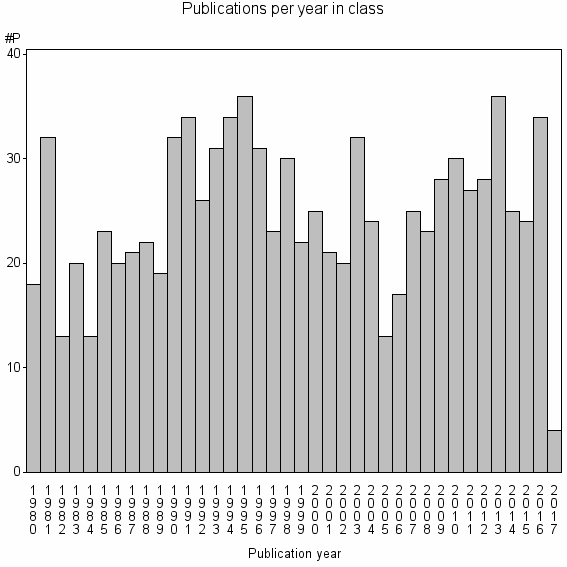
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 12034 | 936 | 32.0 | 57% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 13 | 4 | ZOOLOGY//MARINE & FRESHWATER BIOLOGY//FISHERIES | 1013533 |
| 696 | 3 | OSTRICH//JOURNAL OF AVIAN MEDICINE AND SURGERY//AVIAN | 7063 |
| 3547 | 2 | LYSOZYME//GOOSE TYPE LYSOZYME//G TYPE LYSOZYME | 1351 |
| 12034 | 1 | GOOSE TYPE LYSOZYME//LYSOZYME//G TYPE LYSOZYME | 936 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | GOOSE TYPE LYSOZYME | authKW | 528469 | 2% | 90% | 18 |
| 2 | LYSOZYME | authKW | 510056 | 25% | 7% | 235 |
| 3 | G TYPE LYSOZYME | authKW | 491243 | 2% | 94% | 16 |
| 4 | I TYPE LYSOZYME | authKW | 326217 | 1% | 100% | 10 |
| 5 | C TYPE LYSOZYME | authKW | 319115 | 2% | 65% | 15 |
| 6 | DESTABILASE | authKW | 281940 | 1% | 79% | 11 |
| 7 | LYSOZYME CATALYZED REACTION | authKW | 228352 | 1% | 100% | 7 |
| 8 | DESTABILASE LYSOZYME | authKW | 163109 | 1% | 100% | 5 |
| 9 | INVERTEBRATE LYSOZYME | authKW | 163109 | 1% | 100% | 5 |
| 10 | HUMAN LYSOZYME | authKW | 142404 | 2% | 21% | 21 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 4018 | 54% | 0% | 507 |
| 2 | Food Science & Technology | 795 | 12% | 0% | 108 |
| 3 | Fisheries | 787 | 6% | 0% | 56 |
| 4 | Biophysics | 650 | 10% | 0% | 98 |
| 5 | Biochemical Research Methods | 418 | 8% | 0% | 74 |
| 6 | Biotechnology & Applied Microbiology | 320 | 9% | 0% | 86 |
| 7 | Veterinary Sciences | 293 | 7% | 0% | 68 |
| 8 | Chemistry, Applied | 224 | 5% | 0% | 50 |
| 9 | Agriculture, Dairy & Animal Science | 211 | 4% | 0% | 37 |
| 10 | Marine & Freshwater Biology | 181 | 5% | 0% | 45 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ENZYME SYST SCI | 52190 | 0% | 40% | 4 |
| 2 | LIVESTOCK FISHERY MANAGEMENT | 43494 | 0% | 67% | 2 |
| 3 | PL VET PHYSIOL | 43494 | 0% | 67% | 2 |
| 4 | KATEDRA ZARZADZANIA JAKOSCIA ZYWNOSCI | 36696 | 0% | 38% | 3 |
| 5 | AGR DIRECTORATE LULEBURGAZ DIST | 32622 | 0% | 100% | 1 |
| 6 | BACTERIOL DR ROBERTO GABALDON | 32622 | 0% | 100% | 1 |
| 7 | BEIJING BIOREACT SUBST FUNCT FOODS | 32622 | 0% | 100% | 1 |
| 8 | BIOL BLOOD COAGULAT | 32622 | 0% | 100% | 1 |
| 9 | BIOSCI TECHNOL ANIM GENET | 32622 | 0% | 100% | 1 |
| 10 | BTIS SUBDIC ANIM BIOTECHNOL | 32622 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | FISH & SHELLFISH IMMUNOLOGY | 10633 | 4% | 1% | 38 |
