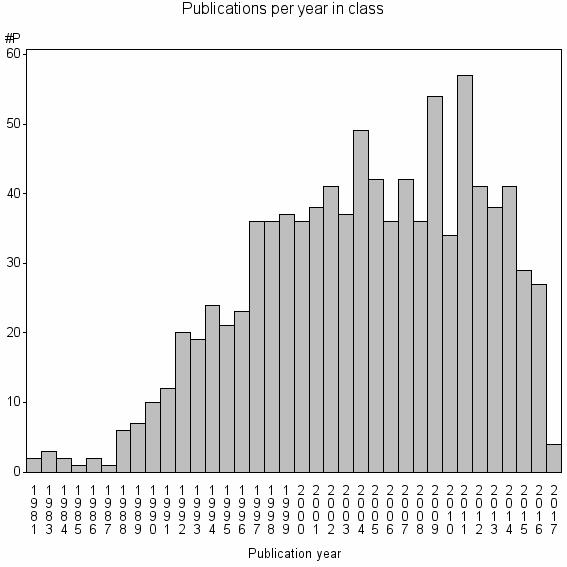
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 11925 | 944 | 41.3 | 89% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 478 | 3 | PHAGE DISPLAY//CELL FREE PROTEIN SYNTHESIS//ANTIBODY ENGINEERING | 23687 |
| 1537 | 2 | CELL FREE PROTEIN SYNTHESIS//INTEIN//PROTEIN SPLICING | 7453 |
| 11925 | 1 | INTEIN//PROTEIN SPLICING//HOMING ENDONUCLEASE | 944 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | INTEIN | authKW | 2979166 | 17% | 56% | 164 |
| 2 | PROTEIN SPLICING | authKW | 2577724 | 11% | 79% | 101 |
| 3 | HOMING ENDONUCLEASE | authKW | 1021373 | 6% | 53% | 60 |
| 4 | SPLIT INTEIN | authKW | 445126 | 2% | 81% | 17 |
| 5 | EXTEIN | authKW | 355798 | 1% | 100% | 11 |
| 6 | PROTEIN TRANS SPLICING | authKW | 288157 | 1% | 64% | 14 |
| 7 | INTRON HOMING | authKW | 217424 | 1% | 61% | 11 |
| 8 | SYNTHET PROT CHEM | address | 192220 | 2% | 26% | 23 |
| 9 | PROTEIN LIGATION | authKW | 176085 | 1% | 39% | 14 |
| 10 | EXPRESSED PROTEIN LIGATION | authKW | 145527 | 2% | 25% | 18 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 6687 | 68% | 0% | 642 |
| 2 | Biotechnology & Applied Microbiology | 569 | 12% | 0% | 110 |
| 3 | Biochemical Research Methods | 453 | 8% | 0% | 77 |
| 4 | Biophysics | 403 | 8% | 0% | 80 |
| 5 | Genetics & Heredity | 401 | 10% | 0% | 93 |
| 6 | Cell Biology | 310 | 11% | 0% | 105 |
| 7 | Microbiology | 114 | 6% | 0% | 57 |
| 8 | Mycology | 59 | 1% | 0% | 11 |
| 9 | Chemistry, Multidisciplinary | 44 | 8% | 0% | 72 |
| 10 | Evolutionary Biology | 25 | 1% | 0% | 14 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | SYNTHET PROT CHEM | 192220 | 2% | 26% | 23 |
| 2 | MACROMOL CRYSTALLOG GRP | 99417 | 2% | 18% | 17 |
| 3 | PROGRAM STRUCT BIOL BIOPHYS | 73254 | 2% | 13% | 18 |
| 4 | PROGRAM MOL BIOPHYS STRUCT DESIGN | 72775 | 0% | 75% | 3 |
| 5 | ADOLF BUTENANDT NEUROBIOCHEM | 32345 | 0% | 100% | 1 |
| 6 | BASKIN COMP ENGN INFORMAT SCI | 32345 | 0% | 100% | 1 |
| 7 | BIO OURCE BIOENVIRONM SCI GENET | 32345 | 0% | 100% | 1 |
| 8 | BIOARCHITECT GRP CELLULAR MOL BIOL | 32345 | 0% | 100% | 1 |
| 9 | BIOMED IRBPROGRAM COMPUTAT BIOL | 32345 | 0% | 100% | 1 |
| 10 | CELLECTIS GENOME SURG | 32345 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | NUCLEIC ACIDS RESEARCH | 9308 | 11% | 0% | 107 |
| 2 | JOURNAL OF MOLECULAR BIOLOGY | 3273 | 5% | 0% | 49 |
| 3 | MOBILE DNA | 1897 | 0% | 2% | 3 |
| 4 | PROTEIN EXPRESSION AND PURIFICATION | 1764 | 2% | 0% | 16 |
| 5 | BIOMOLECULAR NMR ASSIGNMENTS | 1656 | 1% | 1% | 6 |
| 6 | PROTEIN SCIENCE | 1642 | 2% | 0% | 18 |
| 7 | PROTEIN AND PEPTIDE LETTERS | 1525 | 1% | 0% | 11 |
| 8 | PROTEIN ENGINEERING DESIGN & SELECTION | 1513 | 1% | 1% | 7 |
| 9 | ADVANCES IN BIOPHYSICS | 1240 | 0% | 2% | 2 |
| 10 | JOURNAL OF BIOLOGICAL CHEMISTRY | 1073 | 8% | 0% | 74 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | INTEIN | 2979166 | 17% | 56% | 164 | Search INTEIN | Search INTEIN |
