical | 20 | 8% | 0% | 7 |
| 9 | Environmental Sciences | 14 | 8% | 0% | 7 |
| 10 | Horticulture | 13 | 2% | 0% | 2 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ENVIRONM MODELING GENOM | 514643 | 3% | 50% | 3 |
| 2 | ENVIRONM GENOM UNIT | 408431 | 11% | 12% | 10 |
| 3 | ENVIRONM GEN UNIT | 343097 | 1% | 100% | 1 |
| 4 | ENVIRONM MODELING GENOMICS | 343097 | 1% | 100% | 1 |
| 5 | GENOM INTEGRAT BIOL IGIB | 171548 | 1% | 50% | 1 |
| 6 | GENOME INTEGRAT BIOL | 114364 | 1% | 33% | 1 |
| 7 | 209 | 114356 | 6% | 7% | 5 |
| 8 | EDIFICIO DEPARTMENTAL | 85773 | 1% | 25% | 1 |
| 9 | EDIFICIO | 56421 | 6% | 3% | 5 |
| 10 | ENVIRONM GENOM | 34294 | 9% | 1% | 8 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | SYSTEMATIC AND APPLIED MICROBIOLOGY | 3468 | 6% | 0% | 5 |
| 2 | JOURNAL OF MICROBIOLOGICAL METHODS | 1773 | 6% | 0% | 5 |
| 3 | CURRENT MICROBIOLOGY | 1482 | 6% | 0% | 5 |
| 4 | BIORESOURCE TECHNOLOGY | 1269 | 9% | 0% | 8 |
| 5 | BIOENGINEERED | 1259 | 1% | 0% | 1 |
| 6 | WILEY INTERDISCIPLINARY REVIEWS-RNA | 933 | 1% | 0% | 1 |
| 7 | APPLIED AND ENVIRONMENTAL MICROBIOLOGY | 926 | 10% | 0% | 9 |
| 8 | JOURNAL OF COMPUTATIONAL BIOLOGY | 877 | 2% | 0% | 2 |
| 9 | METHODS IN MICROBIOLOGY | 783 | 1% | 0% | 1 |
| 10 | ANTONIE VAN LEEUWENHOEK JOURNAL OF MICROBIOLOGY | 600 | 1% | 0% | 1 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ACTIVATED BIOMASS | 857738 | 6% | 50% | 5 | Search ACTIVATED+BIOMASS | Search ACTIVATED+BIOMASS |
| 2 | EFFLUENT TREATMENT PLANT ETP | 686194 | 2% | 100% | 2 | Search EFFLUENT+TREATMENT+PLANT+ETP | Search EFFLUENT+TREATMENT+PLANT+ETP |
| 3 | STAIRCASE ELECTROPHORESIS | 686194 | 2% | 100% | 2 | Search STAIRCASE+ELECTROPHORESIS | Search STAIRCASE+ELECTROPHORESIS |
| 4 | UNIDENTIFIED BACTERIA | 686194 | 2% | 100% | 2 | Search UNIDENTIFIED+BACTERIA | Search UNIDENTIFIED+BACTERIA |
| 5 | CATABOLIC POTENTIAL | 617573 | 3% | 60% | 3 | Search CATABOLIC+POTENTIAL | Search CATABOLIC+POTENTIAL |
| 6 | LMW RNA | 441122 | 3% | 43% | 3 | Search LMW+RNA | Search LMW+RNA |
| 7 | 5S RRNA SCAFFOLD | 343097 | 1% | 100% | 1 | Search 5S+RRNA+SCAFFOLD | Search 5S+RRNA+SCAFFOLD |
| 8 | ANTI DNA RNA ANTIBODY | 343097 | 1% | 100% | 1 | Search ANTI+DNA+RNA+ANTIBODY | Search ANTI+DNA+RNA+ANTIBODY |
| 9 | APTAMER 5S RRNA CHIMERA | 343097 | 1% | 100% | 1 | Search APTAMER+5S+RRNA+CHIMERA | Search APTAMER+5S+RRNA+CHIMERA |
| 10 | ARHD | 343097 | 1% | 100% | 1 | Search ARHD | Search ARHD |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | VELAZQUEZ, E , RIVAS, R , DEL VILLAR, M , VALVERDE, A , PEIX, A , MATEOS, PF , VELAZQUEZ, E , MARTINEZ-MOLINA, E , (2006) A NEW APPROACH FOR SEPARATING LOW-MOLECULAR-WEIGHT RNA MOLECULES BY STAIRCASE ELECTROPHORESIS IN NON-SEQUENCING GELS.ELECTROPHORESIS. VOL. 27. ISSUE 9. P. 1732-1738 | 20 | 80% | 0 |