| 2 | JOURNAL OF BIOCHEMISTRY | 9151 | 6% | 1% | 55 |
| 3 | ACTA CRYSTALLOGRAPHICA SECTION D-BIOLOGICAL CRYSTALLOGRAPHY | 4976 | 3% | 1% | 26 |
| 4 | PROTEIN ENGINEERING | 2805 | 1% | 1% | 14 |
| 5 | COMPARATIVE BIOCHEMISTRY AND PHYSIOLOGY B-BIOCHEMISTRY & MOLECULAR BIOLOGY | 1780 | 2% | 0% | 23 |
| 6 | DEVELOPMENTAL AND COMPARATIVE IMMUNOLOGY | 1522 | 1% | 0% | 13 |
| 7 | AGRICULTURAL AND BIOLOGICAL CHEMISTRY | 1487 | 2% | 0% | 18 |
| 8 | JOURNAL OF MOLECULAR EVOLUTION | 1425 | 1% | 0% | 13 |
| 9 | BIOSCIENCE BIOTECHNOLOGY AND BIOCHEMISTRY | 1278 | 2% | 0% | 22 |
| 10 | CELLULAR AND MOLECULAR LIFE SCIENCES | 906 | 1% | 0% | 12 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | GOOSE TYPE LYSOZYME | 528469 | 2% | 90% | 18 | Search GOOSE+TYPE+LYSOZYME | Search GOOSE+TYPE+LYSOZYME |
| 2 | LYSOZYME | 510056 | 25% | 7% | 235 | Search LYSOZYME | Search LYSOZYME |
| 3 | G TYPE LYSOZYME | 491243 | 2% | 94% | 16 | Search G+TYPE+LYSOZYME | Search G+TYPE+LYSOZYME |
| 4 | I TYPE LYSOZYME | 326217 | 1% | 100% | 10 | Search I+TYPE+LYSOZYME | Search I+TYPE+LYSOZYME |
| 5 | C TYPE LYSOZYME | 319115 | 2% | 65% | 15 | Search C+TYPE+LYSOZYME | Search C+TYPE+LYSOZYME |
| 6 | DESTABILASE | 281940 | 1% | 79% | 11 | Search DESTABILASE | Search DESTABILASE |
| 7 | LYSOZYME CATALYZED REACTION | 228352 | 1% | 100% | 7 | Search LYSOZYME+CATALYZED+REACTION | Search LYSOZYME+CATALYZED+REACTION |
| 8 | DESTABILASE LYSOZYME | 163109 | 1% | 100% | 5 | Search DESTABILASE+LYSOZYME | Search DESTABILASE+LYSOZYME |
| 9 | INVERTEBRATE LYSOZYME | 163109 | 1% | 100% | 5 | Search INVERTEBRATE+LYSOZYME | Search INVERTEBRATE+LYSOZYME |
| 10 | HUMAN LYSOZYME | 142404 | 2% | 21% | 21 | Search HUMAN+LYSOZYME | Search HUMAN+LYSOZYME |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | CALLEWAERT, L , MICHIELS, CW , (2010) LYSOZYMES IN THE ANIMAL KINGDOM.JOURNAL OF BIOSCIENCES. VOL. 35. ISSUE 1. P. 127-160 | 123 | 54% | 185 |
| 2 | KAWAMURA, S , TOSHIMA, G , CHIJIIWA, Y , TORIKATA, T , ARAKI, T , (2012) AMINO ACID SEQUENCE OF EGYPTIAN GOOSE EGG-WHITE LYSOZYME AND EFFECTS OF AMINO ACID SUBSTITUTION ON THE ENZYMATIC ACTIVITY.BIOSCIENCE BIOTECHNOLOGY AND BIOCHEMISTRY. VOL. 76. ISSUE 4. P. 691 -698 | 35 | 92% | 4 |