| 2 | PROTEIN SPLICING | 2577724 | 11% | 79% | 101 | Search PROTEIN+SPLICING | Search PROTEIN+SPLICING |
| 3 | HOMING ENDONUCLEASE | 1021373 | 6% | 53% | 60 | Search HOMING+ENDONUCLEASE | Search HOMING+ENDONUCLEASE |
| 4 | SPLIT INTEIN | 445126 | 2% | 81% | 17 | Search SPLIT+INTEIN | Search SPLIT+INTEIN |
| 5 | EXTEIN | 355798 | 1% | 100% | 11 | Search EXTEIN | Search EXTEIN |
| 6 | PROTEIN TRANS SPLICING | 288157 | 1% | 64% | 14 | Search PROTEIN+TRANS+SPLICING | Search PROTEIN+TRANS+SPLICING |
| 7 | INTRON HOMING | 217424 | 1% | 61% | 11 | Search INTRON+HOMING | Search INTRON+HOMING |
| 8 | PROTEIN LIGATION | 176085 | 1% | 39% | 14 | Search PROTEIN+LIGATION | Search PROTEIN+LIGATION |
| 9 | EXPRESSED PROTEIN LIGATION | 145527 | 2% | 25% | 18 | Search EXPRESSED+PROTEIN+LIGATION | Search EXPRESSED+PROTEIN+LIGATION |
| 10 | INTRON HOMING SITE | 129381 | 0% | 100% | 4 | Search INTRON+HOMING+SITE | Search INTRON+HOMING+SITE |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | VOLKMANN, G , MOOTZ, HD , (2013) RECENT PROGRESS IN INTEIN RESEARCH: FROM MECHANISM TO DIRECTED EVOLUTION AND APPLICATIONS.CELLULAR AND MOLECULAR LIFE SCIENCES. VOL. 70. ISSUE 7. P. 1185 -1206 | 114 | 90% | 36 |
| 2 | MILLS, KV , JOHNSON, MA , PERLER, FB , (2014) PROTEIN SPLICING: HOW INTEINS ESCAPE FROM PRECURSOR PROTEINS.JOURNAL OF BIOLOGICAL CHEMISTRY. VOL. 289. ISSUE 21. P. 14498 -14505 | 96 | 96% | 14 |
| 3 | SHAH, NH , MUIR, TW , (2014) INTEINS: NATURE'S GIFT TO PROTEIN CHEMISTS.CHEMICAL SCIENCE. VOL. 5. ISSUE 2. P. 446-461 | 84 | 78% | 40 |
| 4 | LI, YF , (2015) SPLIT-INTEINS AND THEIR BIOAPPLICATIONS.BIOTECHNOLOGY LETTERS. VOL. 37. ISSUE 11. P. 2121 -2137 | 91 | 84% | 5 |
| 5 | TOPILINA, NI , MILLS, KV , (2014) RECENT ADVANCES IN IN VIVO APPLICATIONS OF INTEIN-MEDIATED PROTEIN SPLICING.MOBILE DNA. VOL. 5. ISSUE . P. - | 94 | 74% | 14 |
| 6 | ARANKO, AS , WLODAWER, A , IWAI, H , (2014) NATURE'S RECIPE FOR SPLITTING INTEINS.PROTEIN ENGINEERING DESIGN & SELECTION. VOL. 27. ISSUE 8. P. 263 -271 | 68 | 99% | 8 |
| 7 | GUHAN, N , MUNIYAPPA, K , (2003) STRUCTURAL AND FUNCTIONAL CHARACTERISTICS OF HOMING ENDONUCLEASES.CRITICAL REVIEWS IN BIOCHEMISTRY AND MOLECULAR BIOLOGY. VOL. 38. ISSUE 3. P. 199-248 | 142 | 60% | 17 |
| 8 | LIU, XQ , (2000) PROTEIN-SPLICING INTEIN: GENETIC MOBILITY, ORIGIN, AND EVOLUTION.ANNUAL REVIEW OF GENETICS. VOL. 34. ISSUE . P. 61-76 | 73 | 96% | 93 |
| 9 | STODDARD, BL , (2014) HOMING ENDONUCLEASES FROM MOBILE GROUP I INTRONS: DISCOVERY TO GENOME ENGINEERING.MOBILE DNA. VOL. 5. ISSUE . P. - | 73 | 62% | 17 |
| 10 | STODDARD, BL , (2011) HOMING ENDONUCLEASES: FROM MICROBIAL GENETIC INVADERS TO REAGENTS FOR TARGETED DNA MODIFICATION.STRUCTURE. VOL. 19. ISSUE 1. P. 7 -15 | 59 | 69% | 104 |
Classes with closest relation at Level 1 |