| 2 | VELAQUEZ, E , TRUJILLO, ME , PEIX, A , PALOMO, JL , GARCIA-BENAVIDES, P , MATEOS, PF , VENTOSA, A , MARTINEZ-MOLINA, E , (2001) STABLE LOW MOLECULAR WEIGHT RNA ANALYZED BY STAIRCASE ELECTROPHORESIS, A MOLECULAR SIGNATURE FOR BOTH PROKARYOTIC AND EUKARYOTIC MICROORGANISMS.SYSTEMATIC AND APPLIED MICROBIOLOGY. VOL. 24. ISSUE 4. P. 490 -499 | 15 | 54% | 5 |
| 3 | RIVAS, R , VIZCAINO, N , BUEY, RM , MATEOS, PF , MARTINEZ-MOLINA, E , VELAZQUEZ, E , (2001) AN EFFECTIVE, RAPID AND SIMPLE METHOD FOR TOTAL RNA EXTRACTION FROM BACTERIA AND YEAST.JOURNAL OF MICROBIOLOGICAL METHODS. VOL. 47. ISSUE 1. P. 59-63 | 11 | 69% | 18 |
| 4 | CRUZSANCHEZ, JM , VELAZQUEZ, E , MATEOS, PF , VELAZQUEZ, E , MARTINEZMOLINA, E , (1997) ENHANCEMENT OF RESOLUTION OF LOW MOLECULAR WEIGHT RNA PROFILES BY STAIRCASE ELECTROPHORESIS.ELECTROPHORESIS. VOL. 18. ISSUE 11. P. 1909-1911 | 8 | 100% | 8 |
| 5 | VELAZQUEZ, E , CRUZ-SANCHEZ, JM , RIVAS-PALA, T , ZURDO-PINEIRO, JL , MATEOS, PF , MONTE, E , MARTINEZ-MOLINA, E , CHORDI, A , (2001) YEASTIDENT-FOOD/PROLE FOOD, A NEW SYSTEM FOR THE IDENTIFICATION OF FOOD YEASTS BASED ON PHYSIOLOGICAL AND BIOCHEMICAL TESTS.FOOD MICROBIOLOGY. VOL. 18. ISSUE 6. P. 637-646 | 7 | 50% | 5 |
| 6 | TRUJILLO, ME , VELAZQUEZ, E , MATEOS, PF , MARTINEZ-MOLINA, E , CHORDI-CORBO, A , (2001) ANALYSIS OF STABLE LOW MOLECULAR WEIGHT (LMW) RNA PROFILES OF HYDROCARBON METABOLIZING BACTERIA BY STAIRCASE ELECTROPHORESIS.SYSTEMATIC AND APPLIED MICROBIOLOGY. VOL. 24. ISSUE 2. P. 290-293 | 7 | 50% | 1 |
| 7 | PALOMO, JL , VELAZQUEZ, E , MATEOS, PF , GARCIA-BENAVIDES, P , MARTINEZ-MOLINA, E , (2000) RAPID IDENTIFICATION OF CLAVIBACTER MICHIGANENSIS SUBSPECIES SEPEDONICUS BASED ON THE STABLE LOW MOLECULAR WEIGHT RNA (LMW RNA) PROFILES.EUROPEAN JOURNAL OF PLANT PATHOLOGY. VOL. 106. ISSUE 8. P. 789-793 | 6 | 55% | 10 |
| 8 | TUCKER, DL , KAROUIA, F , WANG, J , LUO, Y , LI, TB , WILLSON, RC , FOFANOV, Y , FOX, GE , (2005) EFFECT OF AN ARTIFICIAL RNA MARKER ON GENE EXPRESSION IN ESCHERICHIA COLI.APPLIED AND ENVIRONMENTAL MICROBIOLOGY. VOL. 71. ISSUE 7. P. 4156-4159 | 8 | 35% | 3 |
| 9 | PITULLE, C , DSOUZA, L , FOX, GE , (1997) A LOW MOLECULAR WEIGHT ARTIFICIAL RNA OF UNIQUE SIZE WITH MULTIPLE PROBE TARGET REGIONS.SYSTEMATIC AND APPLIED MICROBIOLOGY. VOL. 20. ISSUE 1. P. 133-136 | 5 | 71% | 6 |
| 10 | RAYL, AJS , (2002) ABOVE AND BEYOND - UNIVERSAL CLASSIFIER DETECTS BACTERIA IN SPACE - AND ON THE HOMELAND.SCIENTIST. VOL. 16. ISSUE 24. P. 26-27 | 3 | 100% | 0 |
Classes with closest relation at Level 1 |