| 3 | UMASUTHAN, N , BATHIGE, SDNK , KASTHURI, SR , WAN, Q , WHANG, I , LEE, JH , (2013) TWO DUPLICATED CHICKEN-TYPE LYSOZYME GENES IN DISC ABALONE HALIOTIS DISCUS DISCUS: MOLECULAR ASPECTS IN RELEVANCE TO STRUCTURE, GENOMIC ORGANIZATION, MRNA EXPRESSION AND BACTERIOLYTIC FUNCTION.FISH & SHELLFISH IMMUNOLOGY. VOL. 35. ISSUE 2. P. 284-299 | 44 | 60% | 6 |
| 4 | WANG, Q , ZHANG, LB , ZHAO, JM , YOU, LP , WU, HF , (2012) TWO GOOSE-TYPE LYSOZYMES IN MYTILUS GALLOPROVINCIALIS: POSSIBLE FUNCTION DIVERSIFICATION AND ADAPTIVE EVOLUTION.PLOS ONE. VOL. 7. ISSUE 9. P. - | 35 | 71% | 11 |
| 5 | WANG, MJ , ZHAO, XL , KONG, XH , WANG, L , JIAO, D , ZHANG, HX , (2016) MOLECULAR CHARACTERIZATION AND EXPRESSING ANALYSIS OF THE C-TYPE AND G-TYPE LYSOZYMES IN QIHE CRUCIAN CARP CARASSIUS AURATUS.FISH & SHELLFISH IMMUNOLOGY. VOL. 52. ISSUE . P. 210 -220 | 31 | 72% | 0 |
| 6 | KAWAMURA, S , CHIJIIWA, Y , MINEMATSU, T , FUKAMIZO, T , VARUM, KM , TORIKATA, T , (2008) THE ROLE OF ARG114 AT SUBSITES E AND F IN REACTIONS CATALYZED BY HEN EGG-WHITE LYSOZYME.BIOSCIENCE BIOTECHNOLOGY AND BIOCHEMISTRY. VOL. 72. ISSUE 3. P. 823 -832 | 29 | 91% | 4 |
| 7 | XU, N , PAN, JL , LIU, SS , XUE, QG , ZHANG, SC , (2014) THREE IN ONE: IDENTIFICATION, EXPRESSION AND ENZYMATIC ACTIVITY OF LYSOZYMES IN AMPHIOXUS.DEVELOPMENTAL AND COMPARATIVE IMMUNOLOGY. VOL. 46. ISSUE 2. P. 508 -517 | 35 | 64% | 3 |
| 8 | SOMBOONPATARAKUN, C , SHINYA, S , KAWAGUCHI, Y , ARAKI, T , FUKAMIZO, T , KLAYNONGSRUANG, S , (2016) A GOOSE-TYPE LYSOZYME FROM OSTRICH (STRUTHIO CAMELUS) EGG WHITE: MULTIPLE ROLES OF HIS101 IN ITS ENZYMATIC REACTION.BIOSCIENCE BIOTECHNOLOGY AND BIOCHEMISTRY. VOL. 80. ISSUE 2. P. 264 -272 | 23 | 85% | 1 |
| 9 | KUWANO, Y , YONEDA, K , KAWAGUCHI, Y , ARAKI, N , ARAKI, T , (2013) THE COMPLETE AMINO ACID SEQUENCE AND ENZYMATIC PROPERTIES OF AN I-TYPE LYSOZYME ISOLATED FROM THE COMMON ORIENT CLAM (MERETRIX LUSORIA).BIOSCIENCE BIOTECHNOLOGY AND BIOCHEMISTRY. VOL. 77. ISSUE 11. P. 2269 -2277 | 27 | 82% | 2 |
| 10 | XUE, QG , HELLBERG, ME , SCHEY, KL , ITOH, N , EYTAN, RI , COOPER, RK , LA PEYRE, JF , (2010) A NEW LYSOZYME FROM THE EASTERN OYSTER, CRASSOSTREA VIRGINICA, AND A POSSIBLE EVOLUTIONARY PATHWAY FOR I-TYPE LYSOZYMES IN BIVALVES FROM HOST DEFENSE TO DIGESTION.BMC EVOLUTIONARY BIOLOGY. VOL. 10. ISSUE . P. - | 41 | 55% | 23 |
Classes with closest relation at Level 1